* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Anabaena - Oxford Academic
Zinc finger nuclease wikipedia , lookup
Transformation (genetics) wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
Genetic code wikipedia , lookup
Real-time polymerase chain reaction wikipedia , lookup
Metalloprotein wikipedia , lookup
Gene desert wikipedia , lookup
Gene nomenclature wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Genetic engineering wikipedia , lookup
Non-coding DNA wikipedia , lookup
Gene expression wikipedia , lookup
Transcriptional regulation wikipedia , lookup
Biochemistry wikipedia , lookup
Molecular cloning wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Cyanobacteria wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Genomic library wikipedia , lookup
Restriction enzyme wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Gene regulatory network wikipedia , lookup
Chloroplast DNA wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Point mutation wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Community fingerprinting wikipedia , lookup
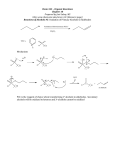
MICROBIOLOGY LETTERS EJSWIER FEMS Microbiology Lettera 133 (1995) 187-193 A comparison of gene organization in the zwf region of the genomes of the cyanobacteria Synechococcus sp. PCC 7942 and Anabaena sp. PCC 7 120 Julie Newman, Haydar Karakaya, David J. Scanlan, Nicholas H. Mann Department oj’Biologica1 qf Sciences, Uniwrsip Received 21 August 1995; revised 7 September Warwick. Cm,entp 1995; accepted * CV4 7AL, UK I4 September I995 Abstract The region of the genome encoding the glucose-6-phosphate dehydrogenase gene zwf was analysed in a unicellular S~~~choco~u.s sp. PCC 7942, and a filamentous, heterocystous cyanobacterium, Anabaena sp. PCC 7 120. Comparison of cyanobacterial zwf se q uences revealed the presence of two absolutely conserved cysteine residues which may be implicated in the light/dark control of enzyme activity. The presence in both strains of a gene flp, encoding fructose- 1,6_bisphosphatase, upstream from zwf strongly suggests that the oxidative pentose phosphate pathway in these organisms may function to completely oxidize glucose 6-phosphate to COz. The amino acid sequence of fructose- I ,6-bisphosphatase does not support the idea of its light activation by a thiol/disulfide exchange mechanism. In the case of Anahaena sp. PCC 7120, the ral gene, encoding transaldolase, lies between ;M?f and &J. cyanobacterium, Kepvordx Glucose-6-phosphate dehydrogenase; ;v$: Fructo se- I ,6-bisphosphatase; 1. Introduction Swechococcus; Anabaena * Corresponding author. Tel.: +44 (1203) 523 526; Fax: f44 (1203) 523 701; E-mail: [email protected]. chococcus sp. PCC 7942 exhibited similar dark respiratory activity, as measured by oxygen uptake, to that of the wild-type [21]. Thus cyanobacteria may employ an alternative respiratory pathway when the OPP is non-functional. The OPP is also thought to be largely responsible for the supply of reductant, in the light, to nitrogenase in the heterocyst [2,24]. Because of the central physiological importance of this transition from phototrophic metabolism to heterotrophic metabolism in the dark, the mechanisms involved in regulating the activity of key reductive and oxidative pentose phosphate cycle enzymes have been the focus of much attention. As is the case with higher plants, cyanobacteria exhibit light/dark activation/ inactivation of Calvin cycle enzymes. In Nostoc sp. MAC experiments with permeabilized cells have 037%1097/95/$09.50 Societies. All rights reserved The dominant nutritional mode of cyanobacteria is photoautotrophy involving the assimilation of CO? through the reductive pentose phosphate pathway (RPP). However, these organisms are also capable of generating maintenance energy during periods of darkness via the dissimilation of fixed carbon stored as glycogen. Dark respiration is thought to proceed exclusively through the oxidative pentose phosphate pathway (OPP) (for review see [22]). However, it has been shown recently that a zwf mutant of Sync- 0 1995 Federation SSDI 037%1097(95)00369-X of European Microbiological suggested the light activation of fructose- I ,6-bisphosphatase, sedoheptulose- 1,7-bisphosphatase, ribulose-5phosphate kinase and NADP-linked glyceraldehyde-3-phosphate dehydrogenase, possibly via a thioredoxin-based mechanism [3]. However, there is as yet little detailed information as regards the mechanism(s) of light/dark enzyme activation/inactivation. In this study, gene organization in the ?Mif regions of the genomes of two cyanobacteria was analysed to establish whether other genes encoding enzymes of dark metabolism are located close to ZVV~ and to verify whether the predicted amino acid sequences of the proteins are consistent with thioredoxin control of activity. DNA sequence analysis was performed using both random and directed cloning of fragments in Ml3 mp18 and M 13 mp19. Reactions were primed using universal - 40 primers or synthetic oligonucleotides, using Sequenase II polymerase (Amersham plc). The nucleotide sequence was determined using the dideoxy chain termination method. Analysis of DNA and protein sequence information was carried out using the Wisconsin Package version 8 [ 171. 2. Materials 3. Results and discussion 2. I. Bacterial tions and methods struins, plasmids and culture condi- Synechococcus sp. PCC7942 and Anabaena sp. PCC7 120 were grown at 30°C under white fluorescent light (20 PE m-2 s- ‘) in liquid BGl 1 medium [ 181. Escherichia coli MC1061 and TG 1 were used for plasmid constructions and DNA sequencing respectively and were grown in LB medium and 2 X YT medium [14]. 2.2. DNA manipulations Chromosomal DNA from the cyanobacterial strains was isolated using a method described previously [ 191. Plasmid isolation from E. cd, restriction digestion, ligation using T4 ligase and transformations in E, cdi were performed using standard molecular biological techniques [ 141. DNA fragments were isolated from agarose gels using the Geneclean kit (Bio 101 Inc). Chromosomal DNA was restricted using various restriction enzymes, according to the manufacturers’ instructions. Southern blotting of this DNA was performed using nitrocellulose (HybondC, Amersham plc) as described by Maniatis et al. [ 141. DNA fragments used for probes were restricted, isolated from low melting point agarose and labelled with [j2P]dCTP using the random priming method [9]. Filters were hybridised under low stringency conditions (.55”C, 5 X SSPE, 5 X Denhardt’s, 0.1% SDS) [ 141 and washed at 55°C in 2 X SSC. unless otherwise stated. 2.3. DNA sequence determinution 3.1. Gene organization 7942 and analysis in Syzechococcus sp. PCC We had previously cloned and sequenced the zyf gene encoding glucose-6-phosphate dehydrogenase from Synechococcus sp. PCC 7942 [20] and were interested in establishing whether genes encoding other enzymes associated with respiratory metabolism were clustered in this region of the genome. Complete sequencing of the 2.8-kb Hind111 fragment containing the Synechococcus zwf gene revealed an incomplete ORF upstream (5’) of Z~I$ A Sal1 site occurs in the middle of the ZW~gene and so the two corresponding Sal1 fragments (approx. 6 kb and 5 kb) were cloned in plasmid pUC19 to yield plasmids pDA and pDB. The sequence of the upstream 1033 bp ORF was completed using oligonucleotide primers and a 3-kb FsfI fragment from pDA sub-cloned into Ml3 mp18 and 19. This ORF was identified as coding for fructose- 1,6-bisphosphatase on the basis of the similarity of its translation product to known sequences including E. coli and several plant sources and was hence designated as jbp. Subsequently, the jbp gene from the cyanobacterium Nostoc sp. strain ATCC 29 133 was cloned and sequenced 1231 and the two cyanobacterial proteins exhibited 8 I % similarity (67% identity). Approximately 4.5 kb of sequence upstream from jbp has been analysed and surprisingly no other ORFs were detected. Sequence information downstream of wf was obtained by sub- .‘_vnechoroccuv sp. PCC 7942 petD petB fapl Anahaena sp P(‘(‘ 7 I20 tal fbP W- 1 kb Fig. I. Gene organization in the :n:f’ region of the genomes 01 Slnrchoc,oc,~u.\ sp. PCC 7942 and A~7ahtrrno sp. PCC 7 120. The nucleotide sequence information on which this diagram is based has the Genbank accession numbers U33282 7 120) and U33285 (S~ilec,ho~oc,c,rrs sp. PCC ( Adxrrmr hp. PCC 7942). cloning a contiguous 1.7-kb Hind111 fragment from pDB into Ml 3 mpl8 and 19 and extended using a 2.8-kb Hind111 fragment from pDB. Three further ORFs were identified. This sequence information is summarized in Fig. 1. The two ORFs furthest downstream from :“:f and on the complementary strand are identified as encoding cytochrome b, ( petB) and subunit IV (perD) of the cytochrome b6/‘f‘ complex by virtue of the similarities with Swechococcw sp. PCC 7002 [61. The ORF immediately downstream from ZW~ potentially encodes a polypeptide of 445 amino acids, and the only protein in the databases with any significant similarity (68%) is that encoded by the ORF immediately downstream of :w$ in Nostoc sp. strain ATCC 29133 [23]. Although no clue to the role. if any, of the protein in the oxidative pentose phosphate pathway was forthcoming from sequence comparisons, it is known to be always co-transcribed with :yf in Nostoc. sp. strain ATCC 29 133 [23]. There is recent evidence [24] (Karakaya. Scanlan. Sundaram. Newman and Mann. unpublished results) that the protein encoded by this ORF (designated opcA) is involved in the functional assembly of glucose-6-phosphate dehydrogenase. 3.2. Gerlr orgcrnix~ion in Anabuena sp, PCC 7942 Synechowccus sp. PCC 7942 is a unicellular strain and is incapable of nitrogen fixation and heterocyst formation. Since the OPP is the major supplier of reductant to nitrogenase in the heterocyst, we decided to examine the zu:f region of the genome of a filamentous, heterocyst-producing strain, namely Anubuetza sp. PCC 7 120. A I.%kb BamHI/HindIII fragment carrying the downstream (3’) half of the Synechococms sp. PCC 7942 ;M,-fgene was used to probe a Southern blot of HindIII-digested DNA from Atmbaenu sp. PCC 7 120. A 7-kb fragment hybridized strongly (data not shown) and a clone carrying this fragment was isolated from a size-fractionated HitId Atzabaenu sp. PCC 7 120 library in pBR325. A 2.4.kb HpuI/HindIII fragment was sub-cloned into M 13 mpl8 and 19 for sequencing. which was completed by random sequencing in Ml3 mpl8. This yielded the complete :\\:f gene and an incomplete ORF upstream. The translation products of the :\~:f genes from Anubaena sp. PCC 7120 and S!,tzpc.hoL,oc,c,ussp. PCC 7942 exhibited 83% similarity (70% identity). The sequence of the upstream ORF was completed by random sequencing of a 1. I-kb EcoRI/HpuI fragment in Ml3 mp18. The ORF encoded a polypeptide of 381 amino acids which on the basis of 50% sequence similarity (30% identity) to the Succhcrrotnyes cerel,isiue protein was identified as encoding transaldolase and was designated rd. Subsequently, it was shown to be 93% similar (83% identical) to the transaldolase of Nosmc sp. strain ATCC 29133 [23]. There was a further ORF upstream (5’) from tal, the sequence of which was completed by analysis of a contiguous 3.5-kb EwRI fragment. This ORF was identified as encoding fructose- 1.6-bisphosphatase on the basis of the similarity of its translation product with that of the Swechococws sp. PCC 7942 ,fbp gene. All this sequence information is summarized in Fig. I. 3.3. Cotnparison of gene nrgani,7don The arrangement of the .fbp. tal and zb;f genes is the same as that reported for Nosmc sp. strain ATCC 29133 [23], which is also a filamentous, heterocystous strain. However, in the unicellular strain Swechocwcms sp. PCC 7942 there is no ml gene between .fbp and :\z:f: An internal 0.5-kb HpaI/ClaI fragment of the rul gene from Ambuenu sp. PCC 7 120 was used to probe a Southern blot of Syncchococcus sp. PCC 7942 DNA under conditions of moderate stringency (hybridization in 5 X SSPE, 55°C; washing in 2 X SSPE, 55°C) and yielded only 190 .I. Newmnn etul./~EMSMicrohiolo~~Letters 133 (IYYSI 1X7-lY3 50 1 ..... Anabaena Nostoc Synechococcus Chloroplast Cytosolic Consensus ... .... ..... . . . .I........ . . . . . . . . . . . . . . . . . . . . . . . . . ,......... . . . . . . . . . . maasaattts shlllsssrh vasssqpsil sprslfsnng kraptgvrnh .. .... .. .. ..... .. . . . ... .. __________ _____..... _________. __________ __________ Anabaena NOStOC Synechococcus Chloroplast Cytosolic consensus 51 ..... ...MA KAsESLDlSV NEsTDkALDR DCTTLSRHVL QQLQSFSaDA 100 . . . . . . ..MA KtpESLEsSI NEiTDPALDR DCTTLSRHVL QQLQSFSpDA . . . . . . ..MA qsttS..... .EthtRdLDR DCTTLSRHVL eQLQSFSpEA qyasgvrcMA VAaDasETkt aarkksgyE1 q..TLtgwlL r.qemkgeid . . . . . . ..Md hAgDsNrT.. . . . . . . ..Dl m..TitRyVL neqskrpesr --------MA KA--SLETS- NE-TDRALDR DCTTLSRHVL QQLQSFS-DA QDLsALNnRI ALAgKIVARR LSRAGLMEGv LGFTGEVNVQ 150 GEsVKKMDVY QDLSAIMnRI ALAgKIVARR MSRAGLMEGV LGFTGHVNVQ GEsVKKMDVY QDLsALMqRI gLAaKLIARR LShAGLvDda LGFTGEINVQ GEaVKrMDVY aELtivMss1 sLAcKqIAs1 vqRAGi.snl tGvqGaINIQ GEdqKKLDVi gDFtiLLsh1 vLgcKFVcsa vnkAGL.akl iGLaGEtNIQ GEeqKKLDVl QDL-ALM-RI ALA-KLVARR LSRAGLMEG- LGFTGE-NVQ GE-VKKMDVY BEMDEPYYIP ENCPIGRYTL 101 Anabaena Nostoc Synechococcus Chloroplast Cytosolic consensus t 151 ANDVFISVFk QSGLVCRLAS l Anabaena Nostoc Synechococcus Chloroplast Cytosolic consensus ANDVFISVFk QSGLVCRLAS sNEVFsncLr sSGrtgiiAS sN!JVFVkaLt SSGrtCiLvS AN-VFISVF- QSGLVCRLAS t dnabaena Nostoc Synechococcus Chloroplast Cytosolic consensu* alignment, Arabidopsis in upper case. redox-sensitive produced sp. PCC thaliancl The 7942 using (this study), cysteines residues 250 .._.. .GdDsDGqAK DLLtnGRkQI .GtDsDGkAt DLLanGRkQl .fyDEsheAK 1GsEEqrciv ...fEtatle -G-DEL%-AK 7 DLLQPGdrQI nvcQPGnnl1 DvLQPGknmV DLLQPGR-QI t 300 RIPdHGaVYS RIPNIiGsVYS qlPNsGqIYS eIPkaGrIYS kIPNkGkIYS RIPNHG-IYS tDnNLSlGS1 FaIRQQE... . VDvdLnvGSI IDaavStGSI IDcgvSiGtI ID-NLS-GSI FaVRrQE... FgIyspnDec FgIymvkD.. F-IRQQED-- . .... ... ivddsddisa . ..___.. ---------- 251 AAGYILYGpS AAGYILYGpc AAGYVLYGaS AAGYcMYssS AAGYcMYGsS AAGYILYG-S c TMLVYTMGtG TMLVYTiGKG TLLVYsMGqG viFVlTLGKG CtLVlstGsG TMLVYTMGKG VHSFTLDPSL GEFILseENI VIiSFvLDPSL GEFILTeENI VHvFvLDPSL GBFVLaqsdI VfSFTLDPmy GEFVLTqENI VngFTLDPSL GEYILThpdI VHSFTLDPSL GBFILT-EN1 l EgyRqYIRem HRrEa....Y SgRYSGALVa DfHRILMQGG 351 VFLYPGTIqN PeGKLRLLYE sAPLAFLIqQ AGGrAtTGLV VFLYPGTIqN PeGKLRLLYE tAPIdFLIEQ AGGrAtTGLV VFLYPeTVFN PtGKLRLLYE aAPMAFLaEQ AGGkAsdGqk IYgYPrdaKs knGKLRLLYE cAPMsFiVEQ AGGkgsdGhs IFLYPGdkKs PnGKLRvLYE VfPMsFLmEQ AGGqAfTGkq VFLYPGTIKN P-GKLRLLYE -APMAFLIEQ AGG-A-TGLV 400 dILDWPkKL nILDWPkKL pILl+qPqaL RVLDIqPtei RaLDlIPtKi RILDVVP-KL 401 429 HQRTPLIIGS KEDVaKVESF iqNGH.... the HQRTPLIIGS KEDVaKVEsF iqNGH HeRcPLIIGS aaDVDfVEac 1Aesvp tEEVEKlEkY lA....... HeRsPvflGS yDDVHdIkaL yAsqekta HQRTPLIIGS KEDVHKVE-F -ANGH---- PILEUP enzyme ._. HQRvPLyIGS Ancrhaenrr implicated in the cytosolic l ._....._.. [ 171, of the amino programme sp. PCC 7120 [I I] and the cytosolic form from Spinoceu cysteine LYDPiDGSSN LYDPLDGSaN vFDPLDGSSN vFDPLDGSSN LYDPLDGSSN fNEGNYqmWD DkLkkYIddl kdpgptgkPY SARYiGsLVg DfHRtLLyGG VNEGNaknWD gpttkYVekc kfptdgsspk SlRYiGsMVa DVHRtLLyGG VNEGNFWQW- ES-R-YIR-- HR-EG---PY SARYSGAMV- DIHRILLQGG Anabaena Nostoc Synechococcus Chloroplast Cytosolic consensus An ENCPIGRYTL ENCPIGRYTL EesysGnYw EpslrGkYcv ENCPIGRYTL 201 tDtNLSlGS1 FsIRQQE... VNEGNFNQNp Armbaena Nostoc Synechococcus Chloroplast Cytosolic consensus 2. BEMEnPWIP EEHEkPYYIP BEeDvPvaV. EEdEEatFI. EEMEEPYYIP 301 350 VNEGNFWQWE ESMReYIRyv HRtEG....Y tARYSGAMVs DIHRILvQGG VNEGNFWQWE ESiReYIRyv HRtEG....Y SARYSGAMVs DIHRILVQGG Anabaena Nostoc Synechococcus Chloroplast Cytosolic consensus S\n&rococcuu QSGLVCRLAS ANqVFISVFr Anabaena Nostoc Synechococcus Chloroplast Cytosolic consensus Fig. 200 LYDPiDGSSN in activation are marked of (this olrrc~ru study), [12]. the chloroplast (* ). Nostoc Amino enzyme acid sequences sp. ATCC acid residues are indicated of 29133 fructose-1,6-bisphosphatase from [23], from agreeing by the chloroplast with vertical enzyme the consensus arrows and those are shown potential 191 J. Newman et al./ FEMS Microhiolog~ Letter.~ 133 ClYY5) 1X7-193 CO!lSeL¶US 1 .mvsLLENPL .mvsLLENPL mtpkLLENPL ----LLENPL RVGLqQqgmP RVGLqQqgmP RIGLrQdkvP RVGL-Q---P 50 EPQIiVIFGA sGDLTwRKLV PAlYkLrrER EPQIiVIFGA sGDLTwRKLV PAlYkLrrER EPQIlVIFGA tGDLTqRKLV PAiYeMhlER EPQI-VIFGA -GDLT-RKLV PA-Y-L--ER Anabaena Nostoc Synechococcus consensus 51 RiPPEtTIVG RiPPEtTIVG RlPPElTIVG R-PPE-TIVG VARREWShEY VARREWShEY VARRDWSdDY VARREWS-EY 100 FREqMqkGmE eahssVelgE 1WqdFsQGLF FREqMqkGmE eahpdVdlgE 1WqdFsQGLF FREhLrqGvE qfgggIqaeE vWntFaQGLF FRE-M--G-E -----V---E -W--F-QGLF Anabaena Nostoc Synechococcus consensus 150 101 YcPGdIDnPe sYQkLknlLs eLDEkRGTRG NRmFYLSVAP nFFpEAiKQL YsPGdIDnPe sYQkLktlLs eLDEkRGTRG NRmFYLSVAP sFFpEAiKQL FaPGnIDdPq fYQtL+drLa nLDElRGTRG NRtFYLSVAP rFFgEAaKQL Y-PG-ID-P- -YQ-L---L- -LDE-RGTRG Ni-FYLSVAP -PF-EA-KQL Anabaena Nostoc S'ynechococcus COIX3HX3U.5 200 151 GgaGMLdDPy KhRLVIEKPF GRDLaSAQsL NaVvQkyCkE hQVYRIDHYL GsgGMLeDPy KhRLVIEKPF GRDLaSAQsL NqVvQkyCkE hQVYRIDHYL GaaGMLaDPa KtRLVVEKPF GRDLsSAQvL NaIlQnvCrE sQIYRIDHYL G--GML-DP- K-RLVIEKPF GP.DL-SAQ-L N-V-Q--C-E -QVYRIDWn Anabaena Nostoc Symchococcus consensus 201 GKETVQNLLV FRFANAIFEP LWWRQFVDHV QITVAET'VGv GKETVQNLLV FRFANAIFEP LWNRQFVDHV QITVABTVGv GKETVQNLLV FRFANAIFEP LWNRQYIDHV QITVAETVGl GKETVQNLLV FRFANAIFEP LWNRQFVDHV QITVAETVG- Anabaena Nostoc Synechococcus consensus 251 GALRDMlQNH GALRDMlQNH GALRDMvQNH GALRDM-QNH LMQLYcLTAM LMQLYcLTAM LMQLFsLTAM LMQLY-LTAM Anabaena Nostoc Synechococcus consensus 301 SrSAIRGQYs SrSAVRGQYs SlSAVRGQYk S-SAVRGQY- AGWMkGqqVP gYRtEpGvDP nSsTPTWgM AGWMkGqaVP gYRtEpGvDP nStTPTYVaM AGWMnGrsVP aYRdEeGaDP qSfTPTYVaM AGWM-G--VP -YR-E-G-DP -S-TPTYV-M Anabaena Nostoc Synechococcus CCJllSellsUs 351 GVPFYLRTGK GVPFYLRTGK GVPFYLRTGK GVPFYLRTGK RMPKKVsEIs RMPKKVsEIa RMPKKVtEIa RMPKKV-EI- IhFrdVPsrM IhFreVPsrM IqFktVPhlM I-F--VP--M Anabaena Nostoc Synechococcus consensus 401 NEGISLRFDV NEGISLRFDV NEGVSLRFEV NEGISLRFDV KmPGaefRsR KmPGaefRtR KtPGssqRtR K-PG---R-R SVDMDFsYgs fgieaTsDAY SVDKDFsYgs fgiqaTsDAY SVDMDFrYdt afgspTqEAY SVDMDF-Y-- -----T-DAY Anabaena Nostoc Synechococcus consensus 451 DQTLFTRADE DQTLFTRADE DQTLFTRADE DQTLFTRADE Anabaena Nostoc Synechococcus consensus 501 INqDG..rrw rRl....... INqDG..rrw rR1 _._... INrDGavgw sRipatqlns IN_DG_____ _R________ Anabaena Nostoc synechococcus l 250 EdRAGYYEkA EdRAGYYEsA EgRAGWEtA E-RAGYYE-A 300 EaPNsMdADs IRtEKVKVlQ ATRLADVhnL EaPNaMdADs IRtEKVKVlQ ATP.LADVhnL SpPNsLgADg IRnEKVKVvQ ATRLADIddL E-PN-M-AD- IR-EKVKV-Q ATRLAD'J-L 350 KFLVDNWRWq KFLVDNWRWk KLLVDNWRWq KFLVDNWRW- 400 FQSAaQqrN. aNILaMRIQP FQSAaQqtN. aNILtMRIQP FQSAtQkvNs pNVLvLRIQP FQSA-Q--N- -NIL-MRIQP 450 dRLFlDCMMG dRLFlDCMMG sRLLvDCMLG -RLF-DCMMG l VEAaWqWTP VEAaWqWTP VSAsWrVVTP VEA-W-WTP aLsvWDsPad aLsvWDaPad 1LesWDdPrq -L--WD-P-- patIpqYEAG pttIpqYEAG aagIsfYEAG ---I--YEAG 500 TWEPaeAEfL TWJZPeqAElL TWEPaeAEqL TWEP--AS-L 525 _.... . .. . . sgdv _____ Fig. 3. An aligment, produced using the PILEUP programme [ 171. of the amino acid sequences of the glucose-6-phosphate dehydrogenases from Anabaena sp. PCC 7120 (this study), Nostoc sp. ATCC 29133 [23] and Syechococcus sp. PCC 7942 [21]. Amino acid residues agreeing with the consensus are shown in upper case. The two conserved cysteine residues are indicated (* ). a faint signal with a 6-kb Hind111 fragment. Thus it is not clear whether Synechococcus sp. PCC 7942 contains a ful gene. Although it is risky to infer physiological properties from DNA sequence information, the close proximity of the ,fbl, gene to :nf in all the cyanobacterial strains examined and their co-transcription in Nosfoc sp. strain ATCC 29133 [23] do suggest something about the way the OPP may be operating. The action of transaldolase and. presumably transketolase, regenerates fructose 6phosphate, which can re-enter the cycle, and glyceraldehyde 3-phosphate. Fructose- I ,6_bisphosphatase ensures that glyceraldehyde 3-phosphate via aldolase will also re-enter the cycle rather than be metabolized to pyruvate. Thus glucose 6-phosphate can be completely oxidized to CO, with the concomitant production of maximal amounts of NADPH. This is in keeping with the observations of BBhme [5], in relation to reductant supply to nitrogenase, that glycolytic degradation of hexose appears to be of minor importance and that aldolase and fructose- I ,6-bisphosphatase function to provide the oxidative pentose phosphate pathway with additional hexose phosphates. 3.4. Implications for regulation of enzyme actir!ity Cyanobacteria, like plants. regulate certain enzyme activities in response to light-dark transitions. The light activation of several enzymes including fructose- I ,6-bisphosphatase, sedoheptulose- I ,7-bisphosphatase, ribulose-5-phosphate kinase and NADP-linked glyceraldehyde-3-phosphate dehydrogenase, possibly via a thioredoxin based mechanism, has been reported for permeabilized cells of Nostoc sp. [3]. In plants there are two distinct fructose-l,6bisphosphatases, chloroplast and cytosolic, with different regulatory properties. The cytosolic enzyme is allosterically regulated by AMP, whereas the chloroplast enzyme is regulated via a thiol/disulfide exchange mechanism involving thioredoxin acting as a protein disulfide reductase [7]. The chloroplast enzyme, compared to the cytosolic form, typically has an insertion of 12-17 amino acids with two adjacent conserved cysteine residues for the light regulation of enzyme activity [ 151. In cyanobacteria, fructose1,6-bisphosphatase is required both for the RPP in the light and also the OPP in the dark. There are conflicting reports regarding the regulatory properties of cyanobacterial fructose- 1,6_bisphosphatase activity. Bishop [4] reported the enzyme from Anucystis nidulurz~ to exhibit regulatory characteristics that were not typical for either form of the enzyme. This conflicts with the report that the regulatory properties of fructose- I ,6-bisphosphatase from Anucystis nidulans resembled that of the chloroplast enzyme with respect to agents such as oxidized and reduced glutathione. ascorbic acid and dithionite [2.5]. Comparison of the cyanobacterial amino acid sequences reported here, and that of Anabuena sp. ATCC 29133 [23], to the plant enzymes reveals them to be of the cytosolic type, in that they lack the insertion and adjacent cysteines typical of the chloroplast form of the enzyme (Fig. 2). Recently it has been demonstrated that even the cytosolic fructose- I ,6-bisphosphatase from sugarbeet exhibits a slow light activation and light-dependent AMP sensitivity [ 131. This observation may be explained by the presence of potential redox-sensitive cysteine pairs predicted by tertiary structure modelling in cytosolic forms of the enzyme [I]. However, these potential redox-sensitive cysteines, apart from one, are not conserved in the cyanobacterial enzymes (Fig. 2). Consequently, it seems likely that if the cyanobacterial fructose- I ,6bisphosphatases are subject to light-dark regulation it is not via a thiol/disulfide exchange mechanism. Several studies have been aimed at elucidating the mechanisms by which glucose 6-phosphate activity is regulated during light-dark transitions in cyanobacteria. Metabolites including NADPH [2,16] and ATP [IO] have been implicated in regulation and thioredoxin control has also been proposed by Cossar et al. [8]. In keeping with the thiol/disulfide exchange mechanism of regulation, sequence analysis of the Swwchococc~l.s sp. PCC 7942 ,-rvf gene revealed the protein to have two cysteine residues which are not present in the enzyme from other prokaryotic sources [20]. Comparison of the glucose6-phosphate dehydrogenase sequences from Anubaena sp. PCC 7120 (this study), Nostoc sp. ATCC 29133 [23] with that of the Synechococws sp. PCC 7942 protein reveals these two cysteines at positions 188 and 447 to be absolutely conserved (Fig. 31, reinforcing the likelihood of their role in the regulation of enzyme activity. and characterization of a cDNA Acknowledgements encoding cytoaolic fructose- 1.6.bisphosphatase from spinach. Plant Mol. Biol. 18, 799X02. Haydar Karakaya was supported by a studentship provided by the Turkish Government through Ondokuz Mayis University. This work benefitted from the use of the SEQNET facility. [I.?1 Khayat. E.. Harn. C. and Daie, J. (1993) Purification and light-dependent molecular modulation of the cytosolic fructose- I .h-bisphosphatase in sugarbeet leaves. Plant Physiol. 101. 57-63. [I41 Maniatis. T., Fritach, E.F. and Sambrook. J. (1981-j Molecular Cloning: A Laboratory Manual. Cold Spring Harbor Laboratory, Cold Spring Harbor, NY. [I51 References Marcus, F.. Moberly. L. and Latshaw. S.P. (1988) Comparative amino acid sequence of fructose I ,h-bisphosphatases: identification of a region unique to the light-regulated chloro- [II Anderson, L.E., Li, D.. Prakash, N. and Stevens, F.J. (1995) Identification of potential redox-sensitive cysteines in cytoso- plast enzyme. Proc. Natl. Acad. Sci. USA 85, 5379-5383. I161 Pelroy. tern II regulation of macromolecule phosphate dehydrogenase. Planta 196, I l8- 124. green alga Aphtruoccptr [21 Apte, S.K., Rowell, P. and Stewart. W.D.P. donation to ferredoxin Anaharna cylindricu. [31 Austin, (1978) Electron in heterocysts of the Nz-fixing Proc. R. Sot. B200, [I71 alga l-25 for the Wisconsin Package. 1994. Genetics Computer Group, Version 575 8, Science IIS1 Rippka, Light/dark R.. Deruellea, J.B., Waterbury, J.B.. Herdman. M. regulation of photosynthetic enzymes within intact cells of and Stanier, R.Y. (1979) the cyanobacterium ries and propertiea of pure cultures of cyanobacteria. J. Gen. Nosroc, sp. MAC. Biochim. Biophys. Microbial. R.H. (1979) Regulatory characteristics of a fructose bisphosphatase from nidulans. bacterium Anac~~~is I I I, l-61. N.G. (1990) Construction of vectors for use in Swrchococcus: Ancrhamrr system from heterocysts of Generic assignments, strain histo- Scanlan, D.J.. Bloye. S.A., Mann. N.H.. Hodgson. D.A. and Carr. H. (I 987) Regulation of electron flow to nitrogenase in a cell-free ~dilis. the blue-green [I91 Arch. Biochem. Biophys. 196. 295-300. [51 Biihme. urri- cation of CO,-regulated DO1 Scanlan. /trcZ promoter probe application to the identifi- promoters. Gene 90. 43-49. D.J.. Newman, J., Sebaihia, M.. Mann, N.H. and Biochim. Biophys. Acta 891, 121-128 Carr. S.N., Tam, X. and Widger, W.R. (1992) Cloning and glucose-h-phosphate dehydrogenase gene from the cyanobac- [61 Brand, sequencing of the p&ID S,wzec~hococcus [71 Clancey, operon from the cyanobacterium C.J. and Gilbert, H.F. (1987) Thiol/disulfide ex- spinach chloroplast fructose-l .6-bisphosphatase. J. (1992) Cloning and sequence analysis of the 880. 1211 Scanlan, D.J.. Sundaram, S., Newman, J., Mann, N.H. ckococcrr.\ sp. strain PCC 7942. J. Bacterial. 177. 2550-2553. WI Smith, A.J. (1982) Modes of cyanobacterial Cossar, J.D.. Rowell, P. and Stewart, W.D.P. (I 984) Thiore- metabolism. In: The Biology of Cyanobacteria doxin as a modulator of glucose-6.phosphate dehydrogenase and Whitton. in a Nz-fixing cyanobacterium. J. Gen. Microbial. 130. 99l- D31 A.P. and Vogelstein, B. (1984) technique for radiolabelling DNA fragments activity. to high specific Addendum: a restriction endonuclease Anal. Biochem. 137, 266-267. R.E. (1975) Eds.), pp. 47-86. Blackwell Scientific Summers, M.L., Regulation of the Meeks, J.C., Chu. S. and Wolf, R.E. Jr. (1995) Nucleotide sequence of an operon in Nostoc sp. strain ATCC 29133 encoding four genes of the oxidative pentose phosphate cycle. Plant Physiol. 107, 267-268. 1241 Summers. Grossman, A. and McGowan, B.A.. carbon (Carr, N.G. Publications. Oxford. 998. I91 Feinberg, and Carr N.G. (1995) Characterization of a :rb:f mutant of Sxne- Biol. Chem. 262, 13545- 13549. N.G. terium S~nrchoc.occl,s PCC 7942. Plant Mol. Biol. 19, X77- sp. PCC 7002. Plant Mol. Biol. 20. 48 I-49 I. change in the thioredoxin catalysed reductive activation of [I II Manual bynthesis in the hlue- 67 14. J. Bacterial. 128, 623-632. Drive, Madison. WI 537 I I P.A., Ross, I.S. and Mills, J.D. (1992) [41 Bishop, [lOI Program September Acta 1099, 226-232. RI R.A.. Kirk, M.R. and Bassham. J.A. (1976) Photosya- lit forms of fructose bisphosphatase and glyceraldehyde-3- M.L., Wallis, J.G., Campbell, E.L. and Meeks, J.C. ( 1995) Genetic evidence of a major role for glucose-6. glucose-6-phosphate dehydrogenase in blue-green algae. Plant phosphate dehydrogenase in nitrogen fixation and dark growth Physiol. 55, 658-662. of the cyanobacterium Horsnell, P.R. and Raines. C.A. (I99 I) Nucleotide sequence Bacterial., in presh. of a cDNA clone sphosphatase from encoding Arabidopsis chloroplast thalianrr. fructose-l,6-biPlant Mol. Biol. 17. 185-186. 1121 Hur, Y.. Unger, E.A. and Vasconcelos. A.C. (1992) Isolation [251 Udvardy, Nostoc sp. strain ATCC J.. Godeh, M.H. and Farkas, G.L. (1982) tory properties of a fructose cyanobacterium 208. 29133. l,h-bisphosphatase r\/~uc~sfis nidulam. J. Bacterial. J. Regulafrom the I5 I. 203-