* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download A genome screen for linkage in Australian sibling-pairs with
Skewed X-inactivation wikipedia , lookup
Medical genetics wikipedia , lookup
Ridge (biology) wikipedia , lookup
Gene expression profiling wikipedia , lookup
Gene expression programming wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Whole genome sequencing wikipedia , lookup
Population genetics wikipedia , lookup
Heritability of IQ wikipedia , lookup
Segmental Duplication on the Human Y Chromosome wikipedia , lookup
Genomic imprinting wikipedia , lookup
Biology and sexual orientation wikipedia , lookup
Genetic engineering wikipedia , lookup
Pathogenomics wikipedia , lookup
Y chromosome wikipedia , lookup
Neocentromere wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Non-coding DNA wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Genomic library wikipedia , lookup
Genome-wide association study wikipedia , lookup
X-inactivation wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Human genetic variation wikipedia , lookup
Human genome wikipedia , lookup
Designer baby wikipedia , lookup
Human Genome Project wikipedia , lookup
Genome editing wikipedia , lookup
Microevolution wikipedia , lookup
History of genetic engineering wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Minimal genome wikipedia , lookup
Genome evolution wikipedia , lookup
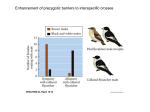
Genes and Immunity (2002) 3, 464–469 & 2002 Nature Publishing Group All rights reserved 1466-4879/02 $25.00 www.nature.com/gene A genome screen for linkage in Australian sibling-pairs with multiple sclerosis M Ban1,2, GJ Stewart2, BH Bennetts3, R Heard2, R Simmons4, M Maranian1, A Compston1 and SJ Sawcer1 1 Neurology Unit, Addenbrooke’s Hospital, University of Cambridge, Cambridge, UK; 2Institute for Immunology and Allergy Research, Westmead Millennium Institute, Westmead Hospital, University of Sydney, NSW, Australia; 3Department of Paediatrics and Child Health, Children’s Hospital at Westmead, University of Sydney, NSW, Australia; 4Australian National Register of Multiple Sclerosis Families, Canberra Hospital, ACT, Australia The role of genetic factors in determining susceptibility to multiple sclerosis is well established but, despite the global distribution of the disease, systematic efforts to locate susceptibility genes have concentrated exclusively on populations from the Northern Hemisphere. We performed a genome wide screen of linkage in the Australian population using a panel of 397 microsatellite markers in 54 affected sibling-pairs. Multipoint linkage analysis revealed four regions of suggestive linkage (on chromosomes 2p13, 4q26-28, 6q26 and Xp11) and 18 additional regions of potential linkage (at 1q43-44, 3q13-24, 4q24, 4q3134, 5q11-13, 6q27, 7q33-35, 8p23-21, 9q21, 13q31-32, 16p13, 16p11, 16q23-24, 17p13, 18p11, 20p12-11, Xp21-11 and Xq2328). Our results contribute to the available data adding new provisional regions of linkage as well as increasing support for areas previously implicated in genetic susceptibility to multiple sclerosis. Genes and Immunity (2002) 3, 464–469. doi:10.1038/sj.gene.6363910 Keywords: multiple sclerosis; linkage genome screen; genetic susceptibility Introduction Multiple sclerosis is a demyelinating disorder of the central nervous system with a complex aetiology in which unknown environmental factors are thought to trigger disease in genetically susceptible individuals.1 Epidemiological analysis of twins,2,3 half-sibs4 and adoptees5 confirm the involvement of genetic factors while the role of environmental agents is demonstrated by the results of migration studies.6 In Australia, studies of the prevalence of multiple sclerosis have been particularly influential showing a latitudinal relationship similar to that seen in the Northern Hemisphere with lower rates in northern regions closer to the equator (11/100 000) than in the south (74/ 100 000).7,8 However, overall, these prevalence rates are lower than those seen in Northern Europe, even though the majority of the Australian population originates from Northern Europe, suggesting that, despite expressing the same susceptibility genes, Australians may be at relatively reduced risk of multiple sclerosis through differential exposure to the causative environmental risk factors. To date seven whole genome screens for linkage have been completed in multiple sclerosis, five in Northern European9–13 and two in Southern European populaCorrespondence: GJ Stewart, Associate Professor, Institute for Immunology and Allergy Research, Westmead Millennium Institute, Westmead Campus, University of Sydney, Westmead, NSW 2145, Australia. E-mail [email protected] tions.14,15 None of these screens identified statistically unequivocal evidence for linkage but nearly all found more regions of potential linkage than would be expected to have occurred by chance alone. Several regions show some degree of consistency between screens, including the major histocompatibility complex (MHC) on chromosome 6p21. A systematic search for linkage in multiple sclerosis has not previously been performed in a population from the Southern Hemisphere. Here, we report a genome screen for linkage in 54 Australian affected sibling-pairs with multiple sclerosis using a panel of 397 microsatellite markers. Results Results of the multipoint non-parametric linkage analysis are shown graphically in Figure 1. A Maximum lod score (MLS) value of greater than 1.8, the threshold of suggestive linkage, was seen on chromosomes 2p13, 4q26-28, 6q26 and Xp11. Another 18 regionsFon chromosomes 1q43-44, 3q13-24, 4q24, 4q31-34, 5q11-13, 6q27, 7q33-35, 8p23-21, 9q21, 13q31-32, 16p13, 16p11, 16q23-24, 17p13, 18p11, 20p12-11, Xp21-11 and Xq23-28Fhad MLS values X0.7, indicating potential linkage. The mean genetic information extracted from the 54 families across the genome was 61%, ranging from 16% on chromosome 19pter to 86% on chromosome 4 in a region covered by particularly informative markers. Australian MS linkage screen M Ban et al 465 Figure 1 MLS calculated by MAPMAKER/SIBS at each point along the genome is shown for each chromosome. The x-axis is proportional to the length of the corresponding chromosome and the map position of the markers is indicated by the tick marks. In each case the y-axis is scaled from 0 to 2.5. Discussion We have completed a genome wide screen for linkage in 54 Australian affected sibling-pairs with multiple sclerosis using 397 markers and identified four regions of suggestive linkage and 18 additional regions of potential linkage. Under the null hypothesis of no susceptibility for multiple sclerosis, a marker map of this density would only be expected to generate a single region with an MLSX1.8 and approximately 10 regions with MLS values X0.7.16,17 As the number of observed peaks greatly exceeds that expected by chance at both MLSX0.7 and X1.8, it is likely that at least some of these regions harbour susceptibility genes. This interpretation is further supported by the fact that two of the four regions providing suggestive linkage have previously featured in multiple sclerosis genome screens: a regions close to chromosome 6q26 was observed in the UK,9 American,10 Nordic13 and Italian14 genome screens, whereas the Figure 1 (continued) suggestive linkage on chromosome Xp11 was previously reported in the UK screen.9 Several of our regions of potential linkage also coincide with those suggested in previous multiple sclerosis genome screens. A region identified in the present study on chromosome 1q43-44 was also seen in the Italian screen,14 and linkage at 4q31, 5q31, 7q33-34 and 18p11 was additionally observed in the American screen.10 Similarly, our potential linkage on chromosome 16p matches that observed in the Nordic13 and American screen,10 as well as in a meta-analysis of the original American, Canadian and British linkage screens.18 Since Genes and Immunity Australian MS linkage screen M Ban et al 466 Figure 1 (continued) 16p was not identified in the two screens in Southern European populations,14,15 this observation emphasises that alternate combinations of genes may determine susceptibility to multiple sclerosis in different populations (allelic and locus heterogeneity). Despite the wellestablished association of the MHC region with susceptibility to multiple sclerosis in several populations including Australia,19,20 we only found modest evidence for linkage in this region. While this result is intriguing Genes and Immunity Figure 1 (continued) as greater than 90% of our subjects were of Northern European origin, it reflects the limited statistical power of the linkage approach in small samples of affected family members. Australian MS linkage screen M Ban et al Figure 1 (continued) Although replication of linkage is the essence of narrowing the search for chromosomal regions of interest, failure to implicate any one chromosomal region across several screens does not necessarily undermine confidence that a susceptibility gene is genuinely encoded at that site. Variation in the evidence for linkage observed between studies can occur for a number of reasons. For example, the genes conferring susceptibility may differ between populations (genetic heterogeneity). More importantly, the limited statistical power of individual linkage screens means that random variation is expected to play a large part in the final result. Since the power is influenced not only by the number of families studied but also by the frequency of susceptibility alleles in the population screened, variation in the extent of evidence for linkage is to be expected.21 The two novel regions of suggestive linkage observed on chromosome 2p13 and 4q26-28 in our screen harbour a number of interesting immunological candidates. For example, IL-2 and IL-21 which are both involved in T and B cell proliferation map to chromosome 4q26-27, while the cluster of genes for the immunoglobulin kappa light chain and CD8 are encoded at chromosome 2p12. These candidates are worthy of further study although previous study of IL-2 in the UK yielded negative results.22 A number of investigators have suggested that some genetic factors influence susceptibility to a general autoimmune disease diathesis against which background additional genes determine which autoimmune condition develops. Studies showing increased prevalence of autoimmunity in first-degree relatives of multiple sclerosis patients23 and a higher than expected concordance of linkage results across genome screens from different autoimmune diseases24 support this hypothesis. A number of regions identified in our study overlap with those found in other autoimmune disease linkage screens. Chromosome 1q43-44 has been implicated in a genome screen of rheumatoid arthritis25 as well as in three independent studies of systemic lupus erythema- tosus.26–28 Chromosome 16q24 also showed evidence of linkage in inflammatory bowel disease29 as well as two independent screens of rheumatoid arthritis,25,30 and the region on 4q24 showed the strongest evidence of linkage outside the HLA region in a genome screen of rheumatoid arthritis.25 In addition, the region of suggestive linkage on chromosome Xp11 overlaps with a region linked to insulin-dependent diabetes mellitus.31,32 Evidence also exists for an autoimmune susceptibility gene on chromosome 4q26-28. An obvious positional candidate within this area is the IL-2 gene, which has been proposed as a possible autoimmunity gene on the basis of linkage to insulin-dependent diabetes mellitus in the NOD mouse and experimental autoimmune encephalomyelitis (the mouse model for human multiple sclerosis).33,34 Our study is consistent with the hypothesis that susceptibility to multiple sclerosis is determined by the action of many genes, each exerting only modest effects individually. Four regions of suggestive linkage (2p13, 4q26-28, 6q26 and Xp11), two of which (2p13 and 4q2628) have not previously been reported, are identified. These provisional results require further exploration through the use of a positional candidate approach and fine mapping of the regions in order to refine characterisation of genetic predisposition to multiple sclerosis in Australian and other Northern European populations. 467 Materials and methods Families The index cases and their affected siblings were recruited by the Australian National Register of Multiple Sclerosis Families, Canberra Hospital (Canberra) and the Institute for Immunology and Allergy Research, Westmead Hospital (Sydney). All patients gave informed written consent for genetic analysis and satisfy criteria for clinically definite (66%), clinically probable (9%), laboratory-supported definite (19%) or laboratory-supported probable (6%) multiple sclerosis.35 Families with affected half-siblings or sib-ships with more than two affected were not included. Although DNA was available from parents and/or unaffected siblings in 60% of families, these additional samples were not genotyped for reasons of genotyping efficiency. Others have shown using these markers that typing parents and unaffected family members results in only a modest increase in the amount of information extraction.13,36 Markers A total of 397 dinucleotide microsatellite markers from the Applied Biosystems Medium Density Linkage Mapping Set (LMS-MD10) were used. The markers were observed to have a mean heterozygosity of 78% and a mean PIC (polymorphism information content) of 75%. Polymerase chain reaction (PCR) amplification of markers DNA was extracted from whole blood using a rapid salting out method.37 PCR amplification of markers was carried out using True Allelet Premix (Applied Biosystems) with the manufacturer’s recommended conditions in a final volume of 15 ml on either a GRI Tetrad or a 9700 thermal cycler (Applied Biosystems). PCR products were Genes and Immunity Australian MS linkage screen M Ban et al 468 pooled in predetermined panels prior to electrophoresis on a 3700 Genetic Analyser (Applied Biosystems). Semiautomated genotyping was performed using the GENESCAN/GENOTYPER software (Applied Biosystems). Statistical methods As the mode of inheritance is unknown in multiple sclerosis, a non-parametric multipoint linkage analysis was performed using the MAPMAKER/SIBS program (version 2.0).38 This calculates the MLS at each point on a chromosome, using all the available genotypes, and computes the proportion of information extracted by the markers typed. Areas of potential linkage were identified as positive MLS values taking X0.7 as the nominal 5% significance threshold and X1.8 as suggestive of linkage. Marker allele frequencies were estimated using the SPLINK program (version 1.07).36 As no parents were genotyped in this study, the data were tested with the SIBERROR program (version 1.0)39 prior to linkage analysis in order to exclude half or unrelated siblings. Acknowledgements We thank S Adams, V Perich, N Hartley, M Bugeja, S Surkus and S Teutsch for the collection and preparation of the DNA samples, J Gray for valuable technical support and the Neurologists of Australia for verifying the diagnosis of the patients. This work was supported by the Multiple Sclerosis Society of Great Britain and Northern Ireland (Grant No 659/01), the Wellcome Trust (Grant No 057097/Z/99Z), and the National Health and Medical Research Council of Australia (Grant No 981580 and 153990). References 1 Hogancamp WE, Rodriguez M, Weinshenker BG. The epidemiology of multiple sclerosis. Mayo Clin Proc 1997; 72: 871–878. 2 Sadovnick AD, Armstrong H, Rice GPA et al. A population based study of multiple sclerosis in twins: updated. Ann Neurol 1993; 33: 281–285. 3 Mumford CJ, Wood NW, Kellar-Wood HF et al. The British Isles survey of multiple sclerosis in twins. Neurology 1994; 44: 11–15. 4 Sadovnick AD, Ebers GC, Dyment DA, Risch NJ, the Canadian Collaborative Study Group. Evidence for genetic basis of multiple sclerosis. Lancet 1996; 347: 1728–1731. 5 Ebers GC, Sadovnick AD, Risch NJ, The Canadian Collaborative Study Group. A genetic basis for familial aggregation in multiple sclerosis. Nature 1995; 377: 150–151. 6 Dean G, McLoughlin H, Brady R, Adelstein A, TallettWilliams J. Multiple sclerosis among immigrants in Greater London. BMJ 1976; 1: 861–864. 7 Hammond SR, de Wytt C, Maxwell IC et al. The epidemiology of multiple sclerosis in Queensland, Australia. J Neurol Sci 1987; 80: 185–204. 8 Hammond SR, McLeod JG, Millingen KS et al. The epidemiology of multiple sclerosis in three Australian cities: Perth, Newcastle and Hobart. Brain 1988; 111: 1–25. 9 Sawcer SJ, Jones HB, Feakes R et al. A genome screen in multiple sclerosis reveals susceptibility loci on chromosomes 6p21 and 17q22. Nat Genet 1996; 13: 464–468. Genes and Immunity 10 Haines JL, Ter-Minassian M, Bazyk A et al. A complete genomic screen for multiple sclerosis underscores a role for the major histocompatibility complex. Nat Genet 1996; 13: 469–471. 11 Ebers GC, Kukay K, Bulman DE et al. A full genome search in multiple sclerosis. Nat Genet 1996; 13: 472–476. 12 Kuokkanen S, Gschwend M, Rioux JD et al. Genomewide scan of multiple sclerosis in Finnish multiplex families. Am J Hum Genet 1997; 61: 1379–1387. 13 Åkesson E, Oturai A, Berg J et al. A genome-wide screen for linkage in Nordic sib-pairs with multiple sclerosis 2002 3: 279–285 14 Broadley S, Sawcer S, D’Alfonso S et al. A genome screen for multiple sclerosis in Italian families. Genes Immun 2001; 2: 205–210. 15 Coraddu F, Sawcer S, D’Alfonso S et al. A genome screen for multiple sclerosis in Sardinian multiplex families. Eur J Hum Genet 2001; 9: 621–662. 16 Lander E, Kruglyak L. Genetic dissection of complex traits: guidelines for interpreting and reporting linkage results. Nat Genet 1995; 11: 241–247. 17 Sawcer SJ, Jones HB, Judge D et al. Empirical genomewide significance levels established by whole genome simulations. Genet Epidemiol 1997; 14: 223–229. 18 The Transatlantic Multiple Sclerosis Genetics Cooperative. A meta-analysis of genome screens in multiple sclerosis. Mult Scler 2001; 7: 3–11. 19 Kellar-Wood H, Wood NW, Holmans P et al. Multiple sclerosis and the HLA-D region: linkage and association studies. J Neuroimmunol 1995; 58: 183–190. 20 Stewart GJ, Teutsch SM, Castle M, Heard RN, Bennetts BH. HLA-DR, -DQA1 and -DQB1 associations in Australian multiple sclerosis patients. Eur J Immunogenet 1997; 24: 81–92. 21 Risch N, Merikangas K. The future of genetic studies of complex human diseases. Science 1996; 273: 1516–1517. 22 Feakes R, Sawcer S, Smillie B et al. No evidence for the involvement of Interleukin 2 or the immunoglobulin heavy chain cluster in determining susceptibility to multiple sclerosis. J Neurol Neurosurg Psychiat 2000; 68: 679. 23 Broadley SA, Deans J, Sawcer SJ, Clayton D, Compston DA. Autoimmune disease in first-degree relatives of patients with multiple sclerosis. A UK survey. Brain 2000; 123: 1102–1111. 24 Becker K, Simon R, Bailey-Wilson J et al. Clustering of nonmajor histocompatibility complex susceptibility candidate loci in human autoimmune diseases. Proc Natl Acad Sci USA 1998; 95: 9979–9984. 25 Jawaheer D, Seldin MF, Amos CI et al. A genomewide screen in multiplex rheumatoid arthritis families suggests genetic overlap with other autoimmune diseases. Am J Hum Genet 2001; 68: 927–936. 26 Tsao BP, Cantor RM, Kalunian KC et al. Evidence for linkage of a candidate chromosome 1 region to human systemic lupus erythematosus. J Clin Invest 1997; 99: 725–731. 27 Gaffney P, Kearns G, Shark K et al. A genome-wide search for susceptibility genes in human systemic lupus erythematosus sib-pair families. Proc Natl Acad Sci USA 1998; 95: 14 875–14 879. 28 Moser K, Neas B, Salmon J et al. Genome scan of human systemic lupus erythematosus: evidence for linkage on chromosome 1q in African-American pedigrees. Proc Natl Acad Sci USA 1998; 95: 14 869–14 874. 29 Cho JH, Nicolae DL, Gold LH et al. Identification of novel susceptibility loci for inflammatory bowel disease on chromosomes 1p, 3q, and 4q: evidence for epistasis between 1p and IBD1. Proc Natl Acad Sci USA 1998; 95: 7502–7507. 30 Cornelis F, Faure S, Martinez M et al. New susceptibility locus for rheumatoid arthritis suggested by a genomewide linkage study. Proc Natl Acad Sci USA 1998; 95: 10 746–10 750. Australian MS linkage screen M Ban et al 469 31 Davies JL, Kawaguchi Y, Bennett ST et al. A genome-wide search for human type 1 diabetes susceptibility genes. Nature 1994; 371: 130–136. 32 Cucca F, Goy JV, Kawaguchi Y et al. A male–female bias in type 1 diabetes and linkage to chromosome Xp in MHC HLA-DR3positive patients. Nat Genet 1998; 19: 301–302. 33 Encinas JA, Lees MB, Sobel RA et al. Genetic analysis of susceptibility to experimental autoimmune encephalomyelitis in a cross between SJL/J and B10.S mice. J Immunol 1996; 157: 2186–2192. 34 Encinas JA, Wicker LS, Peterson LB et al. QTL influencing autoimmune diabetes and encephalomyelitis map to a 0.15-cM region containing IL2. Nat Genet 1999; 21: 158–160. 35 Poser CM, Paty DW, Scheinberg L et al. New diagnostic criteria for multiple sclerosis: guidelines for research protocols. Ann Neurol 1983; 13:227–231. 36 Holmans P, Clayton D. Efficiency of typing unaffected relatives in an affected sib-pair linkage study. Am J Hum Genet 1995; 57: 1221–1232. 37 Lahiri DK, Nurnberger Jr JI. A rapid non-enzymatic method for the preparation of HMW DNA from blood for RFLP studies. Nucl Acids Res 1991; 19: 5444. 38 Kruglyak L, Lander ES. Complete multipoint sib-pair analysis of qualitative and quantitative traits. Am J Hum Genet 1995; 57: 439–454. 39 Ehm M, Wagner M. A test statistic to detect errors in sib-pair relationships. Am J Hum Genet 1998; 62: 181–188. Genes and Immunity