* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download lac
Epigenetics of diabetes Type 2 wikipedia , lookup
Genetic engineering wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Genomic imprinting wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Dominance (genetics) wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Ridge (biology) wikipedia , lookup
Genome evolution wikipedia , lookup
History of genetic engineering wikipedia , lookup
Minimal genome wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Gene expression programming wikipedia , lookup
Designer baby wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Mir-92 microRNA precursor family wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Frameshift mutation wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Genome (book) wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Gene expression profiling wikipedia , lookup
Oncogenomics wikipedia , lookup
Microevolution wikipedia , lookup
List of Mutants in The Hills Have Eyes wikipedia , lookup
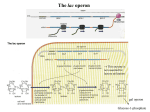
LECTURE 2 OUTLINE GENETIC ANALYSIS OF GENE REGULATION Negative and Positive Regulation -negatively and positively regulated operons behave very differently in genetic tests Negative Regulation Positive Regulation -inactivation of regulator leads -inactivation of regulator leads to CONSTITUTIVE expression to NON-INDUCIBLE -constitutive mutations are expression RECESSIVE to wildtype -non-inducible mutations are RECESSIVE to wildtype -gain-of-function mutations lead -gain-of-function mutations lead to NON-INDUCIBLE to CONSTITUTIVE expression expression - constitutive mutations are DOMINANT to wildtype -non-inducible mutations are DOMINANT to wildtype Identification of Cis and Trans Acting Factors Cis-Acting Factor -DNA sequence that functions exclusively as a DNA sequence -affects only DNA to which it is physically linked -eg. promoter/operator Trans-Acting Factor -diffusible gene product -can affect other DNA molecules -eg. negative or positive regulator -mutations cannot be -mutations can be complemented complemented -mutations may be recessive or -mutations may be recessive or dominant dominant regulatory proteins are trans-acting factors that recognize cisacting elements to control gene expression NEGATIVE REGULATION – THE LAC OPERON -Jacob & Monod’s analysis of negative regulation of the lac operon in the 1950s established the framework for all other gene regulation studies -Nobel prize 1965 The lac operon -bacterial genes organized in clusters according to function, OPERONS -expressed as part of a single POLYSCISTRONIC transcript, subject to coordinate control -enzymes for lactose metabolism are INDUCIBLE -paradigm determined from genetic analyses of lac mutants -based on COMPLEMENTATION ANALYSIS -introduce F’ factors carrying mutated genes (plasmid onto which lac- mutations have been recombined) to create MERODIPLOIDS -determine dominance vs. recessivenss -identify complementation groups/genes Identification of the structural genes -one group of lac mutants unable to grow on lactose (Lac-) -STRUCTURAL GENES identified by complementation analysis -identify genes involved by complementation analysis Strain F’ lacZ Y /lacZ+Y+ F’ lacZ+Y+/lacZ1-Y+ F’ lacZ2-Y+/lacZ+Y+ F’ lacZ2-Y+/lacZ1-Y+ + + Phenotype + Lac Lac+ Lac+ Lac- -most mutants are recessive to wildtype therefore must inactivate genes required for lactose utilization -2 complementation groups/genes lacZ and lacY Identification of the regulator LacI 1) dominant non-inducible Lac- mutants -one group of mutants were dominant Lac- mutants -these mutations make cell Lac- even if another good copy of lac operon is present -lacIs Strain + + + lacI Z Y lacIsZ+Y+ F’ lacI+Z+Y+/lacI+Z+Y+ F’ lacIs Z+Y+/lacI+Y+Z+ Phenotype + Lac LacLac+ Lac- -lacIs mutations alter LacI such that it binds the operator even in the presence of inducer and prevents expression of the operon 2) recessive constitutive mutants -one group of mutants make the cells Lac+ constitutive, rather than Lac-complementation tests determined that these mutants are recessive Strain + + + lacI Z Y lacI-Z+Y+ F’ lacI+Z+Y+/lacI+Z+Y+ F’ lacI-Z+Y+/lacI+Z+Y+ Phenotype Inducible Lac+ Constitutive Lac+ Inducible Lac+ Inducible Lac+ -complementation tests between recessive constitutive mutants determined that they are all in lacI - the mutations knock-out LacI 3) dominant constitutive mutants -one group of mutants make the cells Lac+ constitutive, rather than Lac-complementation tests determined that these mutants are dominant Strain + + + lacI Z Y lacI-dZ+Y+ F’ lacI+Z+Y+/lacI+Z+Y+ F’ lacI-dZ+Y+/lacI+Z+Y+ Phenotype Inducible Lac+ Constitutive Lac+ Inducible Lac+ Constitutive Lac+ -lacI-d mutations alter LacI such that it binds the operator even in the presence of inducer and prevents expression of the operon Identification of cis-acting sites i) promoter mutants -another group of lac mutants unable to grow on lactose could not be complemented -these mutants lie in the lac promoter region -an F’ carrying a p- mutant, but wildtype copies of lacZ and lacY can not be complemented in a strain carrying either mutant lacZ or lacY genes on the chromosome CIS-ACTING REGULATORY SITE Strain + + + p lacZ lacY Phenotype Lac + p-lacZ+lacY+ Lacp+lacZ-lacY+ Lacp+lacZ+lacYLac+ + + + + + F’ p lacZ Y /p lacZ Y Lac+ F’ p-lacZ+Y+/p+lacZ-Y+ LacF’ p-lacZ+Y+/p+lacZ+YLacF’ p-lacZ+Y+/p+lacZ+Y+ Lac+ -these mutations prevent RNAP from initiating transcription at the lac promoter ii) lacOc mutants -another group of mutants resembled the lacI- mutants, they were constitutive -these mutants, however, could not be complemented (cis) and were also dominant Strain Phenotype lacO lacZ Inducible Lac+ lacOclacZ+ Constitutive Lac+ F’ lacO+lacZ+/lacO+lacZ+ Inducible Lac+ F’ lacOclacZ+/lacO+lacZ+ Constitutive Lac+ -mutations were in LacI repressor binding site or OPERATOR -prevent binding of LacI, so expression is always on + + Genotype Phenotype +IPTG -IPTG wild type + LacZLacPLacOc + + LacI+ + LacI(-d) + + LacI(s) LacOcLacI(s) + +