* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Molecular encounters at microtubule ends in the plant cell cortex
Cell membrane wikipedia , lookup
Cell nucleus wikipedia , lookup
Biochemical switches in the cell cycle wikipedia , lookup
Cell encapsulation wikipedia , lookup
Spindle checkpoint wikipedia , lookup
Cytoplasmic streaming wikipedia , lookup
Signal transduction wikipedia , lookup
Cellular differentiation wikipedia , lookup
Extracellular matrix wikipedia , lookup
Programmed cell death wikipedia , lookup
Cell culture wikipedia , lookup
Endomembrane system wikipedia , lookup
Organ-on-a-chip wikipedia , lookup
Cell growth wikipedia , lookup
Kinetochore wikipedia , lookup
Microtubule wikipedia , lookup
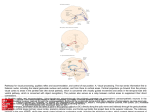
COPLBI-475; NO OF PAGES 7 Molecular encounters at microtubule ends in the plant cell cortex Martine Pastuglia and David Bouchez The cortical arrays that accompany plant cell division and elongation are organized by a subtle interplay between intrinsic properties of microtubules, their self-organization capacity and a variety of cellular proteins that interact with them, modify their behaviour and drive organization of diverse, higher order arrays during the cell cycle, cell growth and differentiation. As a polar polymer, the microtubule has a minus and a plus end, which differ in structure and dynamic characteristics, and to which different sets of partners and activities associate. Recent advances in characterization of minus and plus end directed proteins provide insights into both plant microtubule properties and the way highly organized cortical arrays emerge from the orchestrated activity of individual microtubules. Addresses Institut Jean-Pierre Bourgin, Station de Génétique et d’Amélioration des Plantes UR254, INRA, Centre de Versailles, F-78000 Versailles, France Corresponding author: Pastuglia, Martine ([email protected]) Current Opinion in Plant Biology 2007, 10:1–7 This review comes from a themed issue on Cell Biology Edited by Ben Scheres and Volker Lipka 1369-5266/$ – see front matter # 2007 Elsevier Ltd. All rights reserved. DOI 10.1016/j.pbi.2007.08.001 Introduction Microtubules (MTs) are distinctive components of eukaryotic cells, where they play essential roles in spatially controlled processes like organization of the cytoplasm, cell shape and polarity, flagella/cilia and cell motility, intracellular transport of vesicles and proteins, mitotic and meiotic cell division and cell wall deposition (in plants), to name a few. MTs are non-covalent, polar polymers of a/b tubulin heterodimers assembled headto-tail into linear protofilaments. Side association of (usually) 13 protofilaments forms a hollow cylinder of 25 nm in diameter, a rigid filamentous structure that is intrinsically polar, with a rapidly growing plus end and a slow growing minus end defined as the a-tubulin end (Figure 1a). Properties of tubulin make MTs highly dynamic, with stochastic transitions between growing and shrinking phases, a phenomenon called ‘dynamic instability’. When MT minus ends are anchored or capped, only plus ends undergo dynamic instability, whereas unanchored MTs can display directional repositioning through www.sciencedirect.com ‘treadmilling’ (net polymerization at the plus end, net depolymerization at the minus end); for a recent review on MT assembly, see reference [1]. Thanks to intrinsic properties of MTs and to activities of many cellular proteins that can modify, (de)stabilize, move, cut or bridge them, MTs generate higher order networks that organize the cellular space and adopt different forms during the cell cycle. In expanding plant cells, MTs are cortical, tightly associated with the inner face of the plasma membrane and arranged into parallel arrays transverse to the elongation axis, guiding cellulose deposition on the outside of the cell [2]. At G2/M transition, cortical MTs arrange into a highly condensed ring encircling the nucleus, with radial MTs connecting nucleus and cortex. This transient structure, the preprophase band of MTs (PPB), is specific to land plants and precisely predicts the site of division before disassembling at prophase. To control the distribution and organization of MTs, many eukaryotic cells rely on specific organelles called MT-organizing centres (MTOCs), which are sites for nucleation of new MTs [3]. MTs can spontaneously assemble in vitro under conditions of high concentrations of tubulin, but they form at a much lower tubulin concentration in living cells. MTOCs help overcome the kinetic barrier of limiting tubulin concentration but are also a means for spatial and temporal regulation of MT initiation. In animals, the centrosome consists of a pair of centrioles linked by a matrix and surrounded by a pericentriolar matrix, which promotes de novo nucleation of MTs for the setup of interphase cytoplasmic MT arrays and mitotic spindles. Its capacity to organize MT arrays also depends on tight control of anchoring, capping and release of nucleated MTs. Unlike most other eukaryotes, cells of land plants lack a conspicuous MTOC like a centrosome, except for basal bodies formed in flagellate sperm cells of lower land plants. The absence of centrosomes, centrioles and the mechanisms of MT nucleation in higher plants have prompted many questions as to whether plant cells have conserved, revamped or reinvented functions associated with MTOCs in other eukaryotes. The theory that centrosomes are flexible bodies that adopt a variety of forms was formulated more than 20 years ago [4]. The idea of a distributed centrosome at the membrane’s periphery in plant cells, involved in nucleation, release and attachment of MTs at the cortex, has received wide support in the field, but until recently this had limited direct experimental support. Current Opinion in Plant Biology 2007, 10:1–7 Please cite this article in press as: Pastuglia M, Bouchez D, Molecular encounters at microtubule ends in the plant cell cortex, Curr Opin Plant Biol (2007), doi:10.1016/j.pbi.2007.08.001 COPLBI-475; NO OF PAGES 7 2 Cell Biology Figure 1 Microtubules from nucleation to establishment of cortical arrays. (a) Microtubule assembly and disassembly. MTs are polymers of a/b tubulin heterodimers. A typical MT is composed of 13 linear protofilaments resulting from head-to-tail arrangement of heterodimers. Lateral association of protofilaments forms a hollow cylinder of 25 nm in diameter. The orientation of a/b tubulin dimers in the MT lattice confers a polarity that is crucial for MT organization, with b-tubulin pointing towards the fast growing plus end, and a-tubulin pointing towards the slow growing minus end. a and b-tubulins have N-terminal GTP-binding pockets, but GTP is exchangeable only in b-tubulin in un-incorporated heterodimers. The GTP of the b subunit is buried upon MT polymerization and is rapidly hydrolyzed by a-tubulin catalytic residues into non-exchangeable GDP. The hydrolysis of GTP in b-tubulin is coupled to incorporation in the MT lattice and affects the structure of the a/b heterodimer, which in turn introduces tensions in the protofilament that provide energy for rapid disassembly of the MT. The MT structure is stabilized by a cap of a few layers of GTP-tubulin subunits at the plus end; when the cap is lost, the MT undergoes a rapid depolymerization (termed a catastrophe). Recent models propose that the plus end of a growing MT is an open sheet of protofilaments with lateral contacts, gradually closing into a hollow cylinder [1]. Such a sheet conformation of growing plus ends is expected to provide an open surface for specific interactions at the plus end. Typical curled protofilaments (’protofilament peeling’) are produced during MT shortening. MTs undergo dynamic instability, showing stochastic transitions between growing, pause and shrinking phases. When the minus end is anchored at its nucleation site, only the free plus end exhibits dynamic instability, but free minus ends are also dynamic in non-anchored MTs. This seems to be the rule in the plant cell cortex, where MTs are detached from their nucleation sites and translocated by a ‘hybrid treadmilling’ mechanism [9]. (b) Microtubule nucleation and the g-tubulin complex. The basic process of MT nucleation is probably highly conserved in eukaryotes including plants, though this has to be further confirmed experimentally. In animal cells, g-tubulin is present in a large ring-shaped 2 MDa multi-protein complex, approximately the diameter of a MT (25 nm), called a g-TuRC. The g-TuRC accelerates the rate of MT initiation, and could also have a capping function, inhibiting minus end dynamics. A widely Current Opinion in Plant Biology 2007, 10:1–7 www.sciencedirect.com Please cite this article in press as: Pastuglia M, Bouchez D, Molecular encounters at microtubule ends in the plant cell cortex, Curr Opin Plant Biol (2007), doi:10.1016/j.pbi.2007.08.001 COPLBI-475; NO OF PAGES 7 Molecular encounters at microtubule ends in the plant cell cortex Pastuglia and Bouchez 3 This review is an overview of some progress on the characterization of protein networks at the minus and plus ends of MTs in plant cells. Recent results show that such interactions at MT ends are instrumental in producing organized cortical arrays from concerted activity of individual MTs. g-Tubulin complexes and MT nucleation at the cortex The absence of a well-defined, central MT organizer has long delayed characterization of the molecular basis of MT nucleation in higher plant cells. The third member of the tubulin superfamily, g-tubulin, is a key component of eukaryotic MTOCs where it is found associated to minus ends of MTs in large multi-protein complexes that serve as templates for MT initiation (Figure 1b). Although gtubulin is conserved in plant genomes, its involvement in MT nucleation in acentrosomal plant cells has been controversial until recently. The first evidence of MT nucleation by plant g-tubulin came from heterologous expression of Arabidopsis g-tubulin in fission yeast (Schizosaccharomyces pombe) lacking endogenous g-tubulin. Arabidopsis g-tubulin was properly targeted to MTOCs and able to nucleate MTs in fission yeast [5]. In plant cells, g-tubulin was not restricted to MT ends as expected but seen as a punctate staining along MT walls [6,7]. However, a recent study established that this pattern reflects bona fide MT-dependent nucleation sites [8] and that in the plant cell cortex, MTs are mostly nucleated from the surface of extant MTs, resulting in a branched MT network (Figure 1c). Furthermore, recruitment of cytoplasmic g-tubulin onto existing MTs was necessary to initiate new MTs. Several studies already reported MT nucleation at cortical sites linked to pre-existing MTs in plant cells [9,10], and such branched organization occurs in animal cells [11]. Knocking down both g-tubulin genes in Arabidopsis further supported the involvement of g-tubulin in MT nucleation [12,13]. Regrowth of cortical MTs was particularly impaired in cells with depleted RNAi g-tubulin [12]. In animal cells, g-tubulin is part of large ring-shaped protein complexes called g-tubulin ring complexes (g-TuRCs) (Figure 1b). The g-TuRC contains 10–13 gtubulin molecules per complex and additional subunits named gamma complex proteins (GCPs 2–6) in humans and Dgrip in Drosophila [3] (Table 1). The Arabidopsis genome encodes putative orthologues for all GCPs, including the recently discovered GCP-WD/Nedd1 (Dgp71WD). An antibody directed towards plant Spc98 (GCP3) inhibits MT assembly from isolated tobacco (Nicotiana tabacum) nuclei [14]. However, the absence of co-localization of AtSpc98 and g-tubulin in the cortex raises questions on its role in MT assembly of cortical arrays. Further characterization of the putative plant g-TuRC and functional analysis of these proteins are needed to clarify the processes of MT nucleation in plant cells. The organization of highly structured arrays strongly depends on the spatial and temporal distribution of MT nucleation sites in a cell. Recruitment of nucleation complexes at the surface of extant MTs is not restricted to plant cells. For example, in Drosophila, g-tubulin transiently associates to the spindle and midbody MTs during mitosis, and this interaction requires cap components Dgrip 75, 128, 163 and Dgp71WD and a fully assembled g-TuRC [15]. Whether this recruitment requires specific docking proteins on the MT wall is not known. Such proteins recruiting g-tubulin complexes to cytoplasmic MTs were recently identified in fission yeast [16], but no obvious homologues have been identified in any other eukaryotes. A distributed centrosome at the plant cell cortex? Vertebrate centrosomes are involved in a range of cellular events in addition to MTOCs. They are notably involved in cell cycle transitions, cellular responses to stress and organization of signal transduction pathways. Organization of the plant cortical cytoskeleton is undoubtedly tightly coordinated with the cell cycle, though most players involved in molecular crosstalk between the cell cycle machinery and the cytoskeleton are not identified. Nevertheless, in addition to g-TuRC components, there is converging evidence that a number of functions and proteins are conserved between the animal centrosome (Figure 1 Legend Continued )accepted view of the molecular arrangement of the g-TuRC (the so-called ‘template’ model) is presented here. In this model, 6–7 g-TuSC subcomplexes (each composed of 2 g-tubulins, one GCP2 and one GCP3) are held together in a helicoidal conformation by other GCP subunits that form the cap structure of the g-TuRC. Arrangement of g-tubulin molecules in the complex provides a template for initiating a/b-tubulin heterodimer polymerization and MT assembly. The genome of Arabidopsis (and other plants) possesses clear orthologues of g-TuSC components, as well as equivalents of most cap components of the g-TuRC (Table 1). (c) Emergence of parallel MT arrays at the cortex. The cortex is shown as a two-dimension plane. Stochastic MT dynamic instability. MT Plus ends alternate between phases of polymerization and depolymerization [9]. Shown is a MT undergoing catastrophe, rescued (middle panel), then switching to a polymerization phase (right panel). MT nucleation from pre-existing MT producing branched structures [8]. Encounter between two MTs growing in opposite directions [37]. The MT growing in the minority direction undergoes catastrophe (middle panel) and complete depolymerization (right panel). Release of MT from nucleation site by a putative katanin severing activity. Shallow angle MT encounters usually result in MT co-alignment and bundling [45] (right panel). Steep angle MT encounters usually result in depolymerization of the encountering MT [45] (right panel). Depolymerization of the original MTs after MT nucleation [8]. Recruitment of cytoplasmic g-tubulin on the side of MTs and nucleation of new MT [8]. Cortical arrays contain around 80% of MTs growing in the same direction [37]. A MT growing in the opposite direction can still be incorporated into bundles formed of MTs of opposite polarity. www.sciencedirect.com Current Opinion in Plant Biology 2007, 10:1–7 Please cite this article in press as: Pastuglia M, Bouchez D, Molecular encounters at microtubule ends in the plant cell cortex, Curr Opin Plant Biol (2007), doi:10.1016/j.pbi.2007.08.001 COPLBI-475; NO OF PAGES 7 4 Cell Biology Table 1 g-TuRC components in Arabidopsis. AGI name Name At3g61650 At5g05620 At5g17410 At5g06680 At3g53760 At1g80260 At1g20570 At3g43610 At5g05970 AtTUBG1 AtTUBG2 AtSpc97 AtSpc98 AtGCP4 AtGCP5a AtGCP5b AtGCP6 ? At-Nedd1 MW (kDa) Drosophila Xenopus Human 53 53 77 95 86 106 109 127 85 g-tubulin (2 genes) Dgrip84 Dgrip91 Dgrip75 Dgrip128 g-tubulin Xgrip110 Xgrip109 Xgrip76 Xgrip133 g-tubulin (2 genes) hGCP2 hGCP3 hGCP4 hGCP5 Dgrip163 Dgp71WD Xgrip210 X-Nedd1 hGCP6 GCP-WD (NEDD1) Saccharomyces cerevisiae Schizosaccharomyces pombe Tub4p Tubg1 Spc97p Spc98p Alp4p Alp6p Gfh1p Alp16 The AGI names refer to standard nomenclature for Arabidopsis genome annotation. g-TuRC proteins are conserved across animal species and contain conserved g-ring protein (grip) motifs. AtSpc97 and AtSpc98, as well as AtGCP4, 5a and 5b are clear orthologues of their animal and yeast counterparts on the basis of sequence similarity and phylogeny reconstruction. Although a bona fide member of the GCP family, the position of At3g43610 is less clear with respect to human and Drosophila proteins. Arabidopsis is the only species to have two distinct GCP5 isoforms, sharing 80% similarity. Despite this duplication, it is interesting to note that mutations in AtGCP5a result in embryo lethality (http://www.seedgenes.org/; emb1427-1 and emb1427-2). Depletion of g-tubulin in Arabidopsis also results in gametophytic lethality [12,13]. Other g-TuRC components have not been functionally analyzed up to now in plants. and the organization of the cortical cytoskeleton in plants. For example, several proteins located on, or involved in formation of the PPB have centrosomal counterparts; the FASS protein is a protein phosphatase PP2A regulatory subunit involved in the setup of the PPB [17]. Its homologue in Caenorhabditis elegans, RSA-1, was recently shown to recruit a PP2A complex to the centrosome, where it is involved in spindle assembly and MT outgrowth from the centrosome [18]. Similarly, the cyclindependent kinase CDKA is recruited to the late PPB in plant cells entering mitosis [19]. In mammalian cells, the initial activation of cyclin B1-Cdk1 in early prophase takes place at the centrosome, before spreading to the rest of the cell to induce nuclear and cytoplasmic mitotic events [20]. Interestingly, TONNEAU has been recently shown to co-purify with Arabidopsis CDKA [21]. The TON protein is required for PPB formation and shares a common domain with a human centrosomal protein, FOP [22,23] (Azimzadeh et al., unpublished). Therefore, the view that higher plant cells, being devoid of centrosomes, centrioles, cilia and flagella, have lost the corresponding proteomes (e.g. [24]) may deserve more attention. In particular, it should be considered that centrosomal proteins, especially pericentriolar matrix ones, are notably difficult to analyze with standard bioinformatic methods because of strong compositional biases and richness in coil-coiled regions that prevent efficient similarity searches. MT density by creating seeds for new MT growth [25]. Katanin is a heterodimeric enzyme containing a p60 catalytic subunit and a p80 regulatory subunit and uses energy from ATP hydrolysis to sever MTs. The Arabidopsis p60 subunit is sufficient to sever MTs in vitro [26]. Its overexpression in vivo leads to numerous short bundled MTs, probably resulting from increased severing activity of p60 subunits [27]. Genetic analysis indicated a role of p60 katanin in organizing cortical MTs; in absence of katanin, MT organization at the cortex is impaired, both in rice [28] and Arabidopsis [29,30,31], and MTs are longer and sparser [31]. Proteins at MT plus ends MT assembly/disassembly does not occur by simple helical addition/subtraction of individual a/b subunits, and growing/shrinking MT ends adopt particular conformations, providing structural cues for specific recognition by cellular proteins [1] (Figure 1a). Since their first discovery in 1999 because of improvements in imaging technologies, a growing number of proteins that track MT plus ends (+TIPs) have been identified in animal and yeast cells [32]. Some bind directly to MTs, others locate at MT plus ends via interaction with the first ones. Owing to their specific localization, +TIPs influence MT dynamics, and also enable the growing plus end to search for specific cellular targets like organelles, membranes, microfilaments or other MTs. In animals, these proteins have been involved in MT attachment to the cortex [32]. Severing releases new MT ends Time-lapse observations of individual MTs revealed that most newly nucleated MTs do not remain anchored at their nucleation sites but are released from initiation sites [9] (Figure 1c). The MT-severing katanin is likely to perform such activity. Severing of newly formed MTs then recycles free g-tubulin complexes able to nucleate new MTs. Katanin severing-activity could also increase Current Opinion in Plant Biology 2007, 10:1–7 The study of +TIPs is just beginning in plants, where homology searches indicate that some animal +TIPs are conserved [33]. The most studied +TIP in plants is EB1, a protein now considered a key element among plus end players in animals [32]. In addition to its localization at MT plus end, EB1 is also present at the centrosome of animal cells, independently of MTs [32]. The localization www.sciencedirect.com Please cite this article in press as: Pastuglia M, Bouchez D, Molecular encounters at microtubule ends in the plant cell cortex, Curr Opin Plant Biol (2007), doi:10.1016/j.pbi.2007.08.001 COPLBI-475; NO OF PAGES 7 Molecular encounters at microtubule ends in the plant cell cortex Pastuglia and Bouchez 5 pattern in plants of the EB1 protein family is still debated. Overexpression of AtEB1a-GFP fusions labels not only MT plus ends [34] but also MT minus ends [35] as well as endosomes [36]. Using an inducible promoter to tightly control EB1-GFP expression, Dixit et al. showed that two out of three Arabidopsis EB1 proteins (AtEB1a and AtEB1b) display comet-like structures, whereas AtEB1c, the most distant from human EB1, localized to the nucleus [37]. Beyond their use as markers for time-lapse imaging of MTs, the functional characterization of EB1 proteins in plants still awaits. In this respect, recent isolation of an Arabidopsis mutant in the a-tubulin 5 gene clarifies the interaction of plant EB1 with MTs [38]. This mutation affects a critical residue at the a/b interface and stabilizes mutant MTs by preventing GTP hydrolysis in b-tubulin. Interestingly, in mutant cells, EB1b associates to lateral walls of MTs, indicating that EB1b recognizes the presence of GTP-b-tubulins in the MT lattice, normally restricted to GTP-capped plus ends [38]. Apart from EB1, very few proteins localized at MT plus ends have been characterized in plants. One is SPR1, a member of a plant-specific protein family involved in cortical MT organization and anisotropic cell growth [39]. SPR1 associates to MTs [40] with a preferred accumulation at MT growing ends in interphase cortical arrays [41]. Whether SPR1 binds directly to MT plus ends or rather via interaction with another +TIP is not known. Another plant +TIP is ATK5, a kinesin primarily important for spindle morphogenesis. In addition to spindle assembly, ATK5 may be involved in cortical array organization during PPB formation [42]. Indeed, ATK5 localizes to MTs of the PPB and mutation in atk5 gene affects PPB narrowing [42,43]. In vitro, ATK5 can co-align and bundle MTs after their encounter, suggesting a role in MT capture and co-alignment [42]. Cortical array organization: action at MT ends During interphasic cell growth, MT arrays in the plant cell cortex are typically unfocused, associated to the plasma membrane and organized in parallel arrays transverse to the elongation axis. Surprisingly, this organization can be switched to a radial MT array emanating from a single particle located in the vicinity of the nucleus, simply by overexpression of a particular kinesin fused to GFP [44]. This observation indicates that acting on a limited number of key activities may form profoundly different MT arrays. Studying dynamics of individual MTs in the plant cell cortex has shed light on the way cortical arrays are set up [37,45,46] (Figure 1c). Interactions at MT ends appear to play an important role in addition to the influence of a variety of MT-associated proteins that act on specific parameters of MT dynamic instability and/or favour www.sciencedirect.com bundling [47,48]. MTs at the cell cortex are tightly connected to the plasma membrane, and hence grow in a two-dimensional cortical space; in such a molecularly crowded space, encounters between growing MT plus ends and pre-existing MTs are frequent. A steep angle of collision usually induces disassembly of the encountering MT; when the angle is shallow, the growth trajectory of the encountering MT is adjusted to align to the existing MT [45]. This property is sufficient to explain selforganization of MTs into parallel arrays [45]. Within cortical arrays, MTs tend to acquire a common polarity by selective stabilization of concordant MTs [37]. Discordant MTs branching from pre-existing MTs [8] at a 408 angle may counterbalance this tendency to homogeneity, allowing exploration of the cortical space and adaptation of MT arrays to cellular or environmental cues. Group behaviour of MTs within a cell has recently revealed another level of complexity to cortical MT array organization [46]. According to this study, the plant cell cortex is a mosaic of polarized MTs domains, rotating and reorienting during elongation. Plus end dynamics appear all the more important during PPB formation. At the onset of division, MTs are progressively condensed and bundled into a narrow band, concomitant with gradual disappearance of other cortical MTs. During this stage, MTs outside the PPB become more dynamic and longer, which has been proposed to contribute to PPB formation via a ‘search and capture’ mechanism [49]. This change in dynamics could allow MT plus ends to probe the cortex more efficiently, and the MTs to be selectively stabilized and bundled as they reach the division site [49]. In addition, MT nucleating sites as revealed by g-tubulin localization in PPB microtubules [7] probably insert new MTs into the developing PPB. A search and capture mechanism of MT plus ends may also exist at this stage, for endoplasmic MTs establishing connections between the nucleus and the PPB [50]. Conclusions Despite significant advances concerning proteins that interact with MTs, information is still scattered and disconnected, and there is much to learn about key factors that organize the plant cell cortical cytoskeleton. Many mechanistic, molecular and cellular aspects remain poorly understood, such as connections and crosstalk of the MT cytoskeleton with the cell cycle, the plasma membrane and the cell wall. There are undoubtedly many more players at MT ends than the few identified. Searching for interacting partners of +TIPs could help identify new proteins acting on MT dynamics and connecting MT ends to other cellular structures such as actin, organelles and membranes. At the minus end, beyond g-tubulin complexes, the regulation of the distribution and activity of MT nucleation sites in the cortical space is crucial. Future progress relies on combining approaches such as Current Opinion in Plant Biology 2007, 10:1–7 Please cite this article in press as: Pastuglia M, Bouchez D, Molecular encounters at microtubule ends in the plant cell cortex, Curr Opin Plant Biol (2007), doi:10.1016/j.pbi.2007.08.001 COPLBI-475; NO OF PAGES 7 6 Cell Biology genetic and in vivo cytologic analyses, and the use of cellfree systems. Comparative genomics will help identify shared components of the distributed MT organizer of plant cells and eukaryotic MTOCs but will not replace designing methods to identify plant-specific cellular proteins that interact with the MT wall and the minus and plus ends. In this respect, developing high-throughput methods to characterize subcellular proteomes such as what was achieved for the human centrosome or mitotic spindle proteomes would be of tremendous significance for plant cell biology. Acknowledgements Work in our laboratory is supported by grants from the French Ministry for Research and by the Région Île-de-France for imaging facilities. References and recommended reading Papers of particular interest, published within the annual period of review, have been highlighted as: of special interest of outstanding interest 1. Nogales E, Wang HW: Structural mechanisms underlying nucleotide-dependent self-assembly of tubulin and its relatives. Curr Opin Struct Biol 2006, 16:221-229. 2. Paredez AR, Somerville CR, Ehrhardt DW: Visualization of cellulose synthase demonstrates functional association with microtubules. Science 2006, 312:1491-1495. Using a yellow fluorescent protein fusion to cellulose synthase, the authors demonstrate for the first time that cellulose synthase complexes move in the cell membrane along trajectories tightly coupled to cortical MTs. 12. Binarova P, Cenklova V, Prochazkova J, Doskocilova A, Volc J, Vrlik M, Bogre L: Gamma-tubulin is essential for acentrosomal microtubule nucleation and coordination of late mitotic events in Arabidopsis. Plant Cell 2006, 18:1199-1212. RNAi depletion of g-tubulin in Arabidopsis is shown to compromise cortical MT organization and to strongly affect cytokinesis. MT regrowth experiment in RNAi seedlings and in vitro polymerization assays using gtubulin immunodepleted extracts show that g-tubulin is essential for MT nucleation. 13. Pastuglia M, Azimzadeh J, Goussot M, Camilleri C, Belcram K, Evrard JL, Schmit AC, Guerche P, Bouchez D: Gamma-tubulin is essential for microtubule organization and development in Arabidopsis. Plant Cell 2006, 18:1412-1425. The Arabidopsis genome encodes two g-tubulin genes shown to have redundant function in this study. Knocking down both g-tubulin genes induces gamete lethality and aberrant spindle formation. The use of a less severe combination of alleles leading to seedling death three weeks after germination enabled following MT array dynamics during cell division. The same developmental defects as in RNAi-depleted plants [12] were observed. 14. Erhardt M, Stoppin-Mellet V, Campagne S, Canaday J, Mutterer J, Fabian T, Sauter M, Muller T, Peter C, Lambert AM et al.: The plant Spc98p homologue colocalizes with gamma-tubulin at microtubule nucleation sites and is required for microtubule nucleation. J Cell Sci 2002, 115:2423-2431. 15. Verollet C, Colombie N, Daubon T, Bourbon HM, Wright M, Raynaud-Messina B: Drosophila melanogaster gamma-TuRC is dispensable for targeting gamma-tubulin to the centrosome and microtubule nucleation. J Cell Biol 2006, 172:517-528. 16. Janson ME, Setty TG, Paoletti A, Tran PT: Efficient formation of bipolar microtubule bundles requires microtubule-bound gamma-tubulin complexes. J Cell Biol 2005, 169:297-308. 17. Camilleri C, Azimzadeh J, Pastuglia M, Bellini C, Grandjean O, Bouchez D: The Arabidopsis TONNEAU2 gene encodes a putative novel PP2A regulatory subunit essential for the control of cortical cytoskeleton. Plant Cell 2002, 14:833-845. 18. Schlaitz AL, Srayko M, Dammermann A, Quintin S, Wielsch N, MacLeod I, de Robillard Q, Zinke A, Yates JR 3rd, Muller-Reichert T et al.: The C. elegans RSA complex localizes protein phosphatase 2A to centrosomes and regulates mitotic spindle assembly. Cell 2007, 128:115-127. 3. Wiese C, Zheng Y: Microtubule nucleation: gamma-tubulin and beyond. J Cell Sci 2006, 119:4143-4153. 4. Mazia D: Centrosomes and mitotic poles. Exp Cell Res 1984, 153:1-15. 5. Horio T, Oakley BR: Expression of Arabidopsis gamma-tubulin in fission yeast reveals conserved and novel functions of gamma-tubulin. Plant Physiol 2003, 133:1926-1934. 19. Weingartner M, Binarova P, Drykova D, Schweighofer A, David JP, Heberle-Bors E, Doonan J, Bogre L: Dynamic recruitment of Cdc2 to specific microtubule structures during mitosis. Plant Cell 2001, 13:1929-1943. 6. Panteris E, Apostolakos P, Graf R, Galatis B: Gamma-tubulin colocalizes with microtubule arrays and tubulin paracrystals in dividing vegetative cells of higher plants. Protoplasma 2000, 210:179-187. 20. Jackman M, Lindon C, Nigg EA, Pines J: Active cyclin B1-Cdk1 first appears on centrosomes in prophase. Nat Cell Biol 2003, 5:143-148. 7. Liu B, Marc J, Joshi HC, Palevitz BA: A gamma-tubulin-related protein associated with the microtubule arrays of higher plants in a cell cycle-dependent manner. J Cell Sci 1993, 104:1217-1228. 21. Van Leene J, Stals H, Eeckhout D, Persiau G, Van Slijke E, Van Isterdael G, De Clercq A, Bonnet E, Laukens K, Remmerie N et al.: A tandem affinity purification-based technology platform to study the cell cycle interactome in Arabidopsis thaliana. Mol Cell Proteomics 2007, 6:1226-1238. 8. Murata T, Sonobe S, Baskin TI, Hyodo S, Hasezawa S, Nagata T, Horio T, Hasebe M: Microtubule-dependent microtubule nucleation based on recruitment of gamma-tubulin in higher plants. Nat Cell Biol 2005, 7:961-968. Using a cell-free system enabling MT polymerization on plasma-membrane ghosts, this elegant study establishes that MT nucleation at the cortex involves recruitment of g-tubulin on pre-existing MTs. Using timelapse imaging, the authors show that new MTs branch from pre-existing MT at a well defined 408 angle. 22. Delaval B, Letard S, Lelievre H, Chevrier V, Daviet L, Dubreuil P, Birnbaum D: Oncogenic tyrosine kinase of malignant hemopathy targets the centrosome. Cancer Res 2005, 65:7231-7240. 23. Traas J, Bellini C, Nacry P, Kronenberger J, Bouchez D, Caboche M: Normal differentiation patterns in plants lacking microtubular preprophase bands. Nature 1995, 375:676-677. Shaw SL, Kamyar R, Ehrhardt DW: Sustained microtubule treadmilling in Arabidopsis cortical arrays. Science 2003, 300:1715-1718. 24. Li JB, Gerdes JM, Haycraft CJ, Fan Y, Teslovich TM, May-Simera H, Li H, Blacque OE, Li L, Leitch CC et al.: Comparative genomics identifies a flagellar and basal body proteome that includes the BBS5 human disease gene. Cell 2004, 117:541-552. 10. Van Bruaene N, Joss G, Van Oostveldt P: Reorganization and in vivo dynamics of microtubules during Arabidopsis root hair development. Plant Physiol 2004, 136:3905-3919. 25. Roll-Mecak A, Vale RD: Making more microtubules by severing: a common theme of noncentrosomal microtubule arrays? J Cell Biol 2006, 175:849-851. 11. Reilein A, Yamada S, Nelson WJ: Self-organization of an acentrosomal microtubule network at the basal cortex of polarized epithelial cells. J Cell Biol 2005, 171:845-855. 26. Stoppin-Mellet V, Gaillard J, Vantard M: Functional evidence for in vitro microtubule severing by the plant katanin homologue. Biochem J 2002, 365:337-342. 9. Current Opinion in Plant Biology 2007, 10:1–7 www.sciencedirect.com Please cite this article in press as: Pastuglia M, Bouchez D, Molecular encounters at microtubule ends in the plant cell cortex, Curr Opin Plant Biol (2007), doi:10.1016/j.pbi.2007.08.001 COPLBI-475; NO OF PAGES 7 Molecular encounters at microtubule ends in the plant cell cortex Pastuglia and Bouchez 7 27. Stoppin-Mellet V, Gaillard J, Vantard M: Katanin’s severing activity favors bundling of cortical microtubules in plants. Plant J 2006, 46:1009-1017. 39. Nakajima K, Kawamura T, Hashimoto T: Role of the SPIRAL1 gene family in anisotropic growth of Arabidopsis thaliana. Plant Cell Physiol 2006, 47:513-522. 28. Komorisono M, Ueguchi-Tanaka M, Aichi I, Hasegawa Y, Ashikari M, Kitano H, Matsuoka M, Sazuka T: Analysis of the rice mutant dwarf and gladius leaf 1. Aberrant katanin-mediated microtubule organization causes upregulation of gibberellin biosynthetic genes independently of gibberellin signaling. Plant Physiol 2005, 138:1982-1993. 40. Nakajima K, Furutani I, Tachimoto H, Matsubara H, Hashimoto T: SPIRAL1 encodes a plant-specific microtubule-localized protein required for directional control of rapidly expanding Arabidopsis cells. Plant Cell 2004, 16:1178-1190. 29. Burk DH, Liu B, Zhong R, Morrison WH, Ye ZH: A katanin-like protein regulates normal cell wall biosynthesis and cell elongation. Plant Cell 2001, 13:807-827. 30. Bichet A, Desnos T, Turner S, Grandjean O, Hofte H: BOTERO1 is required for normal orientation of cortical microtubules and anisotropic cell expansion in Arabidopsis. Plant J 2001, 25:137-148. 31. Bouquin T, Mattsson O, Naested H, Foster R, Mundy J: The Arabidopsis lue1 mutant defines a katanin p60 ortholog involved in hormonal control of microtubule orientation during cell growth. J Cell Sci 2003, 116:791-801. 32. Morrison EE: Action and interactions at microtubule ends. Cell Mol Life Sci 2007, 64:307-317. 33. Bisgrove SR, Hable WE, Kropf DL: +TIPs and microtubule regulation. The beginning of the plus end in plants. Plant Physiol 2004, 136:3855-3863. 34. Van Damme D, Bouget FY, Van Poucke K, Inze D, Geelen D: Molecular dissection of plant cytokinesis and phragmoplast structure: a survey of GFP-tagged proteins. Plant J 2004, 40:386-398. 35. Chan J, Calder GM, Doonan JH, Lloyd CW: EB1 reveals mobile microtubule nucleation sites in Arabidopsis. Nat Cell Biol 2003, 5:967-971. 36. Mathur J, Mathur N, Kernebeck B, Srinivas BP, Hulskamp M: A novel localization pattern for an EB1-like protein links microtubule dynamics to endomembrane organization. Curr Biol 2003, 13:1991-1997. 37. Dixit R, Chang E, Cyr R: Establishment of polarity during organization of the acentrosomal plant cortical microtubule array. Mol Biol Cell 2006, 17:1298-1305. The authors utilized GFP-tagged EB1 protein to assess MT polar arrangement within cortical arrays in Arabidopsis epidermal cells and in tobacco BY-2 cells. Establishment of MT array polarity seems to be acquired through stabilization of MTs oriented in a preferential direction and progressive enrichment of such MTs within the array. This study also provides an analysis of the subcellular localization of the three Arabidopsis EB1 isoforms. 38. Ishida T, Kaneko Y, Iwano M, Hashimoto T: Helical microtubule arrays in a collection of twisting tubulin mutants of Arabidopsis thaliana. Proc Natl Acad Sci USA 2007, 104:8544-8549. This interesting study reports on the identification of 32 a-tubulin or btubulin mutants inducing helical growth in Arabidopsis plantlets. All mutations correspond to missense or deletion mutations at conserved residues in tubulins. Helical growth results from dysfunction of cortical MTs that arrange into helical arrays of fixed handedness rather than normal transverse arrays. Strikingly, all right-handed helical growth mutants had lefthanded MT helical arrays. Analysis of the dynamic behaviour of cortical MTs in these mutants supports the view that MT dynamics is one of the major factors contributing to the final handedness of MT arrays. www.sciencedirect.com 41. Sedbrook JC, Ehrhardt DW, Fisher SE, Scheible WR, Somerville CR: The Arabidopsis sku6/spiral1 gene encodes a plus end-localized microtubule-interacting protein involved in directional cell expansion. Plant Cell 2004, 16:1506-1520. 42. Ambrose JC, Cyr R: The kinesin ATK5 functions in early spindle assembly in Arabidopsis. Plant Cell 2007, 19:226-236. A previous study has shown that the ATK5 kinesin displays +TIP activity conferred by regions outside the minus end directed motor domain [43]. This article focused on the role of ATK5 in early spindle assembly. Analysis of MT organization during this stage revealed an essential role of ATK5 in controlling spindle length, width and integrity. 43. Ambrose JC, Li W, Marcus A, Ma H, Cyr R: A minus-end-directed kinesin with plus-end tracking protein activity is involved in spindle morphogenesis. Mol Biol Cell 2005, 16:1584-1592. 44. Goto Y, Asada T: Excessive expression of the plant kinesin TBK5 converts cortical and perinuclear microtubules into a radial array emanating from a single focus. Plant Cell Physiol 2007, 48:753-761. This study shows that overexpression of the plant specific kinesin TBK5 fused to GFP is sufficient to transform cortical MT arrays into a radial MT array emanating from a single particle located in close proximity to the nucleus and formed of assembled GFP-TBK5. Expression of mutant forms or truncated versions of TBK5 identify domains essential for particle formation. 45. Dixit R, Cyr R: Encounters between dynamic cortical microtubules promote ordering of the cortical array through angle-dependent modifications of microtubule behavior. Plant Cell 2004, 16:3274-3284. 46. Chan J, Calder G, Fox S, Lloyd C: Cortical microtubule arrays undergo rotary movements in Arabidopsis hypocotyl epidermal cells. Nat Cell Biol 2007, 9:171-175. Using GFP markers, the authors followed MT reorientation for up to 24 h in elongating hypocotyl cells. This revealed that MT arrays were a mosaic of polarized domains within a cell, which rotate and undergo striking reorientation over time. 47. Sasabe M, Machida Y: MAP65: a bridge linking a MAP kinase to microtubule turnover. Curr Opin Plant Biol 2006, 9:563-570. 48. Sedbrook JC: MAPs in plant cells: delineating microtubule growth dynamics and organization. Curr Opin Plant Biol 2004, 7:632-640. 49. Vos JW, Dogterom M, Emons AM: Microtubules become more dynamic but not shorter during preprophase band formation: a possible ‘search-and-capture’ mechanism for microtubule translocation. Cell Motil Cytoskeleton 2004, 57:246-258. 50. Dhonukshe P, Mathur J, Hulskamp M, Gadella TW Jr: Microtubule plus-ends reveal essential links between intracellular polarization and localized modulation of endocytosis during division-plane establishment in plant cells. BMC Biol 2005, 3:11. Current Opinion in Plant Biology 2007, 10:1–7 Please cite this article in press as: Pastuglia M, Bouchez D, Molecular encounters at microtubule ends in the plant cell cortex, Curr Opin Plant Biol (2007), doi:10.1016/j.pbi.2007.08.001