* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Chromosome Rearrangements Concepts: Chromosome
Molecular cloning wikipedia , lookup
Copy-number variation wikipedia , lookup
Saethre–Chotzen syndrome wikipedia , lookup
Genealogical DNA test wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Gene expression profiling wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Oncogenomics wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Human genome wikipedia , lookup
Non-coding DNA wikipedia , lookup
Comparative genomic hybridization wikipedia , lookup
DNA supercoil wikipedia , lookup
Ridge (biology) wikipedia , lookup
Minimal genome wikipedia , lookup
Genomic library wikipedia , lookup
History of genetic engineering wikipedia , lookup
Hybrid (biology) wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Designer baby wikipedia , lookup
Point mutation wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Gene expression programming wikipedia , lookup
DiGeorge syndrome wikipedia , lookup
Genome evolution wikipedia , lookup
Genomic imprinting wikipedia , lookup
Skewed X-inactivation wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Genome (book) wikipedia , lookup
Microevolution wikipedia , lookup
Segmental Duplication on the Human Y Chromosome wikipedia , lookup
Y chromosome wikipedia , lookup
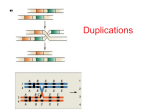
Lecture 21; 2007 Biology 207; Section B2; Good Chromosome Rearrangements Readings: Griffiths et al: 8th Edition: pp. 496-508, 7th Edition: Ch. 17 pp 523-554 Assigned Problems: 8th Ch. 15: 22-28, 40-43, 47, 7th Ch. 17: 1-23 Concepts: How can chromosomes be altered? 1. Chromosomes can undergo physical rearrangements of their DNA, which include deletions, duplications, inversions, and/or translocations of DNA segments. 2. Rearranged chromosomes may pair improperly at meiosis and alter the distribution of chromosomes thereby affecting fertility. 3. Rearrangements can break genes and produce unbalanced gametes (and therefore unbalanced progeny). Chromosome Rearrangements They involve breaks in the DNA duplex -- both strands -- followed by the rejoining of the broken ends. The result is a reorganized chromosome that usually can be transmitted to subsequent generations. If not repaired they are lost and the cell usually dies because it is unbalanced. Example of a normal chromosome : abcdef-o-ghij (with -o- = centromere) Deletions - the loss of a region of DNA from a chromosome abcdef-o-ghij abef-o-ghij where the cd region has been deleted. Duplications - the gain of a region of DNA abcdef-o-ghij if region cd is duplicated it gives abcdcdef-o-ghij Inversions - involve the inversion of a segment of a chromosome abcdef-o-ghij - with breaks between c and d as well as h and i gives abchg-o-fedij Translocations - involve breaks on non-homologous chromosomes with an exchange of parts abcdef-o-ghij and qrst-o-uvw to give abcdef-o-gvw and qrst-o-uhij Deletions There are two types of deletions: interstitial - which have two breaks within the chromosome and the region between there is lost terminal - which involves one break and the segment distal to the break, including the telomere, is lost 1 Lecture 21; 2007 Biology 207; Section B2; Good Pseudo-dominance Deletions remove many genes consecutively positioned along a chromosome. Thus recessive alleles within this hemizygous region will be expressed. The recessive is said to be pseudo-dominant when the homologue is paired with the deletion. Deletions permit mapping the location of genes on a cytogenetic map Deletion loop-absence of a chromosome segment when homologous chromosomes pair (at meiosis) Figure 17-3 (7th) in the text. The location of the deletion maps genes cytogenetically. Thus one can compare the locations of genes on the genetic map (map units -- which represent recombination frequencies) and the cytogenetic map (which represents physical distances --> sequence). To identify a deletion mutation: Deletions can be recognized by several characteristics: 1) Pseudo dominance - involves several adjacent loci 2) Cytologically - deletion loops 3) Usually recessive-lethal - lost essential genes 4) Lack of reverse mutation - unlike many other mutations - cannot revert absence of DNA Duplications Duplications come in two major forms: Tandem - abcdecdef-o-ghij - duplication of a segment is direct Reverse (inverse) - abcde edcf-o-ghij - duplicated segment is reversed Result is extra genes - those within the duplicated region At Meiosis the Duplicated segment is able to pair with homologous region on the same chromosome as well as on the homologue See Fig. 15-19 (8th) 17-1b (7th) ab cde cde f-o-ghij 2 copies X ab cde cde f-o-ghij 2 copies Result: abcdecdecdef-o-ghij 3 copies abcdef-o-ghij 1 copy Duplications can be recognized by: 1) Cytologically - duplication loop Fig 17-9 (7th) 2) Usually not recessive lethal - duplication genes 3) Can revert and at relatively high rate (crossing over) Inversions Chromosome rearrangement with segment inverted. Inversions have two major types, which depend upon the positions of the breakpoints in relation to the centromere 2 Lecture 21; 2007 Biology 207; Section B2; Good Paracentric Inversions have breaks on the same chromosome arms. (| = break) ab|cde|f-o-ghij -> ab edc f o ghij Pericentric Inversions have breaks on different chromosome arms ab|cdef-o-gh|ij -> ab hg-o-fedc ij Pairing at meiosis See Fig 15-22, 15-23 (8th) 17-16, 17-17 (7th). Paracentric inversions – Fig 15-22 (8th) 17-16 (7th) When an inversion homolog pairs with a normal sequence homologue an inversion loop results. The effect of a single cross over event within the loop is the production of an acentric fragment, which is lost and deletion products. These deletion products, if incorporate into a zygote, are usually lethal. Only two of the four gametes would produce viable gametes, both of which are parental in organization: one normal + one inversion. Consequence is that: 1) "recombinants" (vs. parentals) will be reduced in frequency recombinant chromosomes are inviable. 2) Markers within the loop will have an RF of ~ 0 - absolute linkage. The only way a recombinant can be recovered is if there is a double cross over nvolving the same chromatids of the first cross over event. 3) Also inversions inhibit the actual pairing of regions near, or inbetween the break points, then crossing-over can not take place. Pericentric Inversions - Fig 15-23 (8th) 17-17 (7th) The effects are much the same. The actual products are different (no acentric fragments). Note: If both homologues are equivalent (ie. homozygous inversion), then no inversion loop is formed and both chromosomes pair. No abnormal products are formed by crossover events. The only consequence is a linkage map that has an inverted gene order. Translocations – Fig 15-24 (8th) 17-23 (7th) Reciprocal translocation - heterozygous with normal chromosomes. There are consequences for the pairing at meiosis. Two normal homologues (N1, N2) (Brown, Purple) Two translocation chromosomes (T1, T2) Homologous regions of each chromosome synapse Result is a pairing configuration that involves two pairs of homologues. 4 chromosomes in total (8 chromatids) Such a configuration can have several types of segregation depending upon how it lines up on the equatorial plate Homologous paired centromeres disjoin. There are two common patterns of disjunction. Adjacent - 1 segregation T1 with N2 (Top two) N1 with T2 (Bottom two) The result is unbalanced gametes: duplication/deficient 3 Lecture 21; 2007 Biology 207; Section B2; Good Unbalanced gametes are inviable. It is called adjacent because adjacent centromeres move to the same poles. Alternate segregaton It is called alternate because alternate centromeres move to the same pole with the result that: T1 with T2 and N1 with N2 Both products (and all 4 gametes) are balanced and viable. Adjacent-1 and alternate happen equally frequently Only 50% of the products are viable. This reduction in viable gametes reduces the fertility of heterozygote translocation bearing individuals. Only semi-sterile. Translocations and Inversions result in reduced fertility due to inviable gametes. Inviable gametes are meiotic products that are capable of forming sex cells but when joining the normal complementary gametes are unable to form a viable zygote (it is due to the unbalanced gamete). Inversions and Translocations can also affect the linkage between marker loci on the chromosomes involved. Figure 15:19 (Ed. 8 p. 496) 4