* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download PHI-Canto video tutorial slides - PHI-base
Biology and consumer behaviour wikipedia , lookup
X-inactivation wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
History of genetic engineering wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Genetic engineering wikipedia , lookup
Genomic imprinting wikipedia , lookup
Neuronal ceroid lipofuscinosis wikipedia , lookup
Saethre–Chotzen syndrome wikipedia , lookup
Hardy–Weinberg principle wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Metagenomics wikipedia , lookup
Gene therapy wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
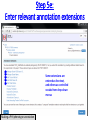
Genome (book) wikipedia , lookup
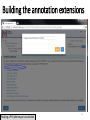
Genome evolution wikipedia , lookup
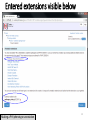
Pharmacogenomics wikipedia , lookup
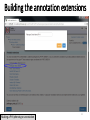
Therapeutic gene modulation wikipedia , lookup
Gene desert wikipedia , lookup
Genome editing wikipedia , lookup
The Selfish Gene wikipedia , lookup
Dominance (genetics) wikipedia , lookup
Gene nomenclature wikipedia , lookup
Gene expression programming wikipedia , lookup
Helitron (biology) wikipedia , lookup
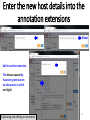
Artificial gene synthesis wikipedia , lookup
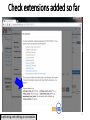
Gene expression profiling wikipedia , lookup
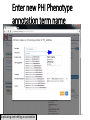
Designer baby wikipedia , lookup
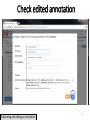
Site-specific recombinase technology wikipedia , lookup
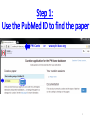
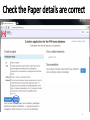
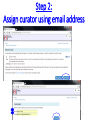
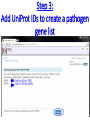
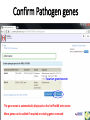
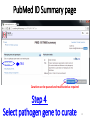
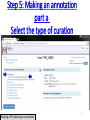
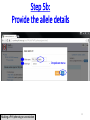
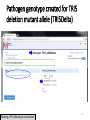
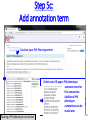
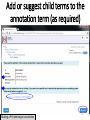
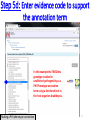
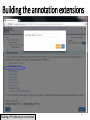
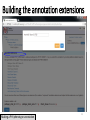
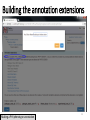
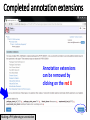
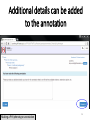
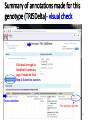
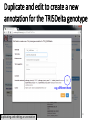
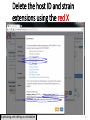
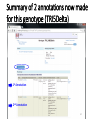
PHI-Canto: The PHI-base Canto tool for author self-curation http://curation.phi-base.org/ 1 PHI-Canto has been adapted from Canto for PomBase (https://curation.pombase.org/) Data capture models Canto captures data like this: gene -> gene annotation -> annotation extensions (for additional annotation detail) In PHI-base, we are trying to capture the data like this: pathogen gene-> host -> pathogen-host interaction annotation For PHI-Canto the tool has been configured to interpret (multi-species PHI-base) data like this: pathogen gene -> pathogen gene annotation -> annotation extensions specifying the host data 2 What you will need to get started The PubMed ID of the publication to be curated The UniProtKB ID(s) of the gene(s) to be curated 3 Example publication for curation One pathogen species: Fusarium graminearum Two pathogen genes: TRI5 and MAP1 Two host species: Arabidopsis and wheat We will annotate 4 PHI phenotype terms and 2 GO terms 4 Step 1: Use the PubMed ID to find the paper PHI-Canto or www.phi-base.org 5 Check the Paper details are correct 6 Step 2: Assign curator using email address 7 Step 3: Add UniProt IDs to create a pathogen gene list UniProt ID for TRI5 UniProt ID for MAP1 8 Confirm Pathogen genes Fusarium graminearum The gene name is automatically displayed as the UniProtKB entry name More genes can be added if required or existing genes removed 9 PubMed ID Summary page TRI5 Curation can be paused and reactivated as required Step 4 Select pathogen gene to curate 10 Step 5: Making an annotation part a Select the type of curation Making a PHI phenotype annotation 11 Step 5b: Provide the allele details Dropdown menu Making a PHI phenotype annotation 12 Pathogen genotype created for TRI5 deletion mutant allele (TRI5Delta) Making a PHI phenotype annotation 13 Step 5c: Add annotation term Curation type: PHI Phenotype term Select one of 9 upper PHI phenotype outcome terms for first annotation. Additional PHI phenotype annotations can be14 made later. Making a PHI phenotype annotation 14 15 Add or suggest child terms to the annotation term (as required) Making a PHI phenotype annotation 16 Step 5d: Enter evidence code to support the annotation term In this example the TRI5Delta genotype resulted in unaffected pathogenicity as a PHI Phenotype annotation term using a functional test in the host organism Arabidopsis. Making a PHI phenotype annotation 17 Step 5e: Enter relevant annotation extensions Some extensions are entered as free text, and others as controlled vocabs from drop down menus Making a PHI phenotype annotation 18 Building the annotation extensions Making a PHI phenotype annotation 19 Entered extensions visible below Making a PHI phenotype annotation 20 Building the annotation extensions Making a PHI phenotype annotation 21 Building the annotation extensions Making a PHI phenotype annotation 22 Building the annotation extensions Making a PHI phenotype annotation 23 Building the annotation extensions Making a PHI phenotype annotation 24 Completed annotation extensions Annotation extensions can be removed by clicking on the red X Making a PHI phenotype annotation 25 Additional details can be added to the annotation Making a PHI phenotype annotation 26 Summary of annotations made for this genotype (TRI5Delta)- visual check Click back through to PubMed ID summary page if ready for final Step 6: Submit to curators One annotation For speedy curation 27 Duplicate and edit to create a new annotation for the TRI5Delta genotype e.g. different host Duplicating and editing an annotation 28 Delete the host ID and strain extensions using the red X Duplicating and editing an annotation 29 Enter the new host details into the annotation extensions Wheat Add in another extension The disease caused by Fusarium graminearum on wheat ears is called ear blight Duplicating and editing an annotation 30 Check extensions added so far Duplicating and editing an annotation 31 Enter new PHI Phenotype annotation term name Duplicating and editing an annotation 32 Check edited annotation Duplicating and editing an annotation 33 Summary of 2 annotations now made for this genotype (TRI5Delta) 1st Annotation 2nd Annotation 34 Genotype summary for publication 2 annotations have now been made to the TRI5Delta pathogen genotype for this Publication 35 PubMed ID summary page TRI5 36 Adding GO term annotation for TRI5 37 Making a GO biological process annotation Adding an annotation for ‘GO biological process’ Making a GO biological process annotation 38 Child term options Making a GO biological process annotation 39 Choose the evidence code for the GO annotation Select from drop down menus Making a GO biological process annotation 40 Add annotation extensions or additional comments for the GO annotation Additional comments Making a GO biological process annotation 41 New GO annotation to the gene TRI5 42 PubMed ID summary page MAP1 Annotating another pathogen gene 43 MAP1 curation MAP1 I have created a map1delta mutant single allele genotype Annotated 2 PHI phenotypes with evidence codes and relevant extensions I have added a GO molecular function annotation term with evidence code to the MAP1 gene Annotating another pathogen gene 44 Summary of annotations made to the MAP1 Gene Annotating another pathogen gene 45 Step 6: Check PubMed ID summary page and submit to curators 2 genes 4 PHI phenotype annotations Submit to curators 2 GO annotations 46 Add any further comments for the curation team 47 Post submission (or a pause) – the author is brought back to this page Recent sessions visible here for quick access Key point 1 : Can only have ONE assigned curator per PubMed ID to prevent duplications Key point 2: Each author can have several papers under curation at the same time 48 Helping the author during curation Key point 3: Each author has various ways to obtain help 49 Our vision Typical Paper: Curation of a single gene deletion in a pathogen tested on two hosts and some in vitro characterisation - Take 15 mins Start with a Pubmed ID use the duplication function manual checking as the annotation extension is built More complex paper: Curation of a single and double gene deletion in a pathogen tested on two or more hosts Detailed microscopy with reporter strains Detailed in vitro characterisation of each strain - Take 30 - 45 mins 50 Thank you for watching this tutorial on PHI-Canto We welcome PHI-Canto feed-back by email to [email protected] 51