* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
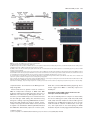
Download ABA overlysensitive5 (ABO5), encoding a pentatricopeptide repeat
Mitochondrial replacement therapy wikipedia , lookup
RNA interference wikipedia , lookup
Signal transduction wikipedia , lookup
Ridge (biology) wikipedia , lookup
Magnesium transporter wikipedia , lookup
Genomic imprinting wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Plant breeding wikipedia , lookup
Point mutation wikipedia , lookup
Transcriptional regulation wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Paracrine signalling wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Secreted frizzled-related protein 1 wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
Gene regulatory network wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Gene expression wikipedia , lookup
Gene expression profiling wikipedia , lookup
The Plant Journal (2010) 63, 749–765 doi: 10.1111/j.1365-313X.2010.04280.x ABA overly-sensitive 5 (ABO5), encoding a pentatricopeptide repeat protein required for cis-splicing of mitochondrial nad2 intron 3, is involved in the abscisic acid response in Arabidopsis Yue Liu1, Junna He1, Zhizhong Chen1, Xiaozhi Ren1, Xuhui Hong1 and Zhizhong Gong1,2,3,* State Key Laboratory of Plant Physiology and Biochemistry, College of Biological Sciences, China Agricultural University, Beijing, 100193, China, 2 China Agricultural University-University of California-Riverside Center for Biological Sciences and Biotechnology, Beijing, 100193, China, and 3 National Center for Plant Gene Research, Beijing, 100193, China 1 Received 21 April 2010; revised 1 June 2010; accepted 8 June 2010; published online 9 July 2010. * For correspondence (fax 86 10 62733733; e-mail [email protected]). SUMMARY To study the molecular mechanism of abscisic acid (ABA) regulation of root development, we screened the root growth of Arabidopsis mutants for sensitivity to ABA. ABA overly-sensitive 5 (ABO5/At1g51965) was identified, and was determined to encode a pentatricopeptide repeat protein required for cis-splicing of mitochondrial nad2 intron 3 (nad2 is one subunit in complex I). Under constant light conditions (24-h light/0-h dark photoperiod), abo5 mutants exhibited various phenotypes and expressed lower transcripts of stressinducible genes, such as RD29A, COR47 and ABF2, and photosynthesis-related genes proton gradient regulation 5 (PGR5) and PGR5-like photosynthetic phenotype (PGRL1), but higher levels of nuclear-encoded genes alternative oxidase 1a (AOX1a) and oxidative signal-inducible 1 (OXI1). Prolonged ABA treatment increased the expression of the cox2 gene in complex IV and nad genes in complex I to a higher level than no ABA treatment in the wild type, but only to a moderate level in abo5, probably because abo5 already expressed high levels of mitochondrial-encoded cox2 and nad genes under no ABA treatment. More H2O2 accumulated in the root tips of abo5 than in the wild type, and H2O2 accumulation was further enhanced by ABA treatment. However, these growth phenotypes and gene-expression defects were attenuated by growing abo5 plants under short-day conditions (12-h light/12-h dark photoperiod). Our results indicate that ABO5 is important in the plant response to ABA. Keywords: mitochondria, PPR protein, ABA signaling, oxidative stress. INTRODUCTION Plants have evolved a series of mechanisms to limit stress and damage caused by unfavorable environmental conditions. Abscisic acid (ABA), a hormone produced when plants are stressed by drought, salt and cold, is an important signal molecule. ABA helps plants cope with these unfavorable stresses, and also plays essential roles in seed development and seedling growth. Genetic screening using seed germination sensitivity to ABA has identified several key mediators in the ABA signaling pathway, including (ABA insensitive 1) ABI1, ABI2, ABI3, ABI4 and ABI5 (Finkelstein and Lynch, 2000; Finkelstein et al., 1998). ABI1 and ABI2, ª 2010 The Authors Journal compilation ª 2010 Blackwell Publishing Ltd which are phosphatase type-2C proteins with negative regulation roles in ABA signaling, physically interact with and inhibit downstream target proteins of ABA signals, such as serine/threonine protein kinase OPEN STOMATA1 (OST1), when ABA content is limited. The increased levels of ABA under abiotic stress cause the ABA receptors PRY1/PYLs to interact with these PP2C proteins and relieve the inhibition on their downstream targeted protein kinases (Fujii et al., 2009; Ma et al., 2009; Nishimura et al., 2009; Park et al., 2009; Santiago et al., 2009). ABI5 is a basic leucine zipper transcription factor that can be phosphorylated and activated 749 750 Yue Liu et al. by SRK2D/SnRK2.2, SRK2E/SnRK2.6/OST1 and SRK2I/ SnRK2.3 to regulate the expression of stress-responsive genes (Nakashima et al., 2009). ABI3 encodes a transcriptional factor that displays high homology with maize viviparous 1 (Giraudat et al., 1992). ABI4 is a member of the ERF/ AP2 transcription factor family (Finkelstein et al., 1998). ABA can stimulate the accumulation of H2O2, which acts as a signal molecule to induce stomatal closure for controlling water consumption by activating calcium channels (Murata et al., 2001; Pei et al., 1997). ABI1 and ABI2 are sensitive to H2O2, which is partially mediated and transduced by a general oxyradical scavenger glutathione peroxidase, ATGPX3 (Miao et al., 2006). The chloroplast and mitochondrion are the two main cellular organelles with independent genomes. Under unfavorable conditions, both chloroplasts and mitochondria cause the accumulation of reactive oxygen species (ROS), which increases oxidative stress and also act as signals to help the plant respond/adapt to the unfavorable environment. Pentatricopeptide repeat (PPR) proteins form a large protein family, are mostly targeted to mitochondria or chloroplasts in plants, and play diverse roles in RNA metabolism (Lurin et al., 2004; O’Toole et al., 2008; Schmitz-Linneweber and Small, 2008). For example, PPR4 from Zea mays is associated with a chloroplast-encoded premRNA, and mediates its splicing (Schmitz-Linneweber et al., 2006). OTP51 is necessary for splicing intron 2 in the putative chloroplast open reading frame 3 (ycf3) mRNA and other introns (de Longevialle et al., 2008). PPR5 binds to the chloroplast trnG-UCC precursor group-II intron and stabilizes it in maize (Beick et al., 2008). PPR10 also stabilizes chloroplast mRNA in Arabidopsis (Pfalz et al., 2009). AtECB2 (Yu et al., 2009), LPA 66 (Cai et al., 2009), RARE1 (Robbins et al., 2009), CHLORORESPIRATORY REDUCTION 22 (CRR22), CRR28 (Okuda et al., 2009) and CRR4 edit certain RNAs in the chloroplast (Kotera et al., 2005). Both OGR1 (opaque and growth retardation 1) and MITOCHONDRIAL RNA EDITING FACTOR 1 (MEF1) are essential for RNA editing in mitochondria of rice (OGR1) or Arabidopsis (MEF1) (Kim et al., 2009; Zehrmann et al., 2009). PPR40 mediates ubiqinol-cytochrome c oxidoreductase activity in complex III, and its impairment leads to the accumulation of ROS and enhanced sensitivity to stress (Zsigmond et al., 2008). PPR-B regulates the translation of orf138 mRNA in radish mitochondria to control cytoplasmic male sterility (Uyttewaal et al., 2008). Most PPR proteins are essential for plant organelle biogenesis, and the disruption of some PPRs leads to embryo lethality (Cushing et al., 2005; Lurin et al., 2004). Mitochondria consist of four major complexes localized on the membranes. Complex I is responsible for transporting electrons from the NADH pool to ubiquinone in the electron transport system. There are 30–40 NADH dehydrogenase subunits encoded by both mitochondrial and nuclear genes. Plant mitochondria have specific alternative respiratory pathways, including the non-proton-pumping NAD(P)H dehydrogenases, which bypass complex I for diminishing ROS production, and an alternative oxidase (AOX), which directly accepts electrons from the ubiquinone pool for reducing ROS production. These alternative pathways increase the tolerance of plant mitochondria to respiratory defects, and loss of complex I has diverse effects on plant cells. For example, mutation in frostbite 1 (fro1), a nuclear gene encoding NADH dehydrogenase co-enzyme in the mitochondrial electron transfer chain, causes reduced transcription of cold-inducible genes, a high accumulation of ROS and decreased cold acclimation (Lee et al., 2002). In contrast, loss of complex-I function in a cytoplasmic malesterile mutant (CMSII) did not cause a large increase in ROS accumulation, but instead caused a marked increase in antioxidant activities and the re-establishment of a new redox homeostasis in tobacco cells (Dutilleul et al., 2003b). Non-chromosomal stripe (NCS) mutants truncated in the nad4 gene of complex I show increased expression of AOX. PPR protein OTP43 is necessary for splicing intron 1 in the nad1, a central anchor component of mitochondrial complex I in Arabidopsis thaliana, and mutation in OTP43 increases levels of AOX (de Longevialle et al., 2007). Mutations affecting different subunits of complex I result in various growth phenotypes (de Longevialle et al., 2007; Dutilleul et al., 2003b; Lee et al., 2002; Nakagawa and Sakurai, 2006), indicating the diverse effects of complex I. Organellar proteins are encoded by both organellar and nuclear genes. Communication between organelles and the nucleus is essential for plant responses to stressful environmental conditions, and for adjusting the requirements for plant development (Pogson et al., 2008). Retrograde signals from organelles that regulate the expression of nuclear genes represent an important feedback control for plant responses to environmental and developmental limitations. Several signaling pathways involved in chloroplast-tonucleus retrograde signaling have been described previously (Surpin et al., 2002; Woodson and Chory, 2008). The existence of multiple mitochondrial retrograde signaling pathways that are capable of initiating specific gene expression in the nucleus is also suggested by plant responses to the dysfunction of specific mitochondrial proteins (Rhoads and Subbaiah, 2007). One target of mitochondrial retrograde signaling is the upregulation of AOX expression, which helps to ameliorate the ROS stress initiated by mitochondrial dysfunction (Umbach et al., 2005). Cross-talk occurs between mitochondria and chloroplasts, and retrograde signals from one organelle that directly or indirectly influence another organelle require further study (Dutilleul et al., 2003b; Jiao et al., 2005; Matsuo and Obokata, 2006). Using a root-bending assay we have identified mutants in which root growth is sensitive to ABA, and have isolated several mutants, including those with a mutation in DNA ª 2010 The Authors Journal compilation ª 2010 Blackwell Publishing Ltd, The Plant Journal, (2010), 63, 749–765 ABA and PPR protein 751 polymerase e and several elongator subunits (Yin et al., 2009; Zhou et al., 2009). Here, we report on a new PPR protein, ABA overly-sensitive 5 (ABO5), that is located in mitochondria and participates in the cis-splicing of intron 3 of nad2 in mitochondrial complex I. abo5 showed various growth phenotypes and changed the expression of both nuclear and mitochondrial genes in a photoperiod-dependent manner. Our results suggest that ABO5 is an important mediator in plant responses to ABA. RESULTS The abo5 mutant shows growth retardation and is sensitive to ABA The abo5 mutant was identified during a genetic screen for mutants exhibiting root-growth sensitivity to 30 lM ABA in a root-bending assay performed on a T-DNA insertion mutant pool. ABA inhibits the early seed germination process, including testa and endosperm rupture, post-germination growth (PGG), and the growth of root and shoot. Here, we define PGG as the percentage of seeds that produce seedlings with green cotyledons during early seedling growth. Genetic analysis indicated that the abo5 mutant was caused by a single recessive mutation. Because PGG is more sensitive to ABA than seedling growth (i.e. growth subsequent to PGG), we first compared PGG on MS medium supplemented with 0.1 lM ABA under a photoperiod of 24-h light/0-h dark (24/0). As shown in Figure 1a,b, PGG without ABA was lower in abo5 than in the wild type. On MS medium containing 0.1 lM ABA, the relative PGG (PGG with ABA expressed as a percentage of PGG without ABA) was much lower for abo5 than for the wild type during 128 h (Figure 1c). Because sugar could increase ABA sensitivity during seed germination, we measured the effects of sugar on PGG of abo5 and the wild type (Figure 1d,e). Without sugar, PGG differed only slightly between abo5 and the wild type. After 72 h on medium containing 3% sucrose, only 12.5% of abo5 exhibited PGG compared with 97.3% of the wild type. After 96 h on MS medium containing 3% sucrose, PGG had increased to 72.2% in abo5. After 72 h on MS medium containing 3% glucose, the germination percentage was 48.6% for abo5 and 97.2% for the wild type. After 96 h on MS medium containing 6% glucose, neither abo5 nor the wild type exhibited any PGG. Next, we transferred 5-day-old seedlings to MS containing different concentrations of ABA and measured the root growth after 6 days. As shown in Figure 1f,g, the root growth of abo5 was more inhibited by ABA than the wildtype root growth. The contents of ABA, sucrose, and total soluble sugars did not differ between abo5 and the wild type (Figure 1i–k). Similarly, the sucrose content of the CMSII mutant is not different from the wild type (Dutilleul et al., 2003a). These results indicate that abo5 is more sensitive than the wild type to both ABA and sugar in terms of PGG, but is especially sensitive to ABA in terms of root growth. Genetic analysis of abo5 with abi1-1, abi2-1, abi3-1, abi4-1 and abi5 mutants To clarify the role of ABO5 in the ABA response, we analyzed the double mutants of abo5 combined with each of five classic ABA-insensitive mutants: abi1-1, abi2-1, abi3-1, abi41 and abi5. abi1-1 and abi2-1 are two dominative negative mutants (Leung et al., 1997; Meyer et al., 1994), whereas abi3-1 (Giraudat et al., 1992), abi4-1 (Finkelstein et al., 1998) and abi5 (Finkelstein and Lynch, 2000) are loss-of-function mutants. Seed germination of all five mutants is insensitive to ABA. The seed germination of the abo5 abi1-1, abo5 abi21, abo5 abi3-1, abo5 abi4-1 and abi5 abo5 double mutants had the same ABA-insensitive phenotype as the abi single mutants (Figure 1h, right lane). When the response of root growth to ABA was measured, abo5 abi1-1 and abo5 abi2-1 were similar to abi1-1 and abi2-1 in that they were insensitive to ABA, but abo5 abi3-1, abo5 abi4-1 and abo5 abi5 differed from abi3-1, abi4-1 and abi5 in that they were as sensitive to ABA as abo5 (Figure 1h, left lane). Given that root growth of abi3-1, abi4-1, and abi5 is insensitive to ABA, we conclude that the sensitivity of root growth to ABA of the abo5 abi3-1, abo5 abi4-1 and abo5 abi5 mutants is mainly caused by the abo5 mutation. These results suggest that ABO5 is part of the general response to ABA. ABO5 encodes a PPR protein localized in the mitochondrion Because the abo5 mutant was obtained from a T-DNA insertion library, we used TAIL-PCR to clone the targeted gene. The T-DNA was inserted in the second exon of At1g51965 (Figure 2a). Insertion of T-DNA caused premature transcription termination, and the product of ABO5 mRNA was shorter in the abo5 mutant than in the wild type (Figure 3a). We acquired three T-DNA insert SALK lines, and each T-DNA was inserted in the promoter region of At1g51965. RT-PCR using the total RNA isolated from each T-DNA and abo5 indicated that the transcripts were equally detected in the wild type and three SALK lines, but not in the abo5 mutant (Figure 2b). SALK lines with homozygous T-DNA did not exhibit any growth phenotype or ABAsensitive phenotypes (data not shown). These results indicate that At1g51965 is not disrupted by T-DNA insertions in three SALK lines. To further confirm that abo5 is caused by At1g51965, we crossed abo5 with the Landsberg accession, and performed map-based cloning using the segregated abo5 mutants identified from F2 seedlings, based on the root sensitivity to ABA. As shown in Figure 2c, abo5 was limited to the same T-DNA locus region. ABO5 cDNA was obtained by RT-PCR from total RNAs isolated from young seedlings. ABO5 encodes a putative P-subfamily PPR protein with 650 amino acids. ABO5 contains one signal peptide for mitochondria at the N terminal, two PPR repeats in the middle and four PPR repeats at the C terminal (Figure 2d). Because the full-length ABO5 ª 2010 The Authors Journal compilation ª 2010 Blackwell Publishing Ltd, The Plant Journal, (2010), 63, 749–765 752 Yue Liu et al. (a) (b) (c) (f) (d) (e) (g) (h) (i) (j) (k) Figure 1. The phenotypes of abo5 mutants. (a) The wild type and abo5 were germinated on and grown on MS medium without or with 0.1 lM ABA, and with a photoperiod of 24/0 (24 h of light and 0 h of dark). Seedlings were photographed after 10 days. (b) Post-germination growth (PGG) of the wild type and abo5 on MS medium. PGG indicates the percentage of total seeds (>40 per plate) that germinated and produced green cotyledons. Values are means SEs (n = 3). (c) PGG of the wild type and abo5 as affected by ABA. Relative PGG is PGG with ABA expressed as a percentage of PGG without ABA. Values are means SE. (d) and (e) Relative PGG of the wild type and abo5 on MS medium containing different concentrations of glucose (d) and sucrose (e). Data are means SEs (n = 3). (f) Root growth of the wild type and abo5 on MS medium containing 0 or 50 lM ABA. (g) Root growth of the wild type and abo5 as affected by ABA. Relative root growth is root length in the presence of ABA expressed as a percentage of root length in the absence of ABA. Values are means SE (n = 35). (h) Genetic analysis of abo5 with abi1-1, abi2-1, abi3-1, abi4-1 and abi5. Four-day-old seedlings were transferred to MS medium (upper lane, left) or MS medium containing 30 lM ABA for 5 days (lower lane, left), or different seeds were directly sown on MS medium (upper lane, right) or MS medium containing 1 lM ABA (lower lane, right) for 8 days before seedlings were photographed. (i) Comparison of soluble sugar contents in the wild type (WT) and abo5 under 24/0 and 12/12 photoperiods. (j) Comparison of sucrose contents in the wild type (WT) and abo5 under 24/0 and 12/12 photoperiods. (k) ABA contents in the wild type and abo5 under a 24/0 photoperiod. fused with GFP failed to emit any florescence, we constructed a fused protein containing the ABO5 N-terminal signal peptide fused in frame with GFP, and transiently expressed it in leaf protoplast cells. The fused protein was co-localized with mitochondrial-specific staining marker (Figure 2e), suggesting that ABO5 is localized in mitochondrion. In the complex-I mutant NADH dehydrogenase (ubiquinone) fragment S subunit 4 (ndufs4) and the complex-III mutant ppr40, a similar delay in seed germination was observed, indicating that the components in complex I and III are critical for seed germination (de Longevialle et al., 2007; Meyer et al., 2009). Seed germination was even more sensitive to ABA in ndufs4 than in abo5 (Figure 2f), suggesting that both ABO5 and NDUFS4 are critical components in plant response to ABA. We overexpressed the cDNA of At1g51965/ABO5 in abo5 under the control of a cauliflower mosaic virus 35S promoter, and isolated five independent transgenic lines: all of them complemented the retarded-growth phenotypes and the ABA-sensitive phenotypes. Figure 2g,h show one transgenic line (line 3) of the T3 generation that ª 2010 The Authors Journal compilation ª 2010 Blackwell Publishing Ltd, The Plant Journal, (2010), 63, 749–765 ABA and PPR protein 753 (a) (f) (b) (g) (c) (d) (h) (e) Figure 2. Cloning of the ABO5 gene and mutant complementation. (a) T-DNA insertion sites in abo5 and three SALK lines. (b) ABO5 transcript in abo5 and three T-DNA insertion lines. The primers used (F and R) are indicated by arrows in (a). actin4 was amplified as the loading control. (c) Map-based cloning of ABO5. See Experimental procedures for details. (d) ABO5 contains one mitochondrial signal peptide (labeled mt and colored black) and six PPR domains (labeled 1–6 and shaded grey). The position of T-DNA insertion in abo5 is indicated. (e) ABO5 N-terminal-GFP subcellular location. The subcellular location of ABO5 mitochondrial signal peptide was determined by transforming ABO5 N terminal fused with GFP in frame into leaf protoplast. The GFP fluorescence was observed with a confocal microscope (left lane). At the same time, mitochondria were stained in red by Mito-tracker (Invitrogen) (middle lane). The right lane shows the merged GFP and Mito-tracker. Scale bar = 10 lM. (f) Seed germination of the wild type, abo5 and ndufs4 on MS medium or MS medium supplemented with 0.3 lM ABA. The seeds were sown on medium and cultured in a growth chamber for 7 days before the seedlings were photographed. (g) Root growth of the wild type, abo5 and abo5 carrying p35S-ABO5 cDNA (line 3). Four-day-old seedlings were transferred from MS medium to MS medium or MS medium containing 50 lM ABA with a 24/0 photoperiod (full light) for 6 days. (h) Growth of the wild-type, abo5, and transgenic plant line 3 seedlings in soil under normal conditions (an 18/6 photoperiod). An arrow points to an abo5 seedling. complemented the abo5 mutant in both ABA hypersensitivity and growth. Because PGG and root growth of abo5 are sensitive to ABA, we analyzed the transcripts of ABO5 under ABA treatment. Northern blot indicated that the expression of ABO5 was not influenced by ABA (Figure 3a). We constructed a promoter-GUS binary vector and transformed it into the wild type. GUS staining was determined using transgenic seedlings expressing pABO5-GUS. As shown in Figure 3b–f, GUS activity was greater in flowers, sliques, root tips and young leaves than in mature leaves, stems and elongated roots. Quantitative RT-PCR (qRT-PCR) indicated that ABO5 was differentially expressed in various parts, which was consistent with GUS staining (Figure 3g). These results suggest that ABO5 is universally expressed in Arabidopsis. abo5 impairs nad2 pre-RNA splicing and increases the transcripts of mitochondrial genes Out of more than 40 proteins in complex I of mitochondria, nine are encoded by genes (nad1, nad2, nad3, nad4, nad4L, nad5, nad6, nad7 and nad9) in the mitochondrial genome (Heazlewood et al., 2003). Among them, the exons of nad1, nad2 and nad5 are scattered on different sites of the genome, and require both cis- and trans-splicing for mRNA maturation. A previous study suggested that the PPR protein ª 2010 The Authors Journal compilation ª 2010 Blackwell Publishing Ltd, The Plant Journal, (2010), 63, 749–765 754 Yue Liu et al. Con (a) WT ABA abo5 WT abo5 (b) (c) ABO5 TUB Relative expression (g) 6 5 4 3 2 1 0 ABO5 (f) (e) Root Stem (d) Figure 3. Expression pattern of the ABO5 gene in seedlings. (a) Expression of ABO5 in the wild type and abo5 before or after ABA treatment. Two-week-old seedlings were treated with 50 lM ABA for 5 h. Tubulin was used as the loading control. A small truncated band was detected in abo5. (b–f) GUS activity of transgenic seedlings carrying the ABO5 promoter-GUS. Activity in stem and mature leaf (b), in a 1-day-old seedling (c), in a root tip (d), in a 1-week-old seedling (e), and in a flower and silique (f). (g) Quantitative RT-PCR for relative expression of the ABO5 gene in root, stem, leaf, flower and silique. Values are means SEs (n = 3). Leaf Flower Silique OTP43 is required for the trans-splicing of the nad1 intron 1 (de Longevialle et al., 2007). Because PPR proteins are mainly involved in organelle RNA metabolism, we first examined the transcripts of mitochondrial genes nad1, nad2 and nad5. For each gene, we selected two fragments that cover two different exons separated by other genes, and we used these fragments as probes. As shown in Figure 4a, the splicing of nad2 was impaired in the abo5 mutant, but a very low level of correctly spliced nad2 transcripts could still be detected. nad2 encodes NADH dehydrogenase subunit 2 in complex I. qRT-PCR was performed to identify the intron(s) in which the splicing was affected. The qRT-PRC conditions used here (which were suitable for amplifying a 300–500-bp fragment) could not amplify the fragment containing the large introns in nad2 (Figure 4bi). We found that cis-splicing of nad2 intron 3 was defective in the abo5 mutant, and the splicing of other introns was not affected by abo5 mutation (Figure 4bii). Instead, of the transcripts covering introns 1 and 2, two were elevated to higher levels in abo5 than in the wild type, whereas the transcripts covering intron 4 were unchanged (Figure 4bii). To determine whether the fragment covering exons 3 and 4 was amplified because of stabilization of nad2 precursor transcripts, we compared the intron-3 transcripts in abo5 with the wild type by using primers inside intron 3. qRT-PCR indicated that there were twice as many intron-3 transcripts in abo5 than in the wild type (Figure 4biii). We also compared two other complex-I cis-splicing genes in abo5 and the wild type, nad4 and nad7 (Figure 1a), and complex IV cis-splicing gene cox2 (Figure 4c), but we did not find a splicing defect. However, all of the genes tested exhibited higher transcriptional levels in abo5 than in the wild type, including nad6, which does not contain any intron (Figure 4d). Complementing the abo5 mutant with a wild-type ABO5 cDNA fully restored the splicing of nad2 (Figure 4e). These results indicate that the abo5 mutation impairs the cis-splicing of the nad2 intron 3, and increases the transcripts of all mitochondrial genes tested, but determining whether ABO5 has other biological roles besides the splicing of the nad2 intron 3 will require further study. The transcripts of nad genes are highly induced by ABA in the wild type, but are only moderately induced in the abo5 mutant during prolonged ABA treatment We analyzed the transcript changes of cox2 and nad genes under ABA treatment for 5 h by northern blot, but did not detect a clear difference between ABA-treated and untreated samples (Figure 4a,c,d). We then monitored the expression of cox2 and nad genes after prolonged ABA treatment. The 7-day-old seedlings of abo5 and the wild type grown on MS medium were transferred to MS medium containing 10 lM ABA or no ABA: after 7 days, the expression of cox2 and nad genes was examined by qRT-PCR (Figure 5a). Consistent with the results in Figure 4a,c,d, the abo5 mutation led to an enhanced expression of all nad genes and cox2 relative to the wild type, but nad and cox2 transcripts in abo5 were only slightly increased by ABA treatment. In contrast, nad and cox2 transcripts were significantly increased by ABA treatment in the wild-type seedlings. The transcripts induced by ABA in the wild type were comparable with those in the abo5 mutant (Figure 5a). These results suggest that the transcripts of cox2 and nad genes are induced by ABA only with prolonged treatment, and that the abo5 mutation reduces the response of cox2 and nad genes to such prolonged ABA treatment. The abo5 mutant increases the transcripts of AOX1a Complex-I dysfunction usually increases alternative pathways as an adaptive response to mitochondrial stress, as shown in several previous studies (de Longevialle et al., 2007; Dutilleul et al., 2003b; Karpova et al., 2002). AOX is a terminal oxidase that can be induced by various stresses. AOX in Arabidopsis includes two subfamilies with five members, including AOX1a, AOX1b, AOX1c, AOX1d and AOX2 (Strodtkotter et al., 2009). AOX1a is a marker for mitochondrial retrograde response. Previous study indicates that the expression of AOX1a is directly regulated by ABI4 (Giraud et al., 2009). We used a northern blot to determine whether the abo5 mutant alters the transcripts of AOX1a (Figure 5b). Under 24 h of light, more transcripts ª 2010 The Authors Journal compilation ª 2010 Blackwell Publishing Ltd, The Plant Journal, (2010), 63, 749–765 ABA and PPR protein 755 (a) (b) (i) (ii) (c) (iii) (e) (d) Figure 4. Expression levels of mitochondria complex-I subunit genes in abo5 and the wild type. (a) Expression and splicing pattern of complex-I subunit genes that require trans- and/or cis-splicing with or without ABA treatment. Probes were designed to detect each independent pre-mature transcript. rRNAs stained with ethidium bromide were used as loading controls (lower lane). Two-week-old seedlings were treated with 50 lM ABA for 5 h. Expression and splicing patterns were not affected by 5 h of ABA treatment. Note that the mature mRNA (1.497 kb) band (indicated by an arrow) of nad2 was greatly reduced in abo5. (b) i, Diagram of nad2 transcripts. The pre-transcript of nad2 requires both cis- and trans-splicing, as indicated. ii, Quantitative RT-PCR for ABO5 potential target position. Primers were designed as shown in the upper diagram for nad2 to amplify the fragment covering each intron. The relative expression of each fragment amplified by qRT-PCR was compared. Fragments 1, 3 and 4 require cis-splicing, whereas fragment 2 requires trans-splicing. Only the mature mRNAs could be amplified; the longer intron of pre-mRNA could not be amplified under the experimental conditions. iii, Measurement of intron 3 by qRT-PCR. The primers for intron 3 amplification are indicated with arrows in b–i. Values are means SEs (n = 3). (c) Expression of complex IV cox2, which requires cis-splicing. Samples were treated with or without 50 lM ABA for 5 h. rRNAs stained with ethidium bromide served as loading controls. (d) Expression pattern of complex I subunit nad6, which does not require splicing. Samples were treated with or without 50 lM ABA for 5 h. rRNAs stained with ethidium bromide served as loading controls. (e) Defect in nad2 splicing was recovered in complementary line 3 (Figure 2g). rRNAs stained with ethidium bromide served as loading controls. were detected in the abo5 mutant than in the wild type. ABA treatment increased AOX1a expression in both abo5 and the wild type. Because the basic level of AOX1a is higher in abo5 than in the wild type, it is likely that ABA did not induce more transcripts of AOX1a in abo5 than in the wild type. qRT-PCR confirmed that the relative increase of AOX1a caused by ABA was similar in abo5 and in the wild type (Figure 9g), suggesting that the high expression of AOX1a in abo5 might not be directly affected by ABI4 transcription because ABI4 expression in abo5 was not changed by ABA treatment (data not shown). The OXI1 gene, which encodes a serine/threonine kinase, is induced by both biotic and abiotic stress, and is an important component in oxidative burst-mediated signaling (Rentel et al., 2004). We compared the expression of OXI1 at different times after ABA treatment (Figure 5c). Before ABA treatment, OXI1 expression was very low in abo5 and almost undetectable in the wild type. ABA treatment for 24 h increased the expression of OXI1 more in abo5 than in the wild type, suggesting that abo5 might suffer from oxidative stress. ª 2010 The Authors Journal compilation ª 2010 Blackwell Publishing Ltd, The Plant Journal, (2010), 63, 749–765 756 Yue Liu et al. (a) (b) (d) (c) (e) Figure 5. Expression pattern of cox2, nad genes, alternative oxidase gene AOX1a, oxidative responsive gene OXI1 and proline-related genes in abo5 under ABA treatment. (a) Relative expression of complex-I subunits nad1, nad4, nad5, nad6 and nad7, and complex-IV subunit cox2 in abo5 and the wild type after 10 lM ABA treatment or without ABA treatment for 7 days. Values are means SEs (n = 3). (b) AOX1a expression in abo5 and the wild type treated with ABA. Two-week-old seedlings grown under a 24/0 photoperiod (full light) were treated with 50 lM ABA for 0, 0.5, 1, 3, 5 and 24 h before they were subjected to northern blot analysis using 32P-labeled AOX1a cDNA as the probe. rRNAs were used as the loading control. (c) OXI1 expression analysis in abo5 and the wild type. Two-week-old seedlings grown under a 24/0 photoperiod (full light) were treated with 50 lM ABA for 0, 0.5, 1, 3, 5, 8, 12 and 24 h. OXI1 expression was detected by northern blot. rRNAs stained with ethidium bromide served as loading controls. (d) The expression of genes participating in proline metabolism was analyzed by northern blot. The transcripts of the P5CR, ProDH and GDH1 genes were higher in abo5 than in the wild type, but were not changed by ABA treatment. Two-week-old seedlings grown under a 24/0 photoperiod (full light) were treated with or without 50 lM ABA for 5 h. Tubulin was used as the loading control. (e) Proline contents in abo5 and the wild type. Seedlings were grown on MS medium containing different concentrations of ABA for 2 weeks under a 24/0 photoperiod (full light). abo5 mutants accumulate increased levels of proline and exhibit increased expression of P5CR and other nuclear genes that encode mitochondrial proteins To adapt to various environmental stresses, plants accumulate the compatible osmolyte proline. In the biosynthesis of proline in the cytosol, the enzymes pyrroline-5-carboxylate synthase (P5CS) and pyrroline-5-carboxylate reductase (P5CR) catalyze the last two successive reductions from glutamate. Northern blot analysis indicated that P5CS was induced to the same level in abo5 and in the wild type by ABA treatment (Figure 5c). P5CR was not induced by ABA, but its expression was higher in abo5 than in the wild type. GDH1 (glutamate dehydrogenase 1) is localized in mitochondria and catalyzes the reversible amination of 2-oxoglutarate to glutamate in vitro. The expression of GDH1 was also higher in abo5 than in the wild type, and was not induced by ABA. Proline is oxidated by proline dehydroge- nase (AtProDH) to produce pyrroline-5-carboxylate (P5C), which is subsequently oxidated by P5C dehydrogenase (P5CDH). The expression of AtProDH was higher in abo5 than in the wild type, and was not induced by ABA. Proline content in both abo5 and the wild type was induced by ABA treatment, and abo5 accumulated more proline than the wild type, regardless of the ABA concentration in the medium (Figure 5d). Similarly, various metabolites including proline are accumulated to a higher level in the complex-I mutant ndufs4 than in the wild type (Meyer et al., 2009). abo5 mutants accumulate increased levels of H2O2 in root tips The growth of abo5 primary roots is sensitive to ABA, and ABO5 is involved in the splicing of intron 3 in the complex-I nad2 gene, suggesting that, like the fro1 mutant, the abo5 mutant might accumulate more ROS than the wild type (Lee et al., 2002). We measured the H2O2 level in root tips by ª 2010 The Authors Journal compilation ª 2010 Blackwell Publishing Ltd, The Plant Journal, (2010), 63, 749–765 ABA and PPR protein 757 transcriptional factor that regulates the expression of genes in the downstream of the ABA signaling pathway (Choi et al., 2000; Fujita et al., 2005; Kim et al., 2004). The expression of ABF2 was less in abo5 than in the wild type (Figure 7c). ABF2regulated RD29B is strongly induced by ABA, and its expression was also less in abo5 than in the wild type (Figure 7d). Other stress-inducible genes such as RD22 (Yamaguchi-Shinozaki and Shinozaki, 1993), RAB18 (Lang and Palva, 1992), DREB2A (Liu et al., 1998) and MYB2 (Abe et al., 2003) were all less induced by ABA in abo5 than in the wild type (Figure 7e–h). These results suggest that abo5 mutants were impaired in ABA-regulated gene expression. 2¢,7¢-dichlorfluorescein diacetate (DCFH-DA) staining (Figure 6a,b). Seedlings (1 week old) were treated with ABA for 5 h and stained with DCFH-DA. Fluorescence was examined by a MicroRadiance laser-scanning confocal microscope, and the data were quantified by METAMORPH 6.1. In the root tip region, abo5 accumulated more H2O2 under a 24/0 photoperiod (Figure 6a,c) than under a 12/ 12 photoperiod (Figure 6b,c), and abo5 accumulated much more H2O2 than the wild type after ABA treatment under a 24/0 photoperiod (Figure 6a,c), but not under a 12/12 photoperiod (Figure 6b,c). We also measured H2O2 content in roots by another quantitative method and obtained similar results (Figure 6d). These results suggest that abo5 accumulates more H2O2 than the wild type in the root tips under a 24/0 photoperiod. abo5 phenotypes are attenuated by reducing the length of light exposure In the mutant CMSII, plant growth was greatly inhibited by strong light, suggesting that the defect in complex I impairs photosynthetic carbon assimilation under higher irradiance (Priault et al., 2006). We found that the retarded growth and ABA-sensitive phenotype of abo5 could be attenuated (i.e. growth could be increased) when the photoperiod was changed from a 24/0 to a 12/12 photoperiod. Wild-type seedlings grew more slowly under a 12/12 photoperiod than under a 24/0 photoperiod (Figure 8a). In contrast, abo5 seedlings grew better under a 12/12 photoperiod than under a 24/0 photoperiod, and the growth rates of abo5 and the wild type were almost the same under a 12/12 photoperiod on MS medium (Figure 8a). A 12/12 photoperiod also reduced the difference in growth (PGG and relative root growth) between abo5 and the wild type on a medium containing ABA (Figure 8b,c). We also measured proline abo5 mutants produced fewer transcripts of some ABA-inducible genes Previous study with fro1 indicates that impairment of complex I increases ROS accumulation while suppressing cold-inducible genes (Lee et al., 2002). We compared the transcripts of several ABA-inducible marker genes between abo5 and the wild type under a 24/0 photoperiod. COR47 and RD29A are induced by various abiotic stresses such as cold, ABA and drought. We found that ABA induced more transcripts of both COR47 and RD29A at 3 h in the wild type than in abo5, and that the response to ABA was delayed in abo5, i.e. the level attained at 3 h in the wild type was attained at 5 h in abo5 (Figure 7a,b). ABRE-binding bZIP proteins (ABF2)/ABSCISIC ACID-RESPONSIVE ELEMENT BINDING PROTEIN 1 (AREB1) is an ABA-induced ABRE-binding bZIP Con (a) ABA Con (b) ABA (i) (ii) (i) (ii) (iii) (iv) (iii) (iv) WT abo5 (d) 120 WT 100 24/0 H2O2 content (nm/g FW) (c) Arbitrary unit Figure 6. Detection of reactive oxygen species (ROS) in the root tips of abo5 and the wild type. (a) Detection of H2O2 in the root-tip region of 1-week-old seedlings grown under a 24/0 photoperiod (full light). (i) Wild type (without ABA treatment); (ii) wild type treated with 50 lM ABA for 5 h; (iii) abo5 (without ABA treatment); (iv) abo5 treated with 50 lM ABA for 5 h. H2O2 was stained by DCFH-DA, as described in Experimental procedures. (b) H2O2 accumulation in the root-tip region of seedlings grown under a 12/12 photoperiod. (i) Wild type; (ii) wild type treated with 50 lM ABA for 5 h; (iii) abo5; (iv) abo5 treated with 50 lM ABA for 5 h. (c) Quantitative analyses of DCFH-DA stain fluorescence in (b) and (c). Values are means SEs (n = 13). (d) Quantitative measurement of H2O2 content in root tips by another method. Con, control without ABA. abo5 24/0 80 WT 60 abo5 12/12 12/12 40 20 0 Con ABA ª 2010 The Authors Journal compilation ª 2010 Blackwell Publishing Ltd, The Plant Journal, (2010), 63, 749–765 WT 200 WT 100 0 24/0 abo5 24/0 12/12 abo5 12/12 Con ABA 758 Yue Liu et al. (a) (b) (c) (f) (d) (g) (e) (h) (a) (b) (c) (d) contents under a 12/12 photoperiod, and found no obvious difference between the wild type and abo5 (Figure 8d). Gene expression patterns change under a 12/12 photoperiod To clarify the impact of photoperiod on gene expression, we used qRT-PCR to analyze mitochondrial gene expression of Figure 7. Expression of stress-responsive genes in abo5. (a) The expression of Cor47 was analyzed by northern blot. Two-week-old seedlings grown under a 24/0 photoperiod (full light) were treated with a 50 lM ABA solution for 0, 1, 3, 5, 8 or 24 h. Tubulin was used as the loading control. (b) RNA gel analysis of RD29A expression in the wild type and abo5. Two-week-old seedlings grown under a 24/0 photoperiod (full light) were treated with a 50 lM ABA solution for 0, 1, 3 or 5 h. rRNA was used as the loading control. (c–h) Quantitative RT-PCR was performed to detect expression of ABF2, RAB18, RD29B, DREB2A, RD22 and MYC2 in wild-type and abo5 seedlings treated with or without ABA for different times. The extracted total RNAs were reverse-transcribed and used for qRT-PCR. Three independent experiments were performed, each with triple replicates. Similar results were obtained. Figure 8. The phenotype of the abo5 mutant cultured under a 12/12 photoperiod. (a) and (b) The wild type and the abo5 mutant phenotype grown on MS medium for 10 days under a 24/0 or 12/12 photoperiod. Relative PGG was calculated based on PGG on MS medium with or without 0.1 lM ABA, and under a 12/12 photoperiod. More than 40 seeds were counted each time. Values are means SEs (n = 3). (c) Relative root growth for wild-type and abo5 mutant seedlings cultured under a 12/12 photoperiod with different concentrations of ABA. Values are means SEs (n = 35). (d) Proline content in abo5 and the wild type. Two-week-old seedlings were grown under a 24/0 or 12/12 photoperiod on MS medium containing 0.1 lM or no ABA. Proline was extracted and quantified. Values are means SE (n = 3). nad1, nad4, nad5, nad6 and nad7 under different photoperiods. Consistent with the northern blot results, the transcript levels of the five genes were greater in abo5 than in the wild type under a 24/0 photoperiod. Under a 12/12 photoperiod, however, transcriptional levels of these genes were similar for abo5 and the wild type (Figure 9a–e) because gene expression levels were much greater under a 12/12 ª 2010 The Authors Journal compilation ª 2010 Blackwell Publishing Ltd, The Plant Journal, (2010), 63, 749–765 ABA and PPR protein 759 (a) (b) (e) (f) (h) (c) (d) (g) (i) (j) Figure 9. Gene expression patterns in abo5 and the wild type under a 12/12 photoperiod. (a–e) Quantitative RT-PCR analysis for nad1 (a), nad4 (b), nad5 (c), nad6 (d) or nad7 (e) gene expression under a 24/0 or a 12/12 photoperiod. Values are means SEs (n = 3). Seedlings were grown on MS medium for 2 weeks. (f) Quantitative RT-PCR for nad2 splicing under a 24/0 or a 12/12 photoperiod. Values are means SE (n = 3). The same sample was used as in (a). (g–h) Quantitative RT-PCR for AOX1a (g) and OXI1 (h) expression under a 24/0 or a 12/12 photoperiod. Values are means SEs (n = 3). Two-week-old seedlings were treated with 50 lM ABA for 24 h. Controls were seedlings that were not treated with ABA. (g–j) Quantitative RT-PCR for PGR5 (i) and PGRL1A (j) expression under a 24/0 or a 12/12 photoperiod. Values are means SE (n = 3). Seedlings were grown on MS medium for 2 weeks. than under a 24/0 photoperiod for the wild type, but remained unchanged for abo5 (Figure 9a–e). For the nad2 gene, we measured the transcripts of different exons covering intron 1, 2, 3 or 4 (Figure 9f). The transcripts covering introns 1 and 2 were higher in abo5 than in the wild type under a 24/0 photoperiod, but were similar under a 12/12 photoperiod. Because abo5 impaired the intron-3 splicing, transcripts covering intron 3 were not detected in abo5 but were detected in the wild type. Because of the impairment of intron-3 splicing, the transcripts covering intron 4 were unaffected by photoperiod in abo5. Transcripts covering introns 1, 2, 3 and 4 were higher under a 12/12 photoperiod than under a 24/0 photoperiod in the wild type, which is consistent with the expression patterns of other nad genes in complex I. We also compared the transcripts of AOX1a and OXI1 in seedlings treated or not treated with 50 lM ABA for 24 h under a 12/12 or 24/0 photoperiod; we found that their expression levels were lower under a 12/12 than under a 24/0 photoperiod in both the wild type and the mutant (Figure 9g,h), and that ABA treatment increased the expression of AOX1a and OXI1 in abo5 and the wild type under both photoperiods. However, expression differences between abo5 and the wild type under ABA treatment occurred under the 24/0 photoperiod but not under the 12/12 photoperiod. There was no clear difference in OXI1 expression without ABA treatment (Figure 9h). AOX1a expression was a little higher in abo5 than in the wild type under the 24/0 photoperiod (Figure 9g), which is consistent with the northern blot result (Figure 5b). Expression of nuclear genes in photosynthesis is changed under constant light conditions in the abo5 mutant Chloroplasts and mitochondria engage in close crosstalk during photosynthetic respiration. Previous studies indicate that impairment of mitochondrion function will reduce the photosynthetic performance in chloroplasts (Noctor et al., 2007). Because abo5 exhibited strong retarded-growth phenotypes under constant light (a 24/0 photoperiod) but not under short days (a 12/12 photoperiod), we suspected that the abo5 mutation might inhibit photosynthesis by influencing the expression of photosynthetic genes. We compared the expression of two nuclear photosynthetic genes in abo5 and the wild type. Proton gradient regulation 5 (PGR5) encodes a thylakoid membrane protein important in the ferredoxin-dependent cyclic electron flow ª 2010 The Authors Journal compilation ª 2010 Blackwell Publishing Ltd, The Plant Journal, (2010), 63, 749–765 760 Yue Liu et al. (CEF) around photosystem I (Munekage et al., 2002). PGR5like photosynthetic phenotype (PGRL1), another thylakoid membrane protein, interacts with PGR5 and facilitates CEF (DalCorso et al., 2008). The expression of PGR5 and PGRL1A was higher in the wild type than in abo5 under a 24/0 photoperiod (Figure 9i), but was similar under a 12/12 photoperiod (Figure 9j). It is conceivable that reduced expression of PGR5 and PGRL1A under a 24/0 photoperiod might reduce CEF, and in turn reduce photosynthesis in abo5. DISCUSSION In this study, we isolated an ABA-sensitive mutant that showed enhanced sensitivity to ABA in both post-germination seedling growth and root growth. The ABO5 gene encodes a PPR protein that is required for the splicing of nad2 intron 3 in mitochondria. nad2 encodes NADH dehydrogenase subunit 2, one of nine mitochondrial geneencoded proteins in complex I. The huge PPR protein family is considered to have expanded before the evolutionary divergence of monocots and dicots (O’Toole et al., 2008). In Arabidopsis and rice, the sizes of the PPR families are similar (450 members in Arabidopsis and 477 in rice) (O’Toole et al., 2008), suggesting that these proteins might have conserved biological roles in the monocot and dicot plants. As of now, however, the biological roles of only a small number of PPR proteins have been characterized. In plants, because of the presence of alternative NAD(P)H dehydrogenases, the electron transport chain can bypass complex I when complex I is not efficiently working or is under stress. In Arabidopsis, six nuclear mutants involved in complex I have been described, including abo5 (this study), fro1/ndufs4 (an 18-kDs subunit of complex I) (Lee et al., 2002; Meyer et al., 2009), a T-DNA insertion mutant in At1g47260 (encoding a carbonic anhydrase) (Perales et al., 2005), opt43 (de Longevialle et al., 2007), css1 (changed sensitivity to cellulose synthesis inhibitors 1, AtnMat1a/ At1g30010, encoding a maturase) (Nakagawa and Sakurai, 2006) and AtnMat2/At5g46920 (Keren et al., 2009). The PPR protein OPT43 is responsible for the trans-splicing of nad1 intron 1, AtnMat1a is responsible for the splicing of mitochondrial nda4, and probably also other mitochondrial genes, and AtnMat2 is responsible for the splicing of cox2, nad1 intron 2 and nad7 intron 2 (de Longevialle et al., 2007; Keren et al., 2009; Nakagawa and Sakurai, 2006). The plant size of the At1g47260 T-DNA mutant is normal, i.e. similar to that of the wild type. The css1 mutant grows slower that the wild type, but finally reaches the same size as the wild-type plant. abo5, fro1/ndufs4, opt43 and AtnMat2 mutants are smaller, and have delayed germination and retarded growth. The inhibition of abo5 germination by high sugar concentration was also observed in the fro1/ndufs4, and css1 mutants (Lee et al., 2002; Meyer et al., 2009; Nakagawa and Sakurai, 2006). Previous studies indicate that sugar has a strong connection with ABA regarding seed germination. Screening for sugar-insensitive (sis) or glucose-insensitive (gin) mutants identified several ABA-insensitive or biosynthesis-deficient mutants, including aba2, abi4 and abi5 (Arenas-Huertero et al., 2000; Laby et al., 2000). Indeed, besides being sensitive to high sugar concentration, the germination of both abo5 and ndufs4 was sensitive to ABA treatment. Furthermore, the seed germination sensitivity to ABA in abo5 could be reversed by each of five classic abi mutations, including two dominant negative mutations in ABI1 and ABI2, and the mutations in ABI3, ABI4 and ABI5. Previous study in analyzing the ABA-sensitive mutant era1 combined with abi1-1 or abi2-1 mutants indicates that the seed germination of abi1 1era1 and abi2 1era1 double mutants is sensitive to ABA (Brady et al., 2003). These results indicate that ABO5 and ERA1 are at different genetic positions with respect to their response to ABA. In the ppr40 mutant that impairs the function of complex III, seed germination is also sensitive to ABA. These results suggest that the energy-requirement processes including both complex I and complex III play important roles in regulation of seed germination by ABA (Meyer et al., 2009; Zsigmond et al., 2008). It is interesting that abo5 was isolated during a genetic screen for mutants in which the roots were sensitive to ABA. ABI1 and ABI2 are two negative regulators in the ABA signaling pathway that have various roles in regulating guard cell movement, plant growth and development, whereas the functions of ABI3, ABI4 and ABI5 have mainly been studied relative to seed germination, but also relative to early seedling development (Bossi et al., 2009; Brady et al., 2003; Lopez-Molina et al., 2001). These results suggest that seed germination and seedling growth respond differently to ABA, and that ABO5 and other ABA-related genes may play different roles in these biological processes. The abo5 mutation increased the expression of nad genes in complex I and of the cox2 gene in complex IV relative to the wild type. However, in the CMSII mutant (lacking a functional mitochondrial complex I) and in the css1 mutant (that impairs the splicing of nad4), the expression of cox2 and/or nad genes was not enhanced (Lelandais et al., 1998; Nakagawa and Sakurai, 2006). These results suggest that unlike the CMSII mutant, which lacks only nad7, the abo5 mutation might not only impair the splicing of nad2 intron 3 but might also play other unidentified biological roles, most probably in mitochondrion. Although the nad genes in complex I and the cox2 gene in complex IV of the mitochondrial genome were expressed at higher levels in abo5 than in the wild type under 24 h of light, the expression of these genes was not increased by a short treatment with ABA. Interestingly, prolonged ABA treatment was able to induce the expression of cox2 and nad genes to higher levels in the wild type, but only to moderate levels in abo5, probably because cox2 and nad genes were already highly expressed as a result of the abo5 mutation. ª 2010 The Authors Journal compilation ª 2010 Blackwell Publishing Ltd, The Plant Journal, (2010), 63, 749–765 ABA and PPR protein 761 These results suggest that the ABA-sensitive phenotype of abo5 might result from the direct impact of ABA on gene expression in the mitochondrion. However, ABO5 expression was not influenced by ABA treatment (5 h in Figure 3a; 7 days, data not shown). The results suggest that ABA affects the expression of cox2 and nad genes in abo5 probably through post-transcription regulation. It is possible that impairment of complex I and/or other biological functions in mitochondria would also impose severe stresses on other cellular components such as chloroplasts, peroxisomes, the cytosol and the nucleus. We found that the expression of some stress-inducible nuclear genes was less induced in abo5 than in the wild type by ABA treatment in a 24/0 photoperiod. Previous study indicates that the coldstress induction of several cold-induced genes is reduced by the fro1 mutation relative to the wild type (Lee et al., 2002). Because lesions of both FRO1/NDUFS4 and ABO5 lead to the accumulation of ROS, it is possible that ROS act as signals to change gene expression levels (Lee et al., 2002; Meyer et al., 2009). Because both ABI1 and ABI2 proteins are sensitive to ROS, the accumulated ROS would inhibit the activities of the negative regulators ABI1 and ABI2 (Meinhard and Grill, 2001; Meinhard et al., 2002). We observed a higher accumulation of ROS in the root tips of abo5 than in the root tips of the wild type, especially in ABA treatment under constant-light conditions. It is therefore possible that the root growth of the ABA-sensitive phenotype might partially result from an increase in ROS accumulation in abo5 relative to the wild type. Consistently, the disruption of two ROS production genes, NADPH oxidase catalytic subunits AtrbohD and AtrbohF in Arabidopsis, impairs the ABA inhibition of seed germination and root growth (Kwak et al., 2003). Disturbance of electron flow in the respiratory chain usually leads to cellular redox imbalance, with a resulting overproduction of ROS. AOX is considered to be important for minimizing ROS generation. Induction of AOX is also closely linked to ROS overproduction in mitochondria. In abo5, more AOX1a was expressed under a 24/0 than under a 12/12 photoperiod, which is consistent with enhanced redox stress caused by a defect in complex I. Previous studies indicate that AOX is necessary for dissipating redox equivalents from chloroplasts (Noctor et al., 2007). Induction of AOX1a in abo5 could help prevent the photoinhibition caused by the overproduction of redox equivalents in chloroplasts. Mutants with a defect in complex I, like opt43, CMSII and maize non-chromosomal stripe (NCS) mutants, have high rates of AOX transcription and translation (de Longevialle et al., 2007; Dutilleul et al., 2003b; Karpova et al., 2002; Vidal et al., 2007). Interestingly, a recent study suggests that the expression of AOX1a is directly regulated by ABI4 (Giraud et al., 2009). However, unlike the CMSII mutant, which does not accumulate more H2O2 than the wild type, the abo5 mutant, like fro1/ndufs4, accumulated slightly more H2O2 than the wild type (Lee et al., 2002; Meyer et al., 2009). Consistent with a slightly higher accumulation of H2O2 in abo5, OXI1, which is another important oxidative burst-mediated gene, was also expressed at a slightly higher level in abo5 than in the wild type under a 24/0 photoperiod. OXI1 is an important mediator for activating two important mitogen-activated protein kinases, MPK3 and MPK6, for ROS-mediated root hair growth (Rentel et al., 2004). The expression of OXI1 is induced by various stresses that stimulate the production of H2O2 (Rentel et al., 2004). Our study suggests that the abo5 mutation, probably partially through impairing complex I as a result of a splicing defect in nad2 intron 3, leads to various stresses at both molecular and growth phenotypic levels, and that the severity of these stresses largely depends on light. A reduced light period alleviates all these different phenotypes, including retarded plant growth and expression of various genes. Complex I in the mitochondrion is the major entry point for the electron transport chain, which burns off the excess reducing NAD(P)H that is produced by photorespiration (from glycine oxidation or by respiration carbon flow), or that is exported directly from chloroplasts during photosynthesis (Noctor et al., 2007). This defect of complex I would alleviate the efficiency of NADH oxidation, which would lead to the accumulated reduced equivalents, which in turn would reduce the photosynthetic activity by regulating the expression of nuclear genes through the retrograde signal pathway. The reduced expression of two important genes, PGR5 and PGRL1A, in photosynthesis supports this hypothesis. Previous studies with the Nicotiana sylvestris CMSII mutant suggest that plants with a loss of function of complex I caused by a mutation of nad7 are not lethal, but decrease the ratio of photosynthesis to respiration with a lower glycine oxidation in a light strength-dependent manner (Noctor et al., 2007; Priault et al., 2006; Sabar et al., 2000). Because mitochondria are closely related to chloroplasts, in that both organelles can convert glycine to serine and provide ATP for cytosol sucrose synthesis during photorespiration, and as required for cellular redox balance, it is reasonable that the mutants defective in complex I or other mitochondrial mutants will greatly affect photosynthetic activities, just as CMSII and the abo5 mutant do (Noctor et al., 2007, 2004; Pogson et al., 2008; Priault et al., 2006; Sabar et al., 2000). However, a biochemical analysis is needed to determine the degree to which the abo5 mutation changes complex I activity. EXPERIMENTAL PROCEDURES Plant growth conditions and mutant isolation Arabidopsis thaliana (Columbia accession) seedlings were grown in forest soil and vermiculite (1:1) under long-day conditions (16-h light/8-h dark photoperiod) at 22C in a glasshouse. Seedlings in plates were grown on MS medium (M5519; Sigma-Aldrich, http:// www.sigmaaldrich.com) containing 3% (w/v) sucrose and 0.8% ª 2010 The Authors Journal compilation ª 2010 Blackwell Publishing Ltd, The Plant Journal, (2010), 63, 749–765 762 Yue Liu et al. (w/v) agar with a 24/0 or a 12/12 photoperiod in a plant growth chamber at 22C. A T-DNA insertion Arabidopsis pool was screened for root bending as described previously (Yin et al., 2009). Seedlings grown on MS medium containing 1% agar for 4 days were transferred onto the medium containing 1% agar and 30 lM ABA. abo5 was selected as an ABA-sensitive mutant. Germination and root growth assay Seeds (>40) were placed on MS medium containing different concentrations of ABA, sucrose and glucose, with three plates for each treatment. Samples were kept at 4C for 2 days and then moved to a plant growth chamber with different photoperiods (24/0 or 12/12). Seedlings with green cotyledons were counted after 2 days, and data were collected every 12 or 24 h. The relative level of PGG (PGG in the presence of ABA expressed as a percentage of PGG in the absence of ABA) was calculated. This experiment was independently repeated three times. Five-day-old seedlings grown on MS medium were transferred onto plates containing different concentrations of ABA under a 24/0 or a 12/12 photoperiod. Root growth was measured after 7 days. Relative root growth (root length in the presence of ABA expressed as a percentage of root length in the absence of ABA) was calculated. This experiment was independently repeated three times. TAIL-PCR for ABO5 cloning The T-DNA flank sequence in abo5 was determined by thermal asymmetric interlaced PCR with pSKI015-specific primers on the left border and a random primer. Primers were as follows: AtLB1, ATACGACGGATCGTAATTTGTC; AtLB2, TAATAACGCTGCGGACATCTAC; AtLB3, TTGACCATCATACTCATTGCTG; random primer DEG1, WGCNAGTNAGWANAAG (W = A or T; N = A, C, G or T). The reaction program for each round was previously described by Qin et al. (2003). Map-based cloning of ABO5 abo5 was crossed with the Landsberg accession, and 1192 abo5 mutants were picked out from the F2 population for their root growth sensitivity on MS medium containing 30 lM ABA. We used simple sequence length polymorphism markers to narrow the mutated site between F5D21 (forward:, 5¢-CTTCTGGATTCTTCCGTTTGCT-3¢; reverse, 5¢-ACCAAAGATTCACGAACTTCGTATC-3¢) and F6D8 (forward, 5¢-GTAGAAGCTGAACGAACCCAAACA-3¢; reverse, 5¢-AGAGTAATTTAGGAAGGACTTCGACAC-3¢), and further mapped using markers T14L22 (forward, 5¢-CCTAACGTACCATAATGATACACCAA-3¢; reverse, 5¢-CCCATTCATCAGCTCAAGGAT-3¢), F19K6 (forward, 5¢-GTCGGATCGATGGCCTAACAAGTGT-3¢; reverse, 5¢-CCGTCTTTGCCGAGATATACTTGGATC-3¢) and F5F19 (forward, 5¢-CCTAAATCGGAAACTGAGTCGACGAC-3¢; reverse, 5¢-GATTGGGCCAAGCCCATAACAC-3¢). The abo5 mutation was located at the top of BAC F5F19. RNA gel blot analyses Seedlings grown on MS medium under a 24/0 or a 12/12 photoperiod were transferred into double-distilled water containing 0 or 50 lM ABA for different times. For prolonged ABA treatment, 7-dayold seedlings were transferred to MS medium with or without 10 lM ABA for 7 days. The photoperiod during incubation in water was the same as for incubation on MS medium. After total RNAs were extracted, 20 lg of RNA from each treatment was separated on a 1.2% (w/v) agarose formaldehyde gel and then hybridized as previously described (Yin et al., 2009). The probes used for RNA gel blot are listed in Table S1. H2O2 assays For the DCFH-DA staining assay, 1-week-old seedlings were treated with 0 or 50 lM ABA. After 5 h, their root tips were removed and incubated in darkness for 10 min in a buffer containing 20 mM K2PO4 K-phosphate (pH 6.0) and 50 lM DCFH-DA. They were then washed three times with 20 mM K-phosphate buffer (pH 6.0) to remove the excess DCFH-DA. Fluorescence was detected with a confocal microscope (Nikon, http://www.nikon. com) with excitation at 488 nm and emission at 525 nm. This experiment was independently repeated three times. Quantitative data were collected for a 1-cm length per root tip using METAMORPH 6.1 software. For quantitative measurement of H2O2, root tips were treated as described above. H2O2 was extracted as described by Rao et al. (2000), and H2O2 content was measured by the Amplex Red Hydrogen Peroxide/Peroxidase Assay kit (Invitrogen, http://www. invitrogen.com). Two independent experiments were performed, each with triple repeats. Similar results were obtained. GUS staining assay The promoter of ABO5 was amplified with forward primer 5¢-PstI (CCACTGACCATCCTTGGTTTGA)-3¢ and reverse primer 5¢-EcoRI (TTTGCGGCGGAGAAGGATACG)-3¢. PCR products were digested and constructed into the pCAMBIA1391 vector. pABO5:GUS was transformed into Arabidopsis by floral dip. Histochemical staining was performed as described previously by Zhou et al. (2009). Twenty-six T2 transgenic lines were subjected to the GUS staining assay. qRT-PCR Total RNA was extracted with TRizol reagent and digested with DNase I. A 6-lg quantity of total RNA was transcripted by M-MLV reverse transcriptase (Promega, http://www.promega.com). qRTPCR was performed as previously described by Zhou et al. (2009). ACTIN4 was used as the internal control to normalize the samples. Each experiment was independently repeated three times, and the primers used for qRT-PCR are listed in Table S2. The primers used to amplify AOX1a, OXI1, cox2 and nads were the same as those used to amplify fragments for the northern blot. Determination of proline, sugar and ABA content Two-week-old seedlings grown on medium containing 0 or 0.1 lM ABA under a 12/12 or a 24/0 photoperiod were ground in liquid nitrogen, and free proline contents were determined according to Chen et al. (2006). Soluble sugar and sucrose contents were determined as previously described (Zhang et al., 2008), and ABA content was determined as described by Chen et al., 2006). These experiments were independently repeated three times. nad2 splicing analysis nad2 premature transcript splicing was analyzed by qRT-PCR. Total RNA was isolated, and its reverse transcription was amplified by qRT-PCR. If introns between the two exons were too long to be amplified, detected PCR products stand for mature transcripts that have been spliced already. Primers used for each fragment were as follows: fragment 1, forward, 5¢-CGTTCGGATCCTCCCACACAT-3¢; reverse, 5¢-GAGCATACCGCGAGTAGGAAGT-3¢; fragment 2, forward, 5¢-CGCTGGCGCACCTCTCCTAACT-3¢; reverse, 5¢-CCACTAGATCGAGCACCAGTGAT-3¢; fragment 3, forward, 5¢-GGGTCTAC TGGAGCTACCCACT-3¢; reverse, 5¢-CTAGAGCGCCCAAATCCGC-3¢; fragment 4, forward, 5¢-GACGATGGATGCATTCGCCATAGT-3¢; reverse, 5¢-CTGAGTGCCATTTGATGAGTAACTGAG-3¢. PCR was ª 2010 The Authors Journal compilation ª 2010 Blackwell Publishing Ltd, The Plant Journal, (2010), 63, 749–765 ABA and PPR protein 763 performed as follows: one cycle at 95C for 5 min followed by 40 cycles of 95C for 20 s, 57C for 20 s and 72C for 20 s. This experiment was independently repeated three times. Subcellular localization A 489-bp fragment at the N terminal of ABO5 was amplified and fused with GFP in frame in front of the 35S promoter. Primers used for amplifying the ABO5 N-terminal fragment were: 5¢-ATGAAGCTTCTCCGCCGCC-3¢; 5¢-CACCATGGAATCGAGAATCGAACGA-3¢. Plasmids were extracted and introduced into protoplasts prepared from Arabidopsis leaves following the protocol previously described by Jin et al. (2001). Mitochondria were stained by Mitotracker (Invitrogen) for co-localization. GFP fluorescence was detected with a confocal microscope, with excitation at 488 nm and emission at 525 nm; for the mito-tracker stain, fluorescence was detected with excitation at 543 nm and emission at 615 nm. Isolation of double mutants of abo5 with each of five classic abi mutants abo5 was crossed with abi1-1, abi2-1, abi3-1, abi4-1 and abi5. The F2 plants were used to identify double mutants. Primers used for confirming the mutation in abi1-1, abi2-1, abi3-1, abi4-1 and abi5 were as follows: for ABI1, forward, 5¢-GATATCTCCGCCGGAGAT-3¢, reverse, 5¢-CCATTCCACTGMTCACTTT-3¢; for ABI2, forward, 5¢-CATCATCTGCTATGGCAGG-3¢, reverse, 5¢-CCGGAGCATGAGCCACAG-3¢; for ABI3, forward, 5¢-CGGTTTCTCTTGCAGAAAGTCTTGAAGCAAGTC-3¢, reverse, 5¢-TTGCCTCTAGCTCCGGCAAGT-3¢; for ABI4, forward, 5¢-ATGGACCCTTTAGCTTCCCAACATC-3¢, reverse, 5¢-AGTTACCGGAACATCAGTGAGCTCG-3¢; for ABI5, forward, 5¢-AGCTGAACAGGGACAAGTAACTGAAGTTTG-3¢, reverse, 5¢-CTCTGACGTCAACTTCGTTTCTCTAGTTACCATTTAT-3¢. PCR products of abi1-1, abi2-1, abi3-1, abi4-1 and abi5 were digested by NcoI, NcoI, SalI, NIaIV and MseI, respectively. abo5 was checked with the same ABO5-F and ABO5-R primers used for qRT-PCR (see Tables S1 and S2). Accession number Sequence data from this article can be found in the GenBank/EMBL data libraries under accession number NM_148578 (At1g51965). ACKNOWLEDGEMENTS We thank Dr Li-jia Qu of Peking University for providing the T-DNA mutant pool, the Arabidopsis Biological Resource Center for providing the T-DNA insertion lines of ABO5 and Dr Harvey Millar of The University of Western Australia for providing ndufs4 mutant seeds. This work was supported by the National Nature Science Foundation of China (90717004, 30721062), the National Transgenic Research Project (2008ZX08009-002) and the Programme of Introducing Talents of Discipline to Universities (B06003) to ZG. SUPPORTING INFORMATION Additional Supporting Information may be found in the online version of this article: Table S1. Primers used to amplify fragments in the northern blot. Table S2. Primers used for quantitative RT-PCR. Please note: As a service to our authors and readers, this journal provides supporting information supplied by the authors. Such materials are peer-reviewed and may be re-organized for online delivery, but are not copy-edited or typeset. Technical support issues arising from supporting information (other than missing files) should be addressed to the authors. REFERENCES Abe, H., Urao, T., Ito, T., Seki, M., Shinozaki, K. and Yamaguchi-Shinozaki, K. (2003) Arabidopsis AtMYC2 (bHLH) and AtMYB2 (MYB) function as transcriptional activators in abscisic acid signaling. Plant Cell, 15, 63–78. Arenas-Huertero, F., Arroyo, A., Zhou, L., Sheen, J. and Leon, P. (2000) Analysis of Arabidopsis glucose insensitive mutants, gin5 and gin6, reveals a central role of the plant hormone ABA in the regulation of plant vegetative development by sugar. Genes Dev. 14, 2085–2096. Beick, S., Schmitz-Linneweber, C., Williams-Carrier, R., Jensen, B. and Barkan, A. (2008) The pentatricopeptide repeat protein PPR5 stabilizes a specific tRNA precursor in maize chloroplasts. Mol. Cell. Biol. 28, 5337–5347. Bossi, F., Cordoba, E., Dupre, P., Mendoza, M.S., Roman, C.S. and Leon, P. (2009) The Arabidopsis ABA-INSENSITIVE (ABI) 4 factor acts as a central transcription activator of the expression of its own gene, and for the induction of ABI5 and SBE2.2 genes during sugar signaling. Plant J. 59, 359–374. Brady, S.M., Sarkar, S.F., Bonetta, D. and McCourt, P. (2003) The ABSCISIC ACID INSENSITIVE 3 (ABI3) gene is modulated by farnesylation and is involved in auxin signaling and lateral root development in Arabidopsis. Plant J. 34, 67–75. Cai, W., Ji, D., Peng, L., Guo, J., Ma, J., Zou, M., Lu, C. and Zhang, L. (2009) LPA66 is required for editing psbF chloroplast transcripts in Arabidopsis. Plant Physiol. 150, 1260–1271. Chen, Z., Zhang, H., Jablonowski, D., Zhou, X., Ren, X., Hong, X., Schaffrath, R., Zhu, J.K. and Gong, Z. (2006) Mutations in ABO1/ELO2, a subunit of holo-Elongator, increase abscisic acid sensitivity and drought tolerance in Arabidopsis thaliana. Mol. Cell. Biol. 26, 6902–6912. Choi, H., Hong, J., Ha, J., Kang, J. and Kim, S.Y. (2000) ABFs, a family of ABAresponsive element binding factors. J. Biol. Chem. 275, 1723–1730. Cushing, D.A., Forsthoefel, N.R., Gestaut, D.R. and Vernon, D.M. (2005) Arabidopsis emb175 and other ppr knockout mutants reveal essential roles for pentatricopeptide repeat (PPR) proteins in plant embryogenesis. Planta, 221, 424–436. DalCorso, G., Pesaresi, P., Masiero, S., Aseeva, E., Schunemann, D., Finazzi, G., Joliot, P., Barbato, R. and Leister, D. (2008) A complex containing PGRL1 and PGR5 is involved in the switch between linear and cyclic electron flow in Arabidopsis. Cell, 132, 273–285. Dutilleul, C., Driscoll, S., Cornic, G., De Paepe, R., Foyer, C.H. and Noctor, G. (2003a) Functional mitochondrial complex I is required by tobacco leaves for optimal photosynthetic performance in photorespiratory conditions and during transients. Plant Physiol. 131, 264–275. Dutilleul, C., Garmier, M., Noctor, G., Mathieu, C., Chetrit, P., Foyer, C.H. and de Paepe, R. (2003b) Leaf mitochondria modulate whole cell redox homeostasis, set antioxidant capacity, and determine stress resistance through altered signaling and diurnal regulation. Plant Cell, 15, 1212– 1226. Finkelstein, R.R. and Lynch, T.J. (2000) The Arabidopsis abscisic acid response gene ABI5 encodes a basic leucine zipper transcription factor. Plant Cell, 12, 599–609. Finkelstein, R.R., Wang, M.L., Lynch, T.J., Rao, S. and Goodman, H.M. (1998) The Arabidopsis abscisic acid response locus ABI4 encodes an APETALA 2 domain protein. Plant Cell, 10, 1043–1054. Fujii, H., Chinnusamy, V., Rodrigues, A., Rubio, S., Antoni, R., Park, S.Y., Cutler, S.R., Sheen, J., Rodriguez, P.L. and Zhu, J.K. (2009) In vitro reconstitution of an abscisic acid signalling pathway. Nature, 462, 660–664. Fujita, Y., Fujita, M., Satoh, R., Maruyama, K., Parvez, M.M., Seki, M., Hiratsu, K., Ohme-Takagi, M., Shinozaki, K. and Yamaguchi-Shinozaki, K. (2005) AREB1 Is a transcription activator of novel ABRE-dependent ABA signaling that enhances drought stress tolerance in Arabidopsis. Plant Cell, 17, 3470– 3488. Giraud, E., Van Aken, O., Ho, L.H. and Whelan, J. (2009) The transcription factor ABI4 is a regulator of mitochondrial retrograde expression of ALTERNATIVE OXIDASE1a. Plant Physiol. 150, 1286–1296. Giraudat, J., Hauge, B.M., Valon, C., Smalle, J., Parcy, F. and Goodman, H.M. (1992) Isolation of the Arabidopsis ABI3 gene by positional cloning. Plant Cell, 4, 1251–1261. Heazlewood, J.L., Howell, K.A. and Millar, A.H. (2003) Mitochondrial complex I from Arabidopsis and rice: orthologs of mammalian and fungal components coupled with plant-specific subunits. Biochim. Biophys. Acta, 1604, 159–169. ª 2010 The Authors Journal compilation ª 2010 Blackwell Publishing Ltd, The Plant Journal, (2010), 63, 749–765 764 Yue Liu et al. Jiao, S., Thornsberry, J.M., Elthon, T.E. and Newton, K.J. (2005) Biochemical and molecular characterization of photosystem I deficiency in the NCS6 mitochondrial mutant of maize. Plant Mol. Biol. 57, 303–313. Jin, J.B., Kim, Y.A., Kim, S.J., Lee, S.H., Kim, D.H., Cheong, G.W. and Hwang, I. (2001) A new dynamin-like protein, ADL6, is involved in trafficking from the trans-Golgi network to the central vacuole in Arabidopsis. Plant Cell, 13, 1511–1526. Karpova, O.V., Kuzmin, E.V., Elthon, T.E. and Newton, K.J. (2002) Differential expression of alternative oxidase genes in maize mitochondrial mutants. Plant Cell, 14, 3271–3284. Keren, I., Bezawork-Geleta, A., Kolton, M., Maayan, I., Belausov, E., Levy, M., Mett, A., Gidoni, D., Shaya, F. and Ostersetzer-Biran, O. (2009) AtnMat2, a nuclear-encoded maturase required for splicing of group-II introns in Arabidopsis mitochondria. RNA, 15, 2299–2311. Kim, S., Kang, J.Y., Cho, D.I., Park, J.H. and Kim, S.Y. (2004) ABF2, an ABRE-binding bZIP factor, is an essential component of glucose signaling and its overexpression affects multiple stress tolerance. Plant J. 40, 75–87. Kim, S.R., Yang, J.I., Moon, S., Ryu, C.H., An, K., Kim, K.M., Yim, J. and An, G. (2009) Rice OGR1 encodes a pentatricopeptide repeat-DYW protein and is essential for RNA editing in mitochondria. Plant J. 59, 738–749. Kotera, E., Tasaka, M. and Shikanai, T. (2005) A pentatricopeptide repeat protein is essential for RNA editing in chloroplasts. Nature, 433, 326– 330. Kwak, J.M., Mori, I.C., Pei, Z.M., Leonhardt, N., Torres, M.A., Dangl, J.L., Bloom, R.E., Bodde, S., Jones, J.D. and Schroeder, J.I. (2003) NADPH oxidase AtrbohD and AtrbohF genes function in ROS-dependent ABA signaling in Arabidopsis. EMBO J. 22, 2623–2633. Laby, R.J., Kincaid, M.S., Kim, D. and Gibson, S.I. (2000) The Arabidopsis sugar-insensitive mutants sis4 and sis5 are defective in abscisic acid synthesis and response. Plant J. 23, 587–596. Lang, V. and Palva, E.T. (1992) The expression of a rab-related gene, rab18, is induced by abscisic acid during the cold acclimation process of Arabidopsis thaliana (L.) Heynh. Plant Mol. Biol. 20, 951–962. Lee, B.H., Lee, H., Xiong, L. and Zhu, J.K. (2002) A mitochondrial complex I defect impairs cold-regulated nuclear gene expression. Plant Cell, 14, 1235– 1251. Lelandais, C., Albert, B., Gutierres, S., De Paepe, R., Godelle, B., Vedel, F. and Chetrit, P. (1998) Organization and expression of the mitochondrial genome in the Nicotiana sylvestris CMSII mutant. Genetics, 150, 873–882. Leung, J., Merlot, S. and Giraudat, J. (1997) The Arabidopsis ABSCISIC ACID-INSENSITIVE2 (ABI2) and ABI1 genes encode homologous protein phosphatases 2C involved in abscisic acid signal transduction. Plant Cell, 9, 759–771. Liu, Q., Kasuga, M., Sakuma, Y., Abe, H., Miura, S., Yamaguchi-Shinozaki, K. and Shinozaki, K. (1998) Two transcription factors, DREB1 and DREB2, with an EREBP/AP2 DNA binding domain separate two cellular signal transduction pathways in drought- and low-temperature-responsive gene expression, respectively, in Arabidopsis. Plant Cell, 10, 1391–1406. de Longevialle, A.F., Meyer, E.H., Andres, C., Taylor, N.L., Lurin, C., Millar, A.H. and Small, I.D. (2007) The pentatricopeptide repeat gene OTP43 is required for trans-splicing of the mitochondrial nad1 Intron 1 in Arabidopsis thaliana. Plant Cell, 19, 3256–3265. de Longevialle, A.F., Hendrickson, L., Taylor, N.L., Delannoy, E., Lurin, C., Badger, M., Millar, A.H. and Small, I. (2008) The pentatricopeptide repeat gene OTP51 with two LAGLIDADG motifs is required for the cis-splicing of plastid ycf3 intron 2 in Arabidopsis thaliana. Plant J. 56, 157–168. Lopez-Molina, L., Mongrand, S. and Chua, N.H. (2001) A postgermination developmental arrest checkpoint is mediated by abscisic acid and requires the ABI5 transcription factor in Arabidopsis. Proc. Natl. Acad. Sci. USA, 98, 4782–4787. Lurin, C., Andres, C., Aubourg, S. et al. (2004) Genome-wide analysis of Arabidopsis pentatricopeptide repeat proteins reveals their essential role in organelle biogenesis. Plant Cell, 16, 2089–2103. Ma, Y., Szostkiewicz, I., Korte, A., Moes, D., Yang, Y., Christmann, A. and Grill, E. (2009) Regulators of PP2C phosphatase activity function as abscisic acid sensors. Science, 324, 1064–1068. Matsuo, M. and Obokata, J. (2006) Remote control of photosynthetic genes by the mitochondrial respiratory chain. Plant J. 47, 873–882. Meinhard, M. and Grill, E. (2001) Hydrogen peroxide is a regulator of ABI1, a protein phosphatase 2C from Arabidopsis. FEBS Lett. 508, 443–446. Meinhard, M., Rodriguez, P.L. and Grill, E. (2002) The sensitivity of ABI2 to hydrogen peroxide links the abscisic acid-response regulator to redox signalling. Planta, 214, 775–782. Meyer, K., Leube, M.P. and Grill, E. (1994) A protein phosphatase 2C involved in ABA signal transduction in Arabidopsis thaliana. Science, 264, 1452– 1455. Meyer, E.H., Tomaz, T., Carroll, A.J., Estavillo, G., Delannoy, E., Tanz, S.K., Small, I.D., Pogson, B.J. and Millar, A.H. (2009) Remodeled respiration in ndufs4 with low phosphorylation efficiency suppresses Arabidopsis germination and growth and alters control of metabolism at night. Plant Physiol. 151, 603–619. Miao, Y., Lv, D., Wang, P., Wang, X.C., Chen, J., Miao, C. and Song, C.P. (2006) An Arabidopsis glutathione peroxidase functions as both a redox transducer and a scavenger in abscisic acid and drought stress responses. Plant Cell, 18, 2749–2766. Munekage, Y., Hojo, M., Meurer, J., Endo, T., Tasaka, M. and Shikanai, T. (2002) PGR5 is involved in cyclic electron flow around photosystem I and is essential for photoprotection in Arabidopsis. Cell, 110, 361–371. Murata, Y., Pei, Z.M., Mori, I.C. and Schroeder, J. (2001) Abscisic acid activation of plasma membrane Ca(2+) channels in guard cells requires cytosolic NAD(P)H and is differentially disrupted upstream and downstream of reactive oxygen species production in abi1-1 and abi2-1 protein phosphatase 2C mutants. Plant Cell, 13, 2513–2523. Nakagawa, N. and Sakurai, N. (2006) A mutation in At-nMat1a, which encodes a nuclear gene having high similarity to group II intron maturase, causes impaired splicing of mitochondrial NAD4 transcript and altered carbon metabolism in Arabidopsis thaliana. Plant Cell Physiol. 47, 772–783. Nakashima, K., Fujita, Y., Kanamori, N. et al. (2009) Three Arabidopsis SnRK2 protein kinases, SRK2D/SnRK2.2, SRK2E/SnRK2.6/OST1 and SRK2I/ SnRK2.3, involved in ABA signaling are essential for the control of seed development and dormancy. Plant Cell Physiol. 50, 1345–1363. Nishimura, N., Sarkeshik, A., Nito, K. et al. (2010) PYR/PYL/RCAR family members are major in-vivo ABI1 protein phosphatase 2C-interacting proteins in Arabidopsis. Plant J. 61, 290–299. Nishimura, N., Hitomi, K., Arvai, A.S., Rambo, R.P., Hitomi, C., Cutler, S.R., Schroeder, J.I. and Getzoff, E.D. (2009) Structural mechanism of abscisic acid binding and signaling by dimeric PYR1. Science, 326, 1373–1379. Noctor, G., Dutilleul, C., De Paepe, R. and Foyer, C.H. (2004) Use of mitochondrial electron transport mutants to evaluate the effects of redox state on photosynthesis, stress tolerance and the integration of carbon/nitrogen metabolism. J. Exp. Bot. 55, 49–57. Noctor, G., De Paepe, R. and Foyer, C.H. (2007) Mitochondrial redox biology and homeostasis in plants. Trends Plant Sci. 12, 125–134. Okuda, K., Chateigner-Boutin, A.L., Nakamura, T., Delannoy, E., Sugita, M., Myouga, F., Motohashi, R., Shinozaki, K., Small, I. and Shikanai, T. (2009) Pentatricopeptide repeat proteins with the DYW motif have distinct molecular functions in RNA editing and RNA cleavage in Arabidopsis chloroplasts. Plant Cell, 21, 146–156. O’Toole, N., Hattori, M., Andres, C., Iida, K., Lurin, C., Schmitz-Linneweber, C., Sugita, M. and Small, I. (2008) On the expansion of the pentatricopeptide repeat gene family in plants. Mol. Biol. Evol. 25, 1120–1128. Park, S.Y., Fung, P., Nishimura, N. et al. (2009) Abscisic acid inhibits type 2C protein phosphatases via the PYR/PYL family of START proteins. Science, 324, 1068–1071. Pei, Z.M., Kuchitsu, K., Ward, J.M., Schwarz, M. and Schroeder, J.I. (1997) Differential abscisic acid regulation of guard cell slow anion channels in Arabidopsis wild-type and abi1 and abi2 mutants. Plant Cell, 9, 409–423. Perales, M., Eubel, H., Heinemeyer, J., Colaneri, A., Zabaleta, E. and Braun, H.P. (2005) Disruption of a nuclear gene encoding a mitochondrial gamma carbonic anhydrase reduces complex I and supercomplex I + III2 levels and alters mitochondrial physiology in Arabidopsis. J. Mol. Biol. 350, 263–277. Pfalz, J., Bayraktar, O.A., Prikryl, J. and Barkan, A. (2009) Site-specific binding of a PPR protein defines and stabilizes 5¢ and 3¢ mRNA termini in chloroplasts. EMBO J. 28, 2042–2052. Pogson, B.J., Woo, N.S., Forster, B. and Small, I.D. (2008) Plastid signalling to the nucleus and beyond. Trends Plant Sci. 13, 602–609. Priault, P., Fresneau, C., Noctor, G., De Paepe, R., Cornic, G. and Streb, P. (2006) The mitochondrial CMSII mutation of Nicotiana sylvestris impairs adjustment of photosynthetic carbon assimilation to higher growth irradiance. J. Exp. Bot. 57, 2075–2085. ª 2010 The Authors Journal compilation ª 2010 Blackwell Publishing Ltd, The Plant Journal, (2010), 63, 749–765 ABA and PPR protein 765 Qin, G., Kang, D. and Dong, Y. (2003) Obtaining and analysis of flanking sequences from T-DNA transformants of Arabidopsis. Plant Sci. 165, 941– 949. Rao, M.V., Lee, H., Creelman, R.A., Mullet, J.E. and Davis, K.R. (2000) Jasmonic acid signaling modulates ozone-induced hypersensitive cell death. Plant Cell, 12, 1633–1646. Rentel, M.C., Lecourieux, D., Ouaked, F. et al. (2004) OXI1 kinase is necessary for oxidative burst-mediated signalling in Arabidopsis. Nature, 427, 858– 861. Rhoads, D.M. and Subbaiah, C.C. (2007) Mitochondrial retrograde regulation in plants. Mitochondrion, 7, 177–194. Robbins, J.C., Heller, W.P. and Hanson, M.R. (2009) A comparative genomics approach identifies a PPR-DYW protein that is essential for C-to-U editing of the Arabidopsis chloroplast accD transcript. RNA, 15, 1142–1153. Sabar, M., De Paepe, R. and de Kouchkovsky, Y. (2000) Complex I impairment, respiratory compensations, and photosynthetic decrease in nuclear and mitochondrial male sterile mutants of Nicotiana sylvestris. Plant Physiol. 124, 1239–1250. Santiago, J., Dupeux, F., Round, A., Antoni, R., Park, S.Y., Jamin, M., Cutler, S.R., Rodriguez, P.L. and Marquez, J.A. (2009) The abscisic acid receptor PYR1 in complex with abscisic acid. Nature, 462, 665–668. Schmitz-Linneweber, C. and Small, I. (2008) Pentatricopeptide repeat proteins: a socket set for organelle gene expression. Trends Plant Sci. 13, 663–670. Schmitz-Linneweber, C., Williams-Carrier, R.E., Williams-Voelker, P.M., Kroeger, T.S., Vichas, A. and Barkan, A. (2006) A pentatricopeptide repeat protein facilitates the trans-splicing of the maize chloroplast rps12 premRNA. Plant Cell, 18, 2650–2663. Strodtkotter, I., Padmasree, K., Dinakar, C. et al. (2009) Induction of the AOX1D isoform of alternative oxidase in A. thaliana T-DNA insertion lines lacking isoform AOX1A is insufficient to optimize photosynthesis when treated with antimycin A. Mol. Plant, 2, 284–297. Surpin, M., Larkin, R.M. and Chory, J. (2002) Signal transduction between the chloroplast and the nucleus. Plant Cell, 14(Suppl.), S327–S338. Umbach, A.L., Fiorani, F. and Siedow, J.N. (2005) Characterization of transformed Arabidopsis with altered alternative oxidase levels and analysis of effects on reactive oxygen species in tissue. Plant Physiol. 139, 1806–1820. Uyttewaal, M., Arnal, N., Quadrado, M., Martin-Canadell, A., Vrielynck, N., Hiard, S., Gherbi, H., Bendahmane, A., Budar, F. and Mireau, H. (2008) Characterization of Raphanus sativus pentatricopeptide repeat proteins encoded by the fertility restorer locus for Ogura cytoplasmic male sterility. Plant Cell, 20, 3331–3345. Vidal, G., Ribas-Carbo, M., Garmier, M., Dubertret, G., Rasmusson, A.G., Mathieu, C., Foyer, C.H. and De Paepe, R. (2007) Lack of respiratory chain complex I impairs alternative oxidase engagement and modulates redox signaling during elicitor-induced cell death in tobacco. Plant Cell, 19, 640– 655. Woodson, J.D. and Chory, J. (2008) Coordination of gene expression between organellar and nuclear genomes. Nat. Rev. Genet. 9, 383–395. Yamaguchi-Shinozaki, K. and Shinozaki, K. (1993) The plant hormone abscisic acid mediates the drought-induced expression but not the seed-specific expression of rd22, a gene responsive to dehydration stress in Arabidopsis thaliana. Mol. Gen. Genet. 238, 17–25. Yin, H., Zhang, X., Liu, J., Wang, Y., He, J., Yang, T., Hong, X., Yang, Q. and Gong, Z. (2009) Epigenetic regulation, somatic homologous recombination, and abscisic acid signaling are influenced by DNA polymerase epsilon mutation in Arabidopsis. Plant Cell, 21, 386–402. Yu, Q.B., Jiang, Y., Chong, K. and Yang, Z.N. (2009) AtECB2, a pentatricopeptide repeat protein, is required for chloroplast transcript accD RNA editing and early chloroplast biogenesis in Arabidopsis thaliana. Plant J. 59, 1011–1023. Zehrmann, A., Verbitskiy, D., van der Merwe, J.A., Brennicke, A. and Takenaka, M. (2009) A DYW domain-containing pentatricopeptide repeat protein is required for RNA editing at multiple sites in mitochondria of Arabidopsis thaliana. Plant Cell, 21, 558–567. Zhang, H., Ohyama, K., Boudet, J., Chen, Z., Yang, J., Zhang, M., Muranaka, T., Maurel, C., Zhu, J.K. and Gong, Z. (2008) Dolichol biosynthesis and its effects on the unfolded protein response and abiotic stress resistance in Arabidopsis. Plant Cell, 20, 1879–1898. Zhou, X., Hua, D., Chen, Z., Zhou, Z. and Gong, Z. (2009) Elongator mediates ABA responses, oxidative stress resistance and anthocyanin biosynthesis in Arabidopsis. Plant J. 60, 79–90. Zsigmond, L., Rigo, G., Szarka, A., Szekely, G., Otvos, K., Darula, Z., Medzihradszky, K.F., Koncz, C., Koncz, Z. and Szabados, L. (2008) Arabidopsis PPR40 connects abiotic stress responses to mitochondrial electron transport. Plant Physiol. 146, 1721–1737. ª 2010 The Authors Journal compilation ª 2010 Blackwell Publishing Ltd, The Plant Journal, (2010), 63, 749–765