* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Tutorial_7 (2016) - Gene Expression
Oncogenomics wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Genetic engineering wikipedia , lookup
Copy-number variation wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
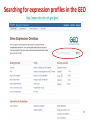
Neuronal ceroid lipofuscinosis wikipedia , lookup
Saethre–Chotzen syndrome wikipedia , lookup
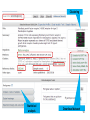
History of genetic engineering wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
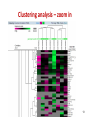
Vectors in gene therapy wikipedia , lookup
Pathogenomics wikipedia , lookup
Long non-coding RNA wikipedia , lookup
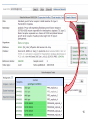
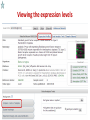
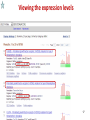
Public health genomics wikipedia , lookup
Genomic imprinting wikipedia , lookup
Gene therapy wikipedia , lookup
Metagenomics wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Gene desert wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Genome (book) wikipedia , lookup
Gene nomenclature wikipedia , lookup
Ridge (biology) wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Genome evolution wikipedia , lookup
Mir-92 microRNA precursor family wikipedia , lookup
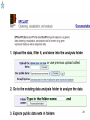
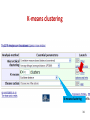
Therapeutic gene modulation wikipedia , lookup
Nutriepigenomics wikipedia , lookup
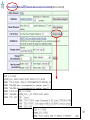
Microevolution wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Designer baby wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Tutorial 8 Gene expression analysis 1 Gene expression analysis • How to interpret an expression matrix • Expression data DBs - GEO • Clustering – Hierarchical clustering – K-means clustering – Tools for clustering - EPCLUST • Functional analysis – Go annotation – DAVID 2 Gene expression data sources Microarrays RNA-seq experiments 3 How to interpret an expression data matrix Sample 1 Sample 2 Sample 3 Sample 4 Sample 5 Sample 6 Gene 1 -1.2 -2.1 -3 -1.5 1.8 2.9 Gene 2 2.7 0.2 -1.1 1.6 -2.2 -1.7 Gene 3 -2.5 1.5 -0.1 -1.1 -1 0.1 Gene 4 2.9 2.6 2.5 -2.3 -0.1 -2.3 Gene 5 0.1 1.9 2.6 2.2 2.7 -2.1 Gene 6 -2.9 -1.9 -2.4 -0.1 -1.9 2.9 • Each column represents all the gene expression levels from a single sample. • Each row represents the expression of a gene across all experiments. 4 Raw data pre-processing Raw data – the data values that we get from the microarray/ sequencer. Raw values are a general term used for the raw measurements made by an instrument. In microarrays the raw data is probe intensities. In sequencing the raw data is counts per gene. • Raw data will almost always need to undergo some kind of processing in order to be in adequate quality and have a biological meaning. – For example high throughput sequencing raw data are the sequenced reads. They need to get mapped to the genome, possibly filtered, and then variant calling is done. 5 Expression profiles DBs • GEO (Gene Expression Omnibus) http://www.ncbi.nlm.nih.gov/geo/ • Human genome browser http://genome.ucsc.edu/ • ArrayExpress http://www.ebi.ac.uk/arrayexpress/ 6 The current rate of submission and processing is over 10,000 samples per month. In 2002 Nature journals announce requirement for microarray data deposit to public databases. 7 Searching for expression profiles in the GEO http://www.ncbi.nlm.nih.gov/geo/ 8 GEO accession IDs GPL**** - platform ID GSM**** - sample ID GSE**** - series ID GDS**** - dataset ID • A Series record defines a set of related samples considered to be part of a group. • A GDS record represents a collection of biologically and statistically comparable GEO samples. Not every experiment has a GDS. 9 Clustering Statistical analysis Download dataset 10 Raw data (soft file) ... ... ... ... Probes Genes Expression values per sample (GSM) Gene annotations 11 Clustering analysis Zoom in 12 Clustering analysis – zoom in 13 Clustering analysis – zoom in 14 15 Viewing the expression levels 16 Viewing the expression levels 17 18 Clustering Grouping together genes with a similar signature 19 Hierarchical Clustering This clustering method is based on distances between expression profiles of different genes. Genes with similar expression patterns are grouped together. 20 Rings a bell?... • In both phylogenetic trees and in clustering we create a tree based on distance matrix. • When computing phylogenetic trees: We compute distances between sequences. • When computing clustering dendograms we compute distances between expression values. ATCTGTCCGCTCG ATGTGTGCGCTTG Score Expr.1 Expr.2 Expr.3 Expr.4 Expr.5 Expr.6 Gene 1 Gene 2 Score 21 Hierarchical clustering methods produce a tree or a dendrogram. They avoid specifying how many clusters are appropriate. The partitions are obtained from cutting the tree at different levels. 2 clusters 4 clusters 6 clusters 22 The more clusters you want the higher the similarity is within each cluster. http://discoveryexhibition.org/pmwiki.php23 /Entries/Seo2009 Hierarchical clustering results You can cluster both samples and genes (separately) http://www.spandidospublications.com/10.3892/ijo.2012.1644 24 Unsupervised Clustering – K-means clustering An algorithm to classify the data into K number of groups. K=4 25 How does it work? 1 k initial "means" (in this casek=3) are randomly selected from the data set (shown in color). 2 k clusters are created by associating every observation with the nearest mean 3 4 The centroid of each of the k clusters becomes the new means. Steps 2 and 3 are repeated until convergence has been reached. The algorithm iteratively divides the genes into K groups and calculates the center of each group. The results are the optimal groups (center distances) for K clusters. 26 How should we determine K? • Trial and error • Take K as square root of gene number 27 Tool for clustering - EPclust http://www.bioinf.ebc.ee/EP/EP/EPCLUST/ 28 29 Choose distance metric Choose algorithm 30 Hierarchical clustering 31 Zoom in by clicking on the nodes 32 33 K-means clustering K-means clustering 34 Samples found in cluster Graphical representation of the cluster Graphical representation of the cluster 35 10 clusters, as requested 36 Now what? Now that we have clusters – we want to know what is the function of each group. There is a need for some kind of generalization for gene functions. 37 Gene Ontology (GO) http://www.geneontology.org/ The Gene Ontology project provides an ontology of defined terms representing gene product properties. The ontology covers three domains: • Biological process • Cellular component • Molecular function Gene Ontology (GO) Cellular Component (CC) - the parts of a cell or its extracellular environment. Molecular Function (MF) - the elemental activities of a gene product at the molecular level, such as binding or catalysis. Biological Process (BP) - operations or sets of molecular events with a defined beginning and end, pertinent to the functioning of integrated living units: cells, tissues, organs, and organisms. 39 The GO tree – a partial example DAVID http://david.abcc.ncifcrf.gov/ Functional Annotation Bioinformatics Microarray Analysis • Identify enriched biological themes, particularly GO terms • Discover enriched functional-related gene/protein groups annotation ID conversion Functional annotation - upload Gene list you want to explore (for example all the genes in a certain cluster) What is the identifier? (probes/ gene names/ gene IDs) You can supply a background list as well 44 Functional annotation - results Different kinds of enrichments are calculated 45 Functional annotation - results Charts for each category Genes from your list involved in this category Minimum number of genes for corresponding term Source of term Enriched terms associated with your genes Maximum EASE score/ E-value Genes from your list involved in this category P-Value Adjusted P-Value Gene expression analysis • How to interpret an expression matrix • Expression data DBs - GEO • Clustering – Hierarchical clustering – K-means clustering – Tools for clustering - EPCLUST • Functional analysis – Go annotation – DAVID 48