* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Lecture 2 Turunen 14.9. - MyCourses
Nutriepigenomics wikipedia , lookup
Nucleic acid tertiary structure wikipedia , lookup
RNA silencing wikipedia , lookup
United Kingdom National DNA Database wikipedia , lookup
Genealogical DNA test wikipedia , lookup
Gel electrophoresis of nucleic acids wikipedia , lookup
Messenger RNA wikipedia , lookup
Genomic library wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Genetic engineering wikipedia , lookup
DNA damage theory of aging wikipedia , lookup
DNA vaccination wikipedia , lookup
History of RNA biology wikipedia , lookup
Designer baby wikipedia , lookup
DNA polymerase wikipedia , lookup
Epigenomics wikipedia , lookup
Molecular cloning wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Nucleic acid double helix wikipedia , lookup
Non-coding RNA wikipedia , lookup
Epitranscriptome wikipedia , lookup
DNA supercoil wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Non-coding DNA wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Point mutation wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Microevolution wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
History of genetic engineering wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Deoxyribozyme wikipedia , lookup
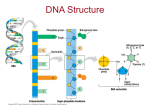
8/29/2016 PowerPoint® Lecture Presentations prepared by Mindy Miller-Kittrell, North Carolina State University Modified by Ossi Turunen, Aalto University CHAPTER 7 Microbial Genetics © 2015 Pearson Education, Ltd. The Structure and Replication of Genomes • Genetics • Study of inheritance and inheritable traits as expressed in an organism's genetic material • Genome • The entire genetic complement of an organism • Includes its genes and nucleotide sequences © 2015 Pearson Education, Ltd. 1 8/29/2016 Figure 7.1 The structure of nucleic acids. Hydrogen bond Sugar Thymine (T) nucleoside Adenine (A) nucleoside A–T base pair (DNA) 5′ end 3′ end Uracil (U) nucleoside Adenine (A) nucleoside A–U base pair (RNA) Cytosine (G) nucleoside Guanine (G) nucleoside G–C base pair (DNA and RNA) 5′ end 3′ end A T G C A T G Guanine 3′ end 5′ end Double-stranded DNA Cytosine Adenine 3′ end Thymine 5 end′ Thymine nucleoside Thymine nucleotide © 2015 Pearson Education, Ltd. The Structure and Replication of Genomes • The Structure of Prokaryotic Genomes • Prokaryotic chromosomes • Main portion of DNA, along with associated proteins and RNA • Prokaryotic cells are haploid (single chromosome copy) • Typical chromosome is circular molecule of DNA in nucleoid © 2015 Pearson Education, Ltd. 2 8/29/2016 Figure 7.2 Bacterial genome. Nucleoid Bacterium Chromosome Plasmid © 2015 Pearson Education, Ltd. The Structure and Replication of Genomes • The Structure of Prokaryotic Genomes • Plasmids • Small molecules of DNA that replicate independently • Not essential for normal metabolism, growth, or reproduction • Can confer survival advantages • Many types of plasmids • Fertility factors • Resistance factors • Bacteriocin factors • Virulence plasmids © 2015 Pearson Education, Ltd. 3 8/29/2016 The Structure and Replication of Genomes • The Structure of Eukaryotic Genomes • Nuclear chromosomes • Typically have more than one chromosome per cell • Chromosomes are linear and sequestered within nucleus • Eukaryotic cells are often diploid (two chromosome copies) © 2015 Pearson Education, Ltd. Figure 7.3 Eukaryotic nuclear chromosomal packaging. 10 nm Nucleosome Active (loosely packed) Histones Linker DNA Inactive (tightly packed) DNA 10 nm 30 nm Nucleosomes Chromatin fiber 700 nm Euchromatin and heterochromatin 1400 nm Highly condensed, duplicated chromosome of dividing nucleus © 2015 Pearson Education, Ltd. 4 8/29/2016 The Structure and Replication of Genomes • The Structure of Eukaryotic Genomes • Extranuclear DNA of eukaryotes • DNA molecules of mitochondria and chloroplasts • Resemble chromosomes of prokaryotes • Code only for about 5% of RNA and proteins • Some fungi, algae, and protozoa carry plasmids © 2015 Pearson Education, Ltd. © 2015 Pearson Education, Ltd. 5 8/29/2016 The Structure and Replication of Genomes • DNA Replication • Key to replication is complementary structure of the two strands • Replication is semiconservative • New DNA composed of one original and one daughter strand • Anabolic polymerization process that requires monomers and energy • Triphosphate deoxyribonucleotides serve both functions © 2015 Pearson Education, Ltd. DNA Replication © 2015 Pearson Education, Ltd. 6 8/29/2016 Figure 7.4 Semiconservative model of DNA replication. Original DNA First replication Second replication Original strand New strands © 2015 Pearson Education, Ltd. Figure 7.5 The dual role of triphosphate deoxyribonucleotides as building blocks and energy sources in DNA synthesis. Guanosine triphosphate deoxyribonucleotide (dGTP) Guanine nucleotide (dGMP) High-energy bond Guanine base Deoxyribose Guanosine (nucleoside) OH Diphosphate released, energy used for synthesis Existing DNA strand Triphosphate nucleotide Longer DNA strand © 2015 Pearson Education, Ltd. 7 8/29/2016 The Structure and Replication of Genomes • DNA Replication • Initial processes in bacterial DNA replication • Replication begins at the origin • DNA polymerase replicates DNA only 5′ to 3′ • Because strands are antiparallel, new strands are synthesized differently • Leading strand synthesized continuously • Lagging strand synthesized discontinuously © 2015 Pearson Education, Ltd. Figure 7.6a DNA replication. Chromosomal proteins (histones in eukaryotes and archaea) removed DNA polymerase III 3′ Replication fork 5′ DNA helicase Stabilizing proteins Initial processes © 2015 Pearson Education, Ltd. 8 8/29/2016 Figure 7.6b-c DNA replication. Primase 3 1 3′ Replication fork 5′ 2 Leading strand P+P Triphosphate nucleotide RNA primer Synthesis of leading strand Replication fork Triphosphate nucleotide RNA primer Okazaki fragment 6 7 Lagging strand 3′ 5′ 8 Primase 9 10 DNA ligase DNA polymerase III DNA polymerase I Synthesis of lagging strand © 2015 Pearson Education, Ltd. The Structure and Replication of Genomes • DNA Replication • Other characteristics of bacterial DNA replication • Bidirectional • Gyrases and topoisomerases remove supercoils in DNA • DNA is methylated • Control of genetic expression • Initiation of DNA replication • Protection against viral infection • Repair of DNA © 2015 Pearson Education, Ltd. 9 8/29/2016 Figure 7.7 The bidirectionality of DNA replication in prokaryotes. Parental strand Origin Replication forks Daughter strand Replication proceeds in both directions Termination of replication © 2015 Pearson Education, Ltd. The Structure and Replication of Genomes • DNA Replication • Replication of eukaryotic DNA • Similar to bacterial replication • Some differences • Uses four DNA polymerases • Thousands of replication origins • Shorter Okazaki fragments • Plant and animal cells methylate only cytosine bases © 2015 Pearson Education, Ltd. 10 8/29/2016 The Structure and Replication of Genomes • Tell Me Why • DNA replication requires a large amount of energy, yet none of a cell's ATP energy supply is used. Why isn't it? © 2015 Pearson Education, Ltd. Gene Function • The Relationship Between Genotype and Phenotype • Genotype • Set of genes in the genome • Phenotype • Physical features and functional traits of the organism • Genotype determines phenotype • Not all genes are active at all times © 2015 Pearson Education, Ltd. 11 8/29/2016 Gene Function • The Transfer of Genetic Information • Transcription • Information in DNA is copied as RNA • Translation • Polypeptides are synthesized from RNA • Central dogma of genetics • DNA is transcribed to RNA • RNA is translated to form polypeptides © 2015 Pearson Education, Ltd. Figure 7.8 The central dogma of genetics. © 2015 Pearson Education, Ltd. 12 8/29/2016 Gene Function • The Events in Transcription • Five types of RNA transcribed from DNA • RNA primers • mRNA • rRNA • tRNA • Regulatory RNA • Occur in nucleoid of prokaryotes • Three steps • Initiation • Elongation • Termination © 2015 Pearson Education, Ltd. Figure 7.9a The events in the transcription of RNA in prokaryotes. 1a RNA polymerase attaches nonspecifically to DNA and RNA polymerase travels down its length until 5′ it recognizes a promoter 3′ sequence. Sigma factor Sigma factor Promoter enhances promoter recognition in bacteria. Attachment of RNA polymerase 1b Upon recognition of the 3′ 5′ DNA Terminator "Bubble" promoter, RNA polymerase 5′ unzips the DNA molecule beginning at the promoter. 3′ 3′ 5′ Unzipping of DNA, movement of RNA polymerase Template DNA strand Initiation of transcription © 2015 Pearson Education, Ltd. 13 8/29/2016 Figure 7.9b The events in the transcription of RNA in prokaryotes. "Bubble" 2 Triphosphate ribonucleotides align with their DNA 5′ complements and RNA 3′ polymerase links them together, synthesizing RNA. Growing RNA molecule No primer is needed. The (transcript) triphosphate ribonucleotides also provide the energy required for RNA synthesis. 3′ 3′ 5′ 5′ 3′ 5′ G C 5′ 3′ Elongation of the RNA transcript Template DNA strand © 2015 Pearson Education, Ltd. Figure 7.10 Concurrent RNA transcription. RNA polymerases Promoter 5′ 3′ 5′ 5′ 3′ 3′ 3′ 3′ 3′ 5′ Sigma factor 3′ 3′ 5′ Template DNA strand RNA 5′ 5′ 5′ © 2015 Pearson Education, Ltd. 14 8/29/2016 Figure 7.9c The events in the transcription of RNA in prokaryotes. 3′ 5′ 5′ 3′ 5′ 3′ Terminator RNA transcript released 3a Self-termination: transcription of GC-rich terminator region produces a hairpin loop, which creates tension, loosening the grip of the polymerase on the DNA. 3b Rho-dependant termination: Rho pushes between polymerase and DNA. This causes release of polymerase, RNA transcript, and Rho. RNA polymerase Rho termination protein Rho protein moves along RNA GC-rich hairpin loop Terminator Terminator C 3′ Template strand Termination of transcription: release of RNA polymerase © 2015 Pearson Education, Ltd. Gene Function • The Events in Transcription • Transcriptional differences in eukaryotes • RNA transcription occurs in the nucleus • Transcription also occurs in mitochondria and chloroplasts • Three types of nuclear RNA polymerases • Numerous transcription factors • mRNA is processed before translation • Capping • Polyadenylation • Splicing © 2015 Pearson Education, Ltd. 15 8/29/2016 Figure 7.11 Processing eukaryotic mRNA. Exons (polypeptide coding regions) 5′ Template DNA strand 3′ Introns (noncoding regions) Transcription Exon 2 Exon 1 Exon 3 Pre-mRNA A AAAAAAAAAAAAA 5′ cap Intron 1 Intron 2 Intron 1 Intron 3 Poly-A tail Processing Spliceosomes 5′ Exon 1 Exon 2 A AAAAAAAAAAAAA 3′ mRNA splicing Exon 3 5′ A AAAAAAAAAAAAA mRNA (codes for 3′ one polypeptide) Nuclear envelope Nucleoplasm Nuclear pore Cytosol mRNA © 2015 Pearson Education, Ltd. Gene Function • Translation • Process in which ribosomes use genetic information of nucleotide sequences to synthesize polypeptides © 2015 Pearson Education, Ltd. 16 8/29/2016 Figure 7.12 The genetic code. © 2015 Pearson Education, Ltd. © 2015 Pearson Education, Ltd. 17 8/29/2016 Gene Function • Translation • Participants in translation • Messenger RNA • Transfer RNA • Ribosomes and ribosomal RNA © 2015 Pearson Education, Ltd. Figure 7.13 A single prokaryotic mRNA can code for several polypeptides. 3′ Gene 2 Gene 1 Promoter Gene 3 Terminator Transcription 5′ Start codon AUG UAA Ribosome binding site (RBS) Start codon AUG Stop RBS codon UAG Start codon AUG Stop RBS codon UAA 5′ Template DNA strand 3′ mRNA Untranslated Stop codon mRNA Translation Polypeptide 1 Polypeptide 2 Polypeptide 3 © 2015 Pearson Education, Ltd. 18 8/29/2016 Figure 7.14 Transfer RNA. OH Acceptor stem 3′ 5′ 5′ Hydrogen bonds 3′ tRNA icon Hairpin loops Anticodon Anticodon © 2015 Pearson Education, Ltd. Figure 7.15 Ribosomal structures. © 2015 Pearson Education, Ltd. 19 8/29/2016 Figure 7.16 Assembled ribosome and its tRNA-binding sites. Large subunit Large subunit mRNA Nucleotide bases 5′ E P A site sitesite tRNAbinding sites 3′ Small subunit Small subunit mRNA Prokaryotic ribosome (angled view) attached to mRNA Prokaryotic ribosome (schematic view) showing tRNA-binding sites © 2015 Pearson Education, Ltd. Gene Function • Translation • Events in translation • Three stages of translation • Initiation • Elongation • Termination • All stages require additional protein factors • Initiation and elongation require energy (GTP) © 2015 Pearson Education, Ltd. 20 8/29/2016 Figure 7.17 The initiation of translation in prokaryotes. 1 fMet 2 Initiator tRNA 3 GTP Large ribosomal subunit fMet fMet tRNA GDP + P fMet Anticodon mRNA 5′ E Start codon A U G U U U A C G P 3′ A U A C A U G U U U A C G P Small ribosomal subunit A U A C A U G U U U A C G P A Initiation complex © 2015 Pearson Education, Ltd. Figure 7.18 The elongation stage of translation. © 2015 Pearson Education, Ltd. 21 8/29/2016 Figure 7.19 In prokaryotes a polyribosome—one mRNA and many ribosomes and polypeptides. Polypeptides mRNA Ribosomes mRNA Ribosomes Polypeptides Direction of transcription © 2015 Pearson Education, Ltd. Gene Function • Translation • Events in translation • Termination • Release factors recognize stop codons • Modify ribosome to activate ribozymes • Ribosome dissociates into subunits • Polypeptides released at termination may function alone or together © 2015 Pearson Education, Ltd. 22 8/29/2016 Gene Function • Translation • Translation differences in eukaryotes • Initiation occurs when ribosomal subunit binds to 5′ guanine cap • First amino acid is methionine rather than f-methionine • Ribosomes can synthesize polypeptides into the cavity of the rough endoplasmic reticulum © 2015 Pearson Education, Ltd. © 2015 Pearson Education, Ltd. 23 8/29/2016 Protein Synthesis © 2015 Pearson Education, Ltd. Gene Function • Regulation of Genetic Expression • Most genes are expressed at all times (constitutive) • Other genes are transcribed and translated when cells need them (inductive) • Allows cell to conserve energy • Quorum sensing regulates production of some proteins • Quorum sensing is a system, in which stimuli and response correlate to population density • Detection of secreted quorum-sensing molecules can signal bacteria to synthesize a certain protein © 2015 Pearson Education, Ltd. 24 8/29/2016 Gene Function • Regulation of Genetic Expression • Regulation of polypeptide synthesis • Typically halts transcription • Can stop translation directly © 2015 Pearson Education, Ltd. Gene Function • Regulation of Genetic Expression • Nature of prokaryotic operons • An operon consists of a promoter and a series of genes • Controlled by a regulatory element called an operator • Typically polycistronic (code for several polypeptides) © 2015 Pearson Education, Ltd. 25 8/29/2016 Figure 7.20 An operon. Operon Regulatory gene Promoter Operator Structural genes 3′ 1 2 3 4 5′ Template DNA strand © 2015 Pearson Education, Ltd. Gene Function • Regulation of Genetic Expression • Induction and Repression in prokaryotic operons • Inducible operons must be activated by inducers • Lactose operon • Repressible operons are transcribed continually until deactivated by repressors • Tryptophan operon © 2015 Pearson Education, Ltd. 26 8/29/2016 Figure 7.21 The lac operon, an inducible operon. lac operon Promoter and regulatory gene Operator (blocked) Promoter 1 3′ 2 3 Lactose catabolism genes Continual transcription 5′ Template DNA strand RNA 2 polymerase cannot bind Repressor mRNA Continual translation 1 Repressor lac operon repressed RNA polymerase 1 3′ Repressor cannot bind Repressor mRNA Transcription proceeds 2 3 5′ Template DNA strand 4 mRNA for lactose catabolism 5′ Repressor 3 Inactivated repressor Inducer (allolactose from lactose) lac operon induced © 2015 Pearson Education, Ltd. Figure 7.22 CAP-cAMP enhances lac transcription. cAMP bound to CAP RNA polymerase Binding of RNA polymerase to the promoter is enhanced by the cAMPbound catabolite activator protein (CAP = cAMP receptor protein) Transcription proceeds CAP binding site Promoter Operator lac genes © 2015 Pearson Education, Ltd. 27 8/29/2016 Lac operon genes lacZ: β-galactosidase, cleaves lactose into glucose and galactose lacY: lactose permease in the cytoplasmic membrane to enable transport of lactose into the cell lacA: galactoside O-acetyltransferase, an enzyme that transfers an acetyl group from acetyl-CoA to β-galactosides © 2015 Pearson Education, Ltd. Figure 7.23 The trp operon, a repressible operon. trp operon with five genes Regulatory gene Promoter Operator 3′ 1 2 3 4 5 5′ Template DNA strand Transcription 3′ mRNA 5′ 5′ 3′ mRNA coding multiple polypeptides Enzymes of tryptophan biosynthetic pathway Inactive repressor trp operon active Trp Tryptophan Movement of RNA polymerase ceases 1 3′ Inactive repressor Trp 2 3 4 5 5′ Operator blocked Trp Trp Trp Trp Tryptophan (corepressor) trp operon repressed Activated repressor trp operon codes for the components for production of tryptophan © 2015 Pearson Education, Ltd. 28 8/29/2016 © 2015 Pearson Education, Ltd. Gene Function • Regulation of Genetic Expression • RNA molecules can control translation • Regulatory RNAs can regulate translation of polypeptides • microRNAs • About 22 ntd long pieces of RNA • Produced by eukaryotic cells • Bind regulatory proteins to form miRNA-induced silencing complex (miRISC) • Bind complementary mRNA and inhibit its translation • Regulates several cellular processes © 2015 Pearson Education, Ltd. 29 8/29/2016 How miRNAs function? - Wikipedia © 2015 Pearson Education, Ltd. Gene Function • Regulation of Genetic Expression • RNA molecules can control translation • Regulatory RNAs can regulate translation of polypeptides • Short interference RNA (siRNA) • RNA molecule complementary to a portion of mRNA, tRNA, or DNA • Binds RISC proteins to form siRISC • siRISC binds and cuts the target nucleic acid • Riboswitch • A regulatory segment of a mRNA that binds a small molecule and then changes shape to help regulate translation © 2015 Pearson Education, Ltd. 30 8/29/2016 Gene Function • Tell Me Why • In bacteria, polypeptide translation can begin even before mRNA transcription is complete. Why can't this happen in eukaryotes? © 2015 Pearson Education, Ltd. Mutations of Genes • Mutation • Change in the nucleotide base sequence of a genome • Rare event • Almost always deleterious • Rarely leads to a protein that improves ability of organism to survive © 2015 Pearson Education, Ltd. 31 8/29/2016 Mutations of Genes • Types of Mutations • Point mutations • One base pair is affected • Substitutions and frameshift mutations • Frameshift mutations • Nucleotide triplets after the mutation are displaced • Creates new sequence of codons • Insertions and deletions of several amino acids © 2015 Pearson Education, Ltd. Figure 7.24 The effects of the various types of point mutations. © 2015 Pearson Education, Ltd. 32 8/29/2016 © 2015 Pearson Education, Ltd. Mutations of Genes • Mutagens • Radiation • Ionizing radiation • Nonionizing radiation • Chemical mutagens • Nucleotide analogs • Disrupt DNA and RNA replication • Nucleotide-altering chemicals • Alter the structure of nucleotides • Result in base-pair substitutions and missense mutations • Frameshift mutagens • Result in nonsense mutations © 2015 Pearson Education, Ltd. 33 8/29/2016 Figure 7.25 A pyrimidine (in this case, thymine) dimer. Ultraviolet light Ultraviolet light induces the formation of covalent linkages by reactions localized on the C=C double bonds Thymine dimer GC T G T=T G GTA CGACAACCAT © 2015 Pearson Education, Ltd. Figure 7.26 The structure and effects of a nucleotide analog. © 2015 Pearson Education, Ltd. 34 8/29/2016 Figure 7.27 The action of a frameshift mutagen. © 2015 Pearson Education, Ltd. Mutations of Genes • Frequency of Mutation • Mutations are rare events • Otherwise, organisms could not effectively reproduce • About 1 of every 10 million genes contains an error • Mutagens increase the mutation rate by a factor of 10 to 1000 times • Many mutations stop transcription or code for nonfunctional proteins © 2015 Pearson Education, Ltd. 35 8/29/2016 Figure 7.28a-b DNA repair mechanisms. Visible light Thymine dimer G AC T=T A G AC T T A C T G A A T C T G A A T Light-activated repair enzyme Light repair Cut G AC T=T A C Repair enzyme C T G A A T G C DNA polymerase I and ligase repair GAC T TAC the gap C T G A A T G C T G A A T G G AC T=T A Dark repair © 2015 Pearson Education, Ltd. Figure 7.28c-d DNA repair mechanisms. Base excision repair enzymes remove incorrect nucleotide DNA polymerase I and ligase repair gap G GC T T AGC G T G GC T T A C G T G GC T T A TC G T C CG A A T AG C A C CG A A T AG C A C CG A A T AG C A Base-excision repair Mismatch repair enzyme removes incorrect segment Mutated DNA (incorrect nucleotide pair) DNA polymerase III correctly repairs the gap C T T A GC G T GC G T C C T A GC G T G G A T CG C A G G A T CG C A G G A T CG C A Mismatch repair © 2015 Pearson Education, Ltd. 36 8/29/2016 Mutations of Genes • Identifying Mutants, Mutagens, and Carcinogens • Mutants • Descendants of a cell that does not repair a mutation • Wild types • Cells normally found in nature • Methods to recognize mutants • Positive selection • Negative (indirect) selection • Ames test © 2015 Pearson Education, Ltd. Figure 7.29 Positive selection of mutants. Penicillinresistant cell Medium with penicillin (only penicillin-resistant cell grows into colony) Penicillinsensitive cells Medium without penicillin (both types of cells form colonies) Mutagen induces mutations Penicillinresistant mutants indistinguishable from nonmutants Medium with penicillin Medium without penicillin © 2015 Pearson Education, Ltd. 37 8/29/2016 Figure 7.30 The use of negative (indirect) selection to isolate a tryptophan auxotroph. 1 Inoculate bacteria onto complete medium containing tryptophan. Mutagen Bacterial suspension X Incubation Bacterial colonies 2 grow. A few may be tryptophan auxotrophs. Most are wild type. X 3 Stamp sterile velvet onto plate, picking up cells from each colony. Sterile velvet surface X Bacteria 4 Stamp replica plates with velvet. X X Complete medium containing tryptophan Medium lacking tryptophan Incubation 5 Identify auxotroph as colony growing on complete medium but not on lacking medium. X X All colonies grow. Tryptophan auxotroph cannot grow. 6 Inoculate auxotroph colony into complete medium. © 2015 Pearson Education, Ltd. Figure 7.31 The Ames test. Experimental tube Liver extract Control tube Suspected mutagen Liver extract Culture of his– Salmonella Medium lacking histidine Incubation © 2015 Pearson Education, Ltd. Colony of revertant (his+) Salmonella No growth 38 8/29/2016 Genetic Recombination and Transfer • Exchange of nucleotide sequences often occurs between homologous sequences • Recombinants • Cells with DNA molecules that contain new nucleotide sequences © 2015 Pearson Education, Ltd. Figure 7.32 Genetic recombination. Homologous sequences 3′ DNA A 5′ 3′ DNA B 5′ Enzyme nicks one strand of DNA at homologous sequence. A B Recombination enzyme inserts the cut strand into second molecule, which is nicked in the process. Ligase anneals nicked ends in new combinations. Molecules resolve into recombinants. Recombinant A © 2015 Pearson Education, Ltd. Recombinant B 39 8/29/2016 Genetic Recombination and Transfer • Horizontal Gene Transfer Among Prokaryotes • Vertical gene transfer • Passing of genes to the next generation • Horizontal gene transfer • Donor cell contributes part of genome to recipient cell • Three types • Transformation • Transduction • Bacterial conjugation © 2015 Pearson Education, Ltd. Genetic Recombination and Transfer • Horizontal Gene Transfer Among Prokaryotes • Transformation • Recipient cell takes up DNA from the environment • Provided evidence that DNA is genetic material • Cells that take up DNA are competent • Results from alterations in cell wall and cytoplasmic membrane that allow DNA to enter cell © 2015 Pearson Education, Ltd. 40 8/29/2016 Figure 7.33 Transformation of Streptococcus pneumoniae. Observations of Streptococcus pneumoniae Griffith's experiment: Living strain R Live cells Injection Capsule + In vitro transformation Heat-treated dead cells of strain S Mouse dies Injection DNA broken into pieces Heat-treated dead cells of strain S DNA fragment from strain S Living strain R Heat-treated dead cells of strain S Injection Mouse dies Some cells take up DNA from the environment and incorporate it into their chromosomes Mouse lives Culture of Streptococcus from dead mouse Strain R live cells (no capsule) Injection Transformed cells acquire ability to synthesize capsules Living cells with capsule (strain S) Mouse lives © 2015 Pearson Education, Ltd. Genetic Recombination and Transfer • Horizontal Gene Transfer Among Prokaryotes • Transduction • Transfer of DNA from one cell to another via replicating virus • Virus must be able to infect both donor and recipient cells • Virus that infects bacteria called a bacteriophage (phage) © 2015 Pearson Education, Ltd. 41 8/29/2016 Figure 7.34 Transduction. Bacteriophage Host bacterial cell (donor cell) Bacterial chromosome 1 Phage injects its DNA. 2 Phage enzymes degrade host DNA. Phage DNA Phage with donor DNA (transducing phage) 3 Cell synthesizes new phages that incorporate phage DNA and, mistakenly, some host DNA. Transducing phage Recipient host cell 4 Transducing phage injects donor DNA. Transduced cell Inserted DNA 5 Donor DNA is incorporated into recipient's chromosome by recombination. © 2015 Pearson Education, Ltd. Genetic Recombination and Transfer • Horizontal Gene Transfer Among Prokaryotes • Transduction • Generalized transduction • Transducing phage carries random DNA segment from donor to recipient • Specialized transduction • Only certain donor DNA sequences are transferred © 2015 Pearson Education, Ltd. 42 8/29/2016 Genetic Recombination and Transfer • Horizontal Gene Transfer Among Prokaryotes • Conjugation • Genetic transfer requires physical contact between the donor and recipient cell • Donor cell remains alive • Mediated by conjugation (sex) pili © 2015 Pearson Education, Ltd. Figure 7.35 Bacterial conjugation. F-plasmid is an episome, a plasmid that can integrate itself into the bacterial chromosome by homologous recombination F plasmid Origin of transfer Chromosome Pilus 1 Donor cell attaches to a recipient cell with its pilus. F+ cell _ F cell 2 Pilus may draw cells together. 3 One strand of F plasmid DNA transfers to the recipient. Pilus 4 The recipient synthesizes a complementary strand to become an F+ cell with a pilus; the donor synthesizes a complementary strand, restoring its complete plasmid. F+ cell F+ cell © 2015 Pearson Education, Ltd. 43 8/29/2016 Figure 7.36 Conjugation involving an Hfr cell. Donor chromosome High-frequency recombination cell = Hfr cell Pilus F+ cell 1 F plasmid integrates into chromosome by recombination. Hfr cell Pilus F+ cell (Hfr) F plasmid 2 Cells join via a pilus. – F recipient Donor DNA Part of F plasmid 3 Portion of F plasmid partially moves into recipient cell trailing a strand of donor's DNA. Incomplete F plasmid; cell remains F− 4 Conjugation ends with pieces of F plasmid and donor DNA in recipient cell; cells synthesize complementary DNA strands. 5 Donor DNA and recipient DNA recombine, making a recombinant F –cell. © 2015 Pearson Education, Ltd. Recombinant cell (still F− ) © 2015 Pearson Education, Ltd. 44 8/29/2016 Genetic Recombination and Transfer • Transposons and Transposition • Transposons • Segments of DNA that move from one location to another in the same or different molecule • Result is a kind of frameshift insertion (transpositions) • Transposons all contain palindromic sequences at each end © 2015 Pearson Education, Ltd. Figure 7.37 Transposition. Transposon Plasmid with transposon DNA Jumping transposons. Transposons move from one place to another on a DNA molecule. Replicating transposons. Transposons may replicate while moving, resulting in more transposons in the cell. Transposons can move onto plasmids. Transposons moving onto plasmids can be transferred to another cell. © 2015 Pearson Education, Ltd. 45 8/29/2016 Genetic Recombination and Transfer • Transposons and Transposition • Simplest transposons • Insertion sequences • Have no more than two inverted repeats and a gene for transposase • Complex transposons • Contain one or more genes not connected with transposition © 2015 Pearson Education, Ltd. Figure 7.38 Transposons. Transposon: Insertion sequence IS1 A CT T AC T GA T T GA A T GAC T A A T CAG T AAG T T AG T CA T T CA Inverted repeat (IR) Transposase gene Inverted repeat (IR) DNA molecule Target site IS1 Target IS1 Transposase Target site Copy Copy of of IS1 target site Original IS1 Complex transposon IR IS1 © 2015 Pearson Education, Ltd. IR Kanamycin- IR resistance gene IR IS1 46