* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Genome-wide transcription profiling of aerobic and anaerobic
Signal transduction wikipedia , lookup
Point mutation wikipedia , lookup
Community fingerprinting wikipedia , lookup
Ridge (biology) wikipedia , lookup
Transcriptional regulation wikipedia , lookup
Gene nomenclature wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Biochemical cascade wikipedia , lookup
Genomic imprinting wikipedia , lookup
Paracrine signalling wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Magnesium transporter wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Secreted frizzled-related protein 1 wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Gene regulatory network wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
Gene expression wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Two-hybrid screening wikipedia , lookup
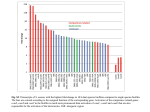
FEMS Microbiology Letters, 364, 2017, fnx006 doi: 10.1093/femsle/fnx006 Advance Access Publication Date: 13 January 2017 Research Letter R E S E A R C H L E T T E R – Physiology & Biochemistry Genome-wide transcription profiling of aerobic and anaerobic Escherichia coli biofilm and planktonic cultures Bihter Bayramoglu, David Toubiana and Osnat Gillor∗ Zuckerberg Institute for Water Research, Blaustein Institutes for Desert Research, Ben Gurion University of the Negev, Midreshet Ben Gurion 84990, Israel ∗ Corresponding author: Zuckerberg Institute for Water Research, Blaustein Institutes for Desert Research, Ben Gurion University of the Negev, Sde Boker Campus, Midreshet Ben Gurion 84990, Israel. Tel: +972-8-6596986; E-mail: [email protected] One sentence summary: This study explores global regulation of Escherichia coli biofilm cultured with and without oxygen. Editor: Jana Jass ABSTRACT Many studies have described the response of the facultative anaerobe, Escherichia coli, to anaerobic conditions, yet they all investigated free-living (planktonic) cells because attempts to cultivate anaerobic E. coli biofilm were mostly unsuccessful. We challenged these findings and cultivated E. coli strain MG1655 biofilm under both aerobic and anaerobic conditions, characterizing the mature biofilm architecture and global gene expression profile. We used RNA sequencing technology to compare stationary phase planktonic cells with mature biofilm, cultured with and without oxygen. Our results suggest that gene expression patterns significantly differ between biofilm and planktonic cultures cultivated under the same oxygenic conditions. The anaerobic E. coli biofilms were slow growing and patchy compared to aerobic biofilms, yet some features were unchanged like the production of extracellular polymeric substances. A closer inspection of the mRNA data revealed that essential cell processes were attenuated in anaerobic biofilms, including protein synthesis, information transfer, cell structure, regulation and transport. Our results suggest that lack of oxygen imposes severe stress on mature biofilms thus limiting the cells’ activity. We further propose that E. coli does not favor growing in anaerobic biofilms and when forced to do so, the cells prevail by attenuating their activity in order to survive. Keywords: sessile; flowcell; transcriptomics INTRODUCTION The gastrointestinal (GI) tract is dominated by diverse anaerobic bacterial communities, even though it is an open system with an oxygen gradient running from its proximal to distal end (Macfarlane and Dillon 2007). The task of sustaining an oxygen-free environment for the dominating gram-positive obligate anaerobes falls onto facultative anaerobes such as Escherichia coli and its close relatives, which metabolize traces of oxygen in the GI biofilm (Jones et al. 2011). In the GI tract, E. coli resides in biofilms—sessile communities of microorganisms associated with the GI epithelial mucosa (Beloin, Roux and Ghigo 2008). Global expression profiles of E. coli biofilms and freeliving (planktonic) cultures showed distinct patterns, negating the assumption that biofilms are sessile cells in stationary phase (Schembri, Kjaergaard and Klemm 2003). Moreover, E. coli cells in a given biofilm may experience different growth conditions and depending on the cells’ position within the biofilm, may also experience aerobic, microaerobic or anaerobic conditions (Karatan and Watnick 2009). The kinetics of E. coli strain MG1655 biofilms grown under aerobic and microaerobic conditions have been Received: 7 May 2016; Accepted: 11 January 2017 C FEMS 2016. All rights reserved. For permissions, please e-mail: [email protected] 1 2 FEMS Microbiology Letters, 2017, Vol. 364, No. 3 described (Beloin et al. 2003; Bjergbæk et al. 2006; Hancock and Klemm 2007), but anaerobic conditions have been suggested to abate this strain’s biofilm formation (Landini and Zehnder 2002; Colón-González, Méndez-Ortiz and Membrillo-Hernández 2004). Under controlled laboratory conditions, planktonic E. coli cells grow rapidly when respiring oxygen. In the absence of oxygen and alternative electron acceptors, the cells switch to anaerobic respiration and fermentation (Partridge et al. 2006). However, it was shown that E. coli anaerobic biofilms could not be formed even when cultures were supplemented with various electron acceptors (Colón-González, Méndez-Ortiz and Membrillo-Hernández 2004), or when longer incubation periods were imposed (Colón-González, Méndez-Ortiz and MembrilloHernández 2004). Furthermore, a survey of over 45 000 random mutants failed to identify an E. coli strain that was able to form a biofilm under anaerobic conditions (Cabellos-Avelar, Souza and Membrillo-Hernández 2006). But, when 77 E. coli strains isolated from various environments were tested, 7 formed a biofilm under anaerobic conditions (Colón-González, Méndez-Ortiz and Membrillo-Hernández 2004). It was further suggested that laboratory E. coli strains could not form a biofilm under anaerobic conditions because lipopolysaccharide and flagellum biosynthesis are attenuated and, therefore, E. coli’s initial attachment efficiency decreases (Landini and Zehnder 2002). As the growth of anaerobic biofilms of laboratory E. coli strains was unsuccessful, studies focused on planktonic cultures, elucidating the transition patterns from aerobic to microaerobic and anaerobic environments (Partridge et al. 2006; Trotter et al. 2011). But, E. coli genome-wide analyses have demonstrated marked differences between the transcriptome of planktonic and biofilm cultures (Beloin, Roux and Ghigo 2008; Wood 2009). Therefore, we hypothesized that E. coli biofilms would respond differently to oxygen limitation than planktonic cells. To address this hypothesis, we developed a method that enables the growth of E. coli strain MG1655 biofilm under anaerobic conditions. We then compared biofilm structure and global gene expression profiles of mature biofilms to stationary-phase planktonic E. coli cultivated with and without oxygen. We hypothesize that the structure and molecular characteristics of the anaerobic biofilm significantly differ from those of the aerobic biofilm, yet the differences between planktonic and biofilm cultures would be more prominent. MATERIAL AND METHODS For detailed descriptions of the material and methods used in this study, see supplementary information file S1. Bacterial strain and culture conditions All experiments were performed with Escherichia coli strain MG1655 (F- lambda- ilvG- rfb-50 rph-1) cultured in M9 minimal medium (Sigma, Rehovot, Israel) supplemented with 4 g L−1 Casein Digest (BD, Franklin Lakes, NJ, USA) at 25◦ C. For each treatment, two biological replicates were prepared and each was replicated at least three times. Overnight planktonic E. coli cultures were refreshed and grown to early exponential phase and then divided for aerobic and anaerobic incubation. The duration of cultivation differs because E. coli cultures grow faster in aerobic compared to anaerobic conditions. The specified growth rates were verified in preliminary experiments (data not shown). Attempts at cultivating E. coli anaerobic biofilms were proven to be unsuccessful when using microtiter plates (Colón-González, Méndez-Ortiz and Membrillo-Hernández 2004) or sand columns (Landini and Zehnder 2002). We applied a flow cell system to culture the biofilms, yet commercial systems are limited by their surface area (as they mostly include a single slide) and by the frequent formation of air bubbles that can lead to the destruction of biofilm architecture (Crusz et al. 2012). To overcome these limitations, we used a custom designed flow cell system (Majeed et al. 2014) in an incubator or anaerobic chamber (Coy Laboratory Products, Grass Lake, MI, USA) for 3 and 6 days, respectively. The duration of cultivation was determined by preliminary experiments (Supplementary files). Extracellular polymeric substances extraction and analysis Extracellular polymeric substances (EPS) was extracted from aerobic and anaerobic biofilms as previously described (Sweity et al. 2011). Confocal laser-scanning microscopy To visualize the biofilm cultures, slides were stained with 300 nM 4-6-diamidino-2-phenylindole (DAPI; Sigma), imaged by confocal laser-scanning microscopy (Zeiss M510, Oberkochen, Germany) and the 3D images were reconstituted using IMARIS software (Bitplane, Zurich, Switzerland). RNA extraction and sequencing Total RNA was extracted (Epicentre, Madison, WI) from two biological replicates of about 1010 planktonic cells and from the slides supporting 35 ± 5 μm3 μm−2 aerobic biofilm and 2.7 ± 0.3 μm3 μm−2 anaerobic biofilm. High-throughput sequencing of the mRNA was carried out by the Genome Center at the Israel Institute of Technology as previously described (Brownstein et al. 2014). Data acquisition and analysis About 90%–95% of all reads were aligned to non-rRNA regions of the E. coli MG1655 genome (accession number U00096.2) using Rockhopper analysis tool (McClure et al. 2013). The sequence reads of all samples were deposited in NCBI’s Gene Expression Omnibus (Edgar 2002) and are accessible through GEO Series accession number GSE72113. Significant differences in gene expression were considered only when expression changed at least 2.5-fold (Fitzgerald, Bonocora and Wade 2014; Shao et al. 2015). Genes were classified with MultiFun (Serres and Riley 2000) and annotated using EcoCyc database (Keseler et al. 2013). An overview of the differentially regulated genes was constructed in heatmaps using R software (Ihaka and Gentleman 1996). Differences of expression levels between the different environments were attempted with statistically stringent approach, integrated in DESeq package (Anders and Huber 2010), and a less stringent approach, integrated in limma package (Ritchie et al. 2015) and Biobase (Gentleman et al. 2004) in R environment. Gene expression distribution among treatments was visualized by boxplot showing that normalized gene counts were reproducible between biological replicates (Fig. S1, Supporting Information). Partitioning around medoids (PAM) clustering was performed to determine the optimal number of clusters of the dataset (Fig. S2, Supporting Information) and principal component analyses (PCA) were performed on the multivariate datasets using pcaMethods Bayramoglu et al. 3 Figure 1. Biofilms of E. coli strain MG1655. Biofilms cultured (A) for 3 days under aerobic conditions and (B) for 6 days under anaerobic conditions. The reconstructed confocal images were obtained after culturing the biofilms in continuous-flow chambers. Each square on the grid of image A is 50 μm and of image B, 80 μm. Table 1. Properties of EPS in E. coli biofilms cultured under aerobic and anaerobic conditions. EPS contents (ppm) Total organic carbon Polysaccharides Total nitrogen Protein Aerobic 44.08 41.2 10.96 12.1 ± ± ± ± 5.27 3.43 1.28 1.51 indicate that E. coli cells maintain their overall biofilm structure despite the oxygen-related stress. Anaerobic 54.24 40.5 13.25 15.6 ± ± ± ± 7.02 3.68 1.79 1.76 package from Bioconductor (http://bioconductor.org) (Stacklies et al. 2007) in the R environment. RESULTS AND DISCUSSION Biofilm formation and structure under anaerobic conditions We cultured the Escherichia coli biofilm with and without oxygen and noted that under anaerobic conditions biofilm development was significantly slower. The mature, 6-days-old, anaerobic biofilm was patchier and thinner compared to the aerobic biofilm (Fig. 1). While the biovolume of aerobic biofilm was 60 ± 10 μm3 μm−2 , anaerobic biofilm had a biovolume of 4 ± 1 μm3 μm−2 . Aerobic biofilms maximum thickness went up to 69.1 ± 15.5 μm with 89.8 ± 5.4% substratum coverage. On the other hand, anaerobic biofilms had a maximum thickness of 50.8 ± 11.5 μm with 70.2 ± 9.5% substratum coverage. This suggests that lack of oxygen limits E. coli biofilm formation but, in contrast to conclusions drawn from previous studies (Landini and Zehnder 2002; Colón-González, MéndezOrtiz and Membrillo-Hernández 2004; Cabellos-Avelar, Souza and Membrillo- Hernández 2006), it does not prevent it. Surprisingly, the EPS properties of the aerobic and anaerobic biofilms including the total organic carbon, polysaccharide, total nitrogen and protein contents (Table 1) were similar in E. coli biofilms grown under both the aerobic and anaerobic conditions (P > 0.05). Moreover, RNA sequencing revealed that the genes promoting EPS production (Ionescu and Belkin 2009), such as the wca, pga and yjb gene operons as well as the dfc pseudo-operon, were similarly expressed in aerobic and anaerobic biofilms (2> q value > 0.5; Table S1, Supporting Information). These results Quantitative analysis of planktonic and biofilm Escherichia coli transcriptomes We first investigated whether the oxygen treatments or growth conditions (planktonic vs biofilm) play a more significant role in differential gene expression profiles of E. coli cells. This was achieved by portraying hierarchal cluster heatmaps of the normalized gene expression (Fig. 2) and by using PCA clustering (Fig. 3). The comparative analyses of the genes differential expression are provided by the heatmaps (Fig. 2) revealing major differences between growth conditions, and to a lesser extent oxygen treatment (Fig. S3, Supporting Information). The optimal number of clusters estimated by the PAM clustering PCA plot suggested a separation into two groups corresponding to the discrepancies between planktonic and biofilm gene expression regardless of the oxygen treatment (Fig. 3). Both clustering analyses (Figs 2 and 3) validated our hypotheses that there are major differences between the E. coli expression profiles under the two cultivation modes (biofilm and planktonic). These differences were considerable amounting to over 40% of the E. coli genome under aerobic conditions and over 60% of the genome under anaerobic conditions (Fig. S4; Tables S2 and S3, Supporting Information). Similarly, global gene expression comparison portrayed by microarray analysis between aerobic biofilm and planktonic Pseudomonas aeruginosa cells revealed dramatic changes, with 26% of the genes showing altered expression rates (Waite et al. 2006). In contrast, a study testing the differences between E. coli cells in the stationary phase and in mature biofilms under aerobic conditions using the microarray technology revealed only a 4% difference in gene expression (Beloin et al. 2003). We could not attribute these dramatic differences among studies to the difference in applied analyses, as an agreement between microarray and RNA-seq has been demonstrated (Malone and Oliver 2011); rather, the different outcomes could be attributed to differences in the biofilm-culturing protocols. Dötsch et al. (2012) reported that there is little consistency in gene expression regulation patterns across various studies comparing planktonic 4 FEMS Microbiology Letters, 2017, Vol. 364, No. 3 Figure 2. Whole-gene expression profile of planktonic and biofilm cultures growing under aerobic and anaerobic conditions. A heatmap of a hierarchal cluster of normalized gene expression for genes that were differentially regulated in planktonic and biofilm E. coli cultured with and without oxygen. The color key represents the overall normalized gene expression pattern indicated by Z-score analysis. A comprehensive list of absolute and differential gene expression for all genes included in this figure is provided in Table S1. Quantitative analysis of aerobic and anaerobic Escherichia coli transcriptomes Figure 3. Cluster analysis of normalized gene expression in E. coli biofilm and planktonic cultures for genes expressed under aerobic and anaerobic conditions. PCA of gene expression levels of biofilms cultured under aerobic and anaerobic conditions (red circles); planktonic cultures under aerobic and anaerobic conditions (blue triangles). PC1 accounts for 77% and PC2 for 16.1% of the variance observed between expression levels. All experiments exert similar contributions to PC1 as indicated by the direction and length of the arrows. Bio = biofilm; Pl = planktonic; aero = aerobic; anaero = anaerobic. and biofilm growth conditions in P. aeruginosa. The authors compared five studies testing the global gene expression under the two growth conditions and pointed to substantial differences among studies. They suggested that culturing conditions of the tested biofilms have a major impact on the emerging transcriptome profiles (Dötsch et al. 2012). This may provide an explanation for the differences between our results and Beloin et al. (2003) as we used a custom-made flow-cell system (Majeed et al. 2014), while they used a continuous-culture system in which the biofilm is formed in a microfermenter. We thus suggest that further research is required to standardize biofilm-culturing conditions. Planktonic cultures Our results show that 80 genes out of 4321 significantly differ in expression and were either up- or downregulated in the stationary-phase planktonic E. coli cultures grown under aerobic vs anaerobic conditions (Fig. 4a; Table S4, Supporting Information). Among the 52 E. coli genes downregulated under anaerobic conditions, 23 were of unknown or putative function, including 4 that were identified as Salmonella orthologs regulated by rpoS (Keseler et al. 2013). In addition, at least seven of the known downregulated genes were identified as rpoS-regulated (Table S4). A total of 28 genes were upregulated in anaerobic stationaryphase cultures (Fig. 4a; Table S4) including stress-related genes involving cold shock (cspAG) and osmoprotectants (proVW) (Keseler et al. 2013). In addition, genes encoding a multisubunit complex responsible for anaerobic oxidation of formic acid to carbon dioxide and molecular hydrogen (hycABCDEFGHI and fdhF) (Sanchez-Torres et al. 2009) were upregulated. Our results correspond with other research done on E. coli (Myers et al. 2013) where the expression of the hyc gene family was also increased when the strains were cultured to exponential phase under anaerobic conditions. Biofilm cultures The RNA-seq analyses of aerobic and anaerobic mature biofilms showed very different gene expression profiles compared to planktonic stationary-phase cultures (Figs 2 and 3) and exponential cultures (Myers et al. 2013). In fact, the majority of the differentially expressed genes were downregulated in the anaerobic biofilms, with only three upregulated genes (Fig. 4b; Tables 2 and S5, Supporting Information). Only one of the three upregulated genes was characterized safA, while the other two yobI and yqgC are of an unknown function (Keseler et al. 2013). The analysis of the downregulated genes in anaerobic compared to aerobic E. coli biofilm showed that most are Bayramoglu et al. 5 Figure 4. Overview of differential gene expression in (A) planktonic and (B) biofilm E. coli under oxic compared to anoxic conditions. Genes displaying >2.5-fold or <2.5-fold change ratios were significantly altered (q < 0.01). Each data points correspond to b-numbers and plotted in chronological order. Genes displaying >10-fold or <10-fold change are at the outer limits that enable the data to be viewed at a reasonable scale. essential to basic cell functions thereby the downregulation of these genes may attenuate key cell processes (Table 2) cutting down on the cells’ energy costs. This included the expression of genes encoding major fundamental components of rRNA function such as 30S (rpsBCDEFGHIJKLMNOPQRSTU) and 50S (rplABCDEFGIJKLMNOPQRSTUVY and rpmABCDEFGHIJ) ribosomal subunit proteins (Keseler et al. 2013). Similarly, some tRNA-related genes were downregulated including those for methionine (metTWZ), isoleucine (ileTUV), lysine (lysT), leucine (leuQ), threonine (thrU), valine (valTUXYZ), argenine (argX), glutamate (gltUV), serine (serS) and alanine (alaW) as well as essential tRNA-encoding factors (trmD, def and frr) (Table 2). Additionally, genes encoding key enzymes in the biosynthesis of fatty acids (accCD and fabABFGI) were downregulated together with genes involved in ATP synthase (atpABCDEFGH) and lipid biosynthesis (lpxACD, skp and fabZ). The expressions of genes that regulate a myriad of other functions were downregulated, such as motility (flxA, flgBCKLM, fliAC, motA), cell division (ftsJ, sulA, zapA and zipA), DNA recombination and repair (dinI, ihfA, ihfB, recA, yebG, ssb and lexA) and cold-shock-related genes (cspABCDG) (Keseler et al. 2013). Among the downregulated genes we detected major transcription factors (Table 2) regulating genes involved in numerous processes and responses (Table S6, Supporting Information). In fact, over 10% of regulators in the E. coli genome (Fis, H-NS and Lrp) were repressed. Part of this group included transcription factor genes that play a major role in biofilm formation (Fis, Crp and IscR) along with factor regulating genes involved in the transition from aerobic to anaerobic growth (Fnr and ArcA) (Keseler et al. 2013). In addition, bacterial cell surface components playing and important role in biofilm development and maturation including type 1 fimbriae and flagellin [fimA (Rodrigues and Elimelech 2009) and fliC (Zhou et al. 2014), respectively] were significantly downregulated (Table S2). The inhibition of these essential factors may have imposed the attenuated growth and the patchy morphology of the anaerobic biofilms (Fig. 1). A large number (∼5%) of small RNA (sRNA) and related genes were repressed (Table S6) in the anaerobic E. coli biofilms. Prominent among them was hfq, an RNA chaperone that facilitates sRNA base pairing with their target, has been found necessary for the translation of RpoS (Gottesman and Storz 2015) and is also a master regulator of the general stress responses in E. coli (Chambers and Sauer 2013). Moreover, we detected downregulated sRNAs (including srsD, ryeA and psrD) that are associated to hfq and by proxy to rpoS (Gottesman and Storz 2015). The downregulation of the hfq proteins could be linked with the overall repression of key regulatory mechanisms and key cell functions (Fig. 4b; Table S6) as well as the slower growth rate (Vytvytska et al. 1998) of the anaerobic biofilms. CONCLUSION Facultative anaerobes, like Escherichia coli, consume oxygen and modify their hosts’ environments yet, efficient oxygen scavenging may impose anaerobic respiration conditions on E. coli cells (Morris and Schmidt 2013). In this study, we demonstrated that under anaerobic conditions E. coli biofilm attenuates essential processes but could still persist as a biofilm (Fig. 1), whereas planktonic cells were less affected by the lack of oxygen. We report that unlike the sessile cultures only a few pathways were affected by oxygen depletion in the planktonic cultures (Figs 3 and 4). In an attempt to alleviate the oxygen-depletion stress in the anaerobic biofilm, E. coli cells minimize activity and mitigate the expression of essential genes (Table 2). The effect of oxygen availability was likewise tested in Pseudomonas aeruginosa, where cells were grown under aerobic, microaerobic and anaerobic conditions. It was demonstrated that the P. aeruginosa cells favored microaerobic conditions, while anaerobic conditions inhibited planktonic growth and biofilm development thus significantly altering gene expression profiles (Alvarez-Ortega and Harwood 2007). In contrast to previous reports, the results presented here may suggest that E. coli cells are constrained by anaerobic conditions. Perhaps E. coli cells, like P. aeruginosa, could better tolerate microaerobic over anaerobic conditions that might better serve the enterics’ main role in the mammalian colon, i.e. removal of oxygen traces and providing an oxygen-free environment to the dominant anaerobic populations (Jones et al. 2011). Hence, we aim to examine E. coli biofilm under microaerobic conditions exploring the means by which oxygen-limited (but probably not deprived) E. coli cells successfully populate the colon. 6 FEMS Microbiology Letters, 2017, Vol. 364, No. 3 Table 2. Key differentially expressed genes in anaerobic E. coli biofilm (for the complete genes list, please see Table S4). Function Amino acids Asparagine Aspartate Glutamine Histidine Isoleucine and valine Lysine Phenylalanine Serine Threonine Tyrosine Carbon metabolism Asparagine Carbohydrate transport Carbon catabolite repression Fructose Glucose Glycine Pentose Pyruvate Succinate Elongation factor Fatty acids/lipids Biotin biosynthesis Key protein Synthesis of unsaturated fatty acids Lipid biosynthesis Membrane structure Name asnA & B pyrB glnB glnA gltD hisL ilvC ilvL dapA dapD pheL aroG serA serC glyA thrL aroF Product Asparagine synthetase A & B Aspartate carbamoyltransferase, catalytic subunit Regulatory protein P-II for glutamine synthetase Glutamine synthetase Glutamate synthase, 4Fe-4S protein, small subunit His operon leader peptide Ketol-acid reductoisomerase, NAD(P) binding ilvG operon leader peptide Dihydrodipicolinate synthase 2,3,4,5-Tetrahydropyridine-2-carboxylate N-succinyltransferase pheA gene leader peptide 3-Deoxy-D-arabino-heptulosonate-7-phosphate synthase, phenylalanine repressible D-3-Phosphoglycerate dehydrogenase 3-Phosphoserine/phosphohydroxythreonine aminotransferase Serine hydroxymethyltransferase thr operon leader peptide 3-Deoxy-D-arabino-heptulosonate-7-phosphate synthase, tyrosine-repressible Fold difference ∗ − 2.69 − 2.79 − 5.17 − 2.54 − 3.55 − 3.04 − 6.57 − 2.66 − 4.1 − 4.4 − 2.69 − 4.21 − 2.62 − 4.47 − 4.39 − 4.1 − 2.66 − 4.02 gpmA Isoaspartyl peptidase Phosphohistidinoprotein-hexose phosphotransferase component of PTS system (Hpr) Phosphoglyceromutase 1 fbaA crr eno tpiA gapA gcvH talB aceE aceF cptB Fructose-bisphosphate aldolase, class II Glucose-specific enzyme IIA component of PTS Enolase Triosephosphate isomerase Glyceraldehyde-3-phosphate dehydrogenase A Glycine cleavage complex lipoylprotein Transaldolase B Pyruvate dehydrogenase, decarboxylase component E1, thiamin-binding Pyruvate dehydrogenase, dihydrolipoyltransacetylase component E2 Antitoxin of CptAB toxin-antitoxin pair − 4.13 − 3.31 − 4.08 − 2.72 − 7.04 − 3.31 − 4.35 − 5.4 − 3.92 − 2.66 efp ihfA & B tsf tufa & B tufB Polyproline-specific translation elongation factor EF-P Integration host factor (IHF), DNA-binding protein, subunits α & β Potein chain elongation factor EF-Ts Protein chain elongation factor EF-Tu Protein chain elongation factor EF-Tu (duplicate of) − 2.57 accC accD acpP fabA fabB & F fabG fabI fabZ lpxA lpxC lpxD cvpA gnsB lpp pal slyB spr mipA safA Acetyl-CoA carboxylase, biotin carboxylase subunit Acetyl-CoA carboxylase, beta (carboxyltransferase) subunit Acyl carrier protein (ACP) Beta-hydroxydecanoyl thioester dehydrase 3-Oxoacyl-[acyl-carrier-protein] synthase I, II 3-Oxoacyl-[acyl-carrier-protein] reductase Enoyl-[acyl-carrier-protein] reductase, NADH-dependent (3R)-Hydroxymyristol acyl carrier protein dehydratase UDP-N-acetylglucosamine acetyltransferase UDP-3-O-acyl N-acetylglucosamine deacetylase UDP-3-O-(3-hydroxymyristoyl)-glucosamine N-acyltransferase Membrane protein required for colicin V production Qin prophage; multicopy suppressor of secG(Cs) and fabA6(Ts) Murein lipoprotein Peptidoglycan-associated outer membrane lipoprotein Outer membrane lipoprotein Mutational suppressor of prc thermosensitivity, outer membrane lipoprotein Scaffolding protein for murein synthesizing machinery Two-component system connector membrane protein, EvgSA to PhoQP − 2.55 − 2.68 − 9.61 − 4.34 iaaA ptsH − 4.71 ∗ − 5.73 ∗ ∗ − 4.5 − 2.69 − 2.86 − 2.61 − 4.01 − 2.53 − 2.86 − 2.67 − 8.8 − 5.47 − 3.09 − 3.12 − 3.38 4.38 7 Bayramoglu et al. Table 2. (Continued). Function Membranebound ATP synthase Motility Nucleic acids Purine biosynthesis Component of ribosome 16S subunit Component of ribosome 23S subunit Components of ribosome 30S subunit Components of ribosome 50S subunit SOS response atp B, E, F Fold difference F1 sector, subunit α, ε, β, γ , δ, respectively ∗ F0 sector, subunit a, c, b, respectively ∗ Biofilm-dependent modulation protein Pleiotropic regulatory protein for carbon source metabolism Major type 1 subunit fimbrin (pilin) Flagellar component of cell-proximal portion of basal-body rod Flagellar hook-filament junction protein 1 RNA polymerase, sigma 28 (sigma F) factor Flagellar filament structural protein (flagellin) Proton conductor component of flagella motor Conserved protein with nucleoside triphosphate hydrolase domain − 2.61 − 4.4 − 22.52 ∗ ∗ − 3.79 − 10.48 − 3.01 − 4.13 − 3.2 −3 − 3.26 − 2.93 cmk pyrC pyrI rimM 16S rRNA processing protein − 4.71 rlmE 23S rRNA U2552 2’-O-ribose methyltransferase, SAM-dependent − 2.57 rpsB, C, D, E, F, G, H, I, J, K, L, M, N, O, P, Q, R, S, T, U rplA, B, C, D, E, F, I, J, K, L, M, N, O, P, Q, R, S, T, U, V, W, X, Y rpmA, B, C, D, E, F, G, H, I, J rpoA rpoZ frr hpf ssb sulA tisB yebG csrB glmZ psrD psrO ryeA ryeB ryfA sibC sibE tff Global sRNA chaperone Product Adenylate kinase Guanylate kinase Phosphoribosylpyrophosphate synthase Carbamoyl phosphate synthetase small subunit, glutamine amidotransferase Cytidylate kinase Dihydro-orotase Aspartate carbamoyltransferase, regulatory subunit adk gmk prs carA dinI lexA recA sRNA Transcriptional regulator atp A, C, D, G, H bdm csrA fimA flgB & C flgK & L fliA fliC motA phoH Pyrimidine biosynthesis rRNA Name hfq − 2.72 − 3.56 − 5.89 Proteins S2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21 respectively ∗ Proteins L1, 2, 3, 4, 5, 6, 9, 10, 11, 7/12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25 respectively ∗ Proteins L27, 28, 29, 30, 31, 32, 33, 34, 35, 36 respectively ∗ RNA polymerase, alpha subunit RNA polymerase, omega subunit Ribosome recycling factor Ribosome hibernation promoting factor HPF; stabilizes 70S dimers (100S) − 5.96 − 4.194 − 3.74 − 3.11 DNA damage-inducible protein I DNA-binding transcriptional repressor of SOS regulon DNA strand exchange and recombination protein with protease and nuclease activity Single-stranded DNA-binding protein SOS cell division inhibitor Toxic membrane persister formation peptide, LexA-regulated Conserved protein regulated by LexA Regulating carbon source metabolism Posttranscriptional regulation of glmS Unknown Unknown Unknown Unknown Unknown Unknown Unknown Unknown − 6.35 − 4.15 − 6.49 HF-I, host factor for RNA phage Q beta replication − 6.78 − 2.67 − 4.83 − 2.96 − 4.89 − 6.75 − 6.4 − 4.91 − 2.98 − 4.58 − 5.25 − 2.67 − 3.13 − 2.72 − 9.01 8 FEMS Microbiology Letters, 2017, Vol. 364, No. 3 Table 2. (Continued). Function Stress response Cold shock Heat shock High pH Osmotic stress Oxidative stress Starvation and low pH Temperature Universal stress response Sulfate assimilation Transport tRNA Alanine Arginine Aspargine and phenylalanine Glutamic acid Isoleucine Leucine Lysine Methionine Threonine Name Product Fold difference − 2.88 − 10.84 − 5.9 − 3.31 − 5.33 − 3.06 − 3.49 − 3.25 − 2.55 ycgZ cspA cspB cspD cspG cspI grpE cpxP osmB RcsB connector protein for regulation of biofilm and acid-resistance RNA chaperone and anti-terminator, cold-inducible Cold shock protein Inhibitor of DNA replication, cold shock protein homolog Cold shock protein homolog, cold-inducible Cold shock protein Heat shock protein Inhibitor of the cpx response; periplasmic adaptor protein Lipoprotein bcp icd lpd tpx trxA can sspA Peroxiredoxin; thiol peroxidase, thioredoxin-dependent e14 prophage; isocitrate dehydrogenase, specific for NADP+ Lipoamide dehydrogenase, E3 component is part of three enzyme complexes Lipid hydroperoxide peroxidase Thioredoxin 1 Carbonic anhydrase 2 Stringent starvation protein A cspC sodA uspA sodB groS cysA cysD cysH cysJ cysK cysN cysP cysW Stress protein, member of the CspA-family Superoxide dismutase, Mn Universal stress global response regulator Superoxide dismutase, Fe Cpn10 chaperonin GroES, small subunit of GroESL Sulfate/thiosulfate transporter subunit Sulfate adenylyltransferase, subunit 2 3’-Phosphoadenosine 5’-phosphosulfate reductase Sulfite reductase, alpha subunit, flavoprotein Cysteine synthase A, O-acetylserine sulfhydrolase A subunit Sulfate adenylyltransferase, subunit 1 Thiosulfate-binding protein Sulfate/thiosulfate ABC transporter subunit − 6.09 − 3.7 − 5.59 − 3.54 − 2.9 − 4.8 − 6.2 − 2.81 − 3.56 − 15.47 − 4.22 − 5.11 − 3.52 AcrAB-TolC efflux pump accessory protein, membrane-associated Cytochrome o ubiquinol oxidase subunit II Membrane spanning protein in TonB-ExbB-ExbD complex Cystine transporter subunit Glutamine transporter subunit Maltose transporter subunit Outer membrane porin protein C Outer membrane porin 1a (Ia;b;F) Outer membrane protein X Glycine betaine transporter subunit Sulfate transporter subunit Protein export chaperone Preprotein translocase membrane subunit Periplasmic chaperone TatABCE protein translocation system subunit Periplasmic protein Nucleoside channel, receptor of phage T6 and colicin K Ala tRNA Arg tRNA tRNA m(1)G37 methyltransferase, SAM-dependent − 3.84 − 3.159 − 2.679 − 3.19 − 4.084 − 2.835 − 4.72 − 12.51 − 7.32 − 3.55 − 2.88 − 4.235 acrZ cyoA exbB fliY glnH malE ompC ompF ompX proV sbp secB secE & G skp tatA tolB tsx alaW argX trmD gltU & V ileT, U, V leuQ & W lysT metT, W, Z thrU Glu tRNA Ile tRNA Leu tRNA Lys tRNA Met tRNA Thr tRNA − 2.51 − 5.23 − 4.71 − 9.22 − 4.49 − 3.85 − 3.54 ∗ − 6.4 − 2.63 − 2.606 − 2.56 − 2.917 − 4.726 − 5.1 ∗ ∗ ∗ − 5.44 ∗ − 4.16 9 Bayramoglu et al. Table 2. (Continued). Function tRNA modification tRNA modification Valine Name Product Fold difference dusB tRNA-dihydrouridine synthase B − 3.363 miaA Delta(2)-isopentenylpyrophosphate tRNA-adenosine transferase − 3.46 valT, U, X, Y, Z ∗ Val tRNA Fold difference values indicate downregulation in anaerobic compared to aerobic conditions. ∗ Please refer to Table S4 for the fold difference values. SUPPLEMENTARY DATA Supplementary data are available at FEMSLE online. ACKNOWLEDGEMENTS We are grateful to Moshe Herzberg for his help with the EPS analyses. We also thank Ahuva Vonshak and Lusine Ghazaryan for their help in performing the experiments. We greatly appreciate the insightful comments of Maya Benami and Adam Stovicek to the manuscript. Conflict of interest. None declared. REFERENCES Alvarez-Ortega C, Harwood CS. Responses of Pseudomonas aeruginosa to low oxygen indicate that growth in the cystic fibrosis lung is by aerobic respiration. Mol Microbiol 2007;65:153–65. Anders S, Huber W. Differential expression analysis for sequence count data. Genome Biol 2010;11:R106. Beloin C, Roux A, Ghigo JM. Escherichia coli biofilms. Curr Top Microbiol Immunol 2008;322:249–89. Beloin C, Valle J, Latour-Lambert P et al. Global impact of mature biofilm lifestyle on Escherichia coli K-12 gene expression: Mature biofilm formation in E. coli. Mol Microbiol 2003;51:659– 74. Bjergbæk LA, Haagensen JAJ, Reisner A et al. Effect of oxygen and growth medium on in vitro biofilm formation by Escherichia coli. Biofilms 2006;3:1–10. Brownstein Z, Abu-Rayyan A, Karfunkel-Doron D et al. Novel myosin mutations for hereditary hearing loss revealed by targeted genomic capture and massively parallel sequencing. Eur J Hum Genet 2014;22:768–75. Cabellos-Avelar T, Souza V, Membrillo-Hernández J. Spent media from cultures of environmental isolates of Escherichia coli can suppress the deficiency of biofilm formation under anoxic conditions of laboratory E. coli strains: Biofilm formation in E. coli under anaerobic conditions. FEMS Microbiol Ecol 2006;58:414–24. Chambers JR, Sauer K. Small RNAs and their role in biofilm formation. Trends Microbiol 2013;21:39–49. Colón-González M, Méndez-Ortiz MM, Membrillo-Hernández J. Anaerobic growth does not support biofilm formation in Escherichia coli K-12. Res Microbiol 2004;155:514–21. Crusz SA, Popat R, Rybtke MT et al. Bursting the bubble on bacterial biofilms: a flow cell methodology. Biofouling 2012;28:835– 42. Dötsch A, Eckweiler D, Schniederjans M et al. The Pseudomonas aeruginosa transcriptome in planktonic cultures and static biofilms using RNA sequencing. PLoS One 2012;7:e31092. Edgar R. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res 2002;30:207–10. Fitzgerald DM, Bonocora RP, Wade JT. Comprehensive mapping of the Escherichia coli flagellar regulatory network. PLoS Genet 2014;10:e1004649. Gentleman RC, Carey VJ, Bates DM et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol 2004;5:R80. Gottesman S, Storz G. RNA reflections: converging on Hfq. RNA 2015;21:511–2. Hancock V, Klemm P. Global gene expression profiling of asymptomatic bacteriuria Escherichia coli during biofilm growth in human urine. Infect Immun 2007;75:966–76. Ihaka R, Gentleman R. R: a language for data analysis and graphics. J Comput Graph Stat 1996;5:299–314. Ionescu M, Belkin S. Overproduction of exopolysaccharides by an Escherichia coli K-12 rpoS mutant in response to osmotic stress. Appl Environ Microb 2009;75:483–92. Jones SA, Gibson T, Maltby RC et al. Anaerobic respiration of Escherichia coli in the mouse intestine. Infect Immun 2011;79:4218–26. Karatan E, Watnick P. Signals, regulatory networks, and materials that build and break bacterial biofilms. Microbiol Mol Biol R 2009;73:310–47. Keseler IM, Mackie A, Peralta-Gil M et al. EcoCyc: fusing model organism databases with systems biology. Nucleic Acids Res 2013;41:D605–12. Landini P, Zehnder AJB. The global regulatory hns gene negatively affects adhesion to solid surfaces by anaerobically grown Escherichia coli by modulating expression of flagellar genes and lipopolysaccharide production. J Bacteriol 2002;184:1522–9. McClure R, Balasubramanian D, Sun Y et al. Computational analysis of bacterial RNA-Seq data. Nucleic Acids Res 2013; 41:e140. Macfarlane S, Dillon JF. Microbial biofilms in the human gastrointestinal tract. J Appl Microbiol 2007;102:1187–96. Majeed H, Ghazaryan L, Herzberg M et al. Bacteriocin expression in sessile and planktonic populations of Escherichia coli. J Antibiot (Tokyo) 2014, DOI: 10.1038/ja.2014.84. Malone JH, Oliver B. Microarrays, deep sequencing and the true measure of the transcriptome. BMC Biol 2011;9:34. Morris RL, Schmidt TM. Shallow breathing: bacterial life at low O2. Nat Rev Microbiol 2013;11:205–12. Myers KS, Yan H, Ong IM et al. Genome-scale analysis of Escherichia coli FNR reveals complex features of transcription factor binding. PLoS Genet 2013;9: e1003565. 10 FEMS Microbiology Letters, 2017, Vol. 364, No. 3 Partridge JD, Scott C, Tang Y et al. Escherichia coli transcriptome dynamics during the transition from anaerobic to aerobic conditions. J Biol Chem 2006;281: 27806–15. Ritchie ME, Phipson B, Wu D et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 2015;43:e47. Rodrigues DF, Elimelech M. Role of type 1 fimbriae and mannose in the development of Escherichia coli K12 biofilm: from initial cell adhesion to biofilm formation. Biofouling 2009;25: 401–11. Sanchez-Torres V, Maeda T, Wood TK. Protein Engineering of the Transcriptional Activator FhlA To Enhance Hydrogen Production in Escherichia coli. Appl Environ Microbiol 2009; 75:5639–46. Schembri MA, Kjaergaard K, Klemm P. Global gene expression in Escherichia coli biofilms: Gene expression in E. coli biofilms. Mol Microbiol 2003;48:253–67. Serres MH, Riley M. MultiFun, a multifunctional classification scheme for Escherichia coli K-12 gene products. Microb Comp Genomics 2000;5:205–22. Shao ZH, Ren SX, Liu XQ et al. A preliminary study of the mechanism of nitrate-stimulated remarkable increase of rifamycin production in Amycolatopsis mediterranei U32 by RNA-seq. Microb Cell Fact 2015;14:75. Stacklies W, Redestig H, Scholz M et al. pcaMethods–a bioconductor package providing PCA methods for incomplete data. Bioinformatics 2007;23:1164–7. Sweity A, Ying W, Ali-Shtayeh MS et al. Relation between EPS adherence, viscoelastic properties, and MBR operation: Biofouling study with QCM-D. Water Res 2011;45:6430–40. Trotter EW, Rolfe MD, Hounslow AM et al. Reprogramming of Escherichia coli K-12 metabolism during the initial phase of transition from an anaerobic to a micro-aerobic environment. PLoS One 2011;6:e25501. Vytvytska O, Jakobsen JS, Balcunaite G et al. Host factor I, Hfq, binds to Escherichia coli ompA mRNA in a growth ratedependent fashion and regulates its stability. P Natl Acad Sci USA 1998;95:14118–23. Waite RD, Paccanaro A, Papakonstantinopoulou A et al. Clustering of Pseudomonas aeruginosa transcriptomes from planktonic cultures, developing and mature biofilms reveals distinct expression profiles. BMC Genomics 2006;7:162. Wood TK. Insights on Escherichia coli biofilm formation and inhibition from whole-transcriptome profiling. Environ Microbiol 2009;11:1–15. Zhou M, Guo Z, Yang Y et al. Flagellin and F4 fimbriae have opposite effects on biofilm formation and quorum sensing in F4ac+ enterotoxigenic Escherichia coli. Vet Microbiol 2014;168:148–53.