* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download MS1 MolBio Genetics Outline
Oncogenomics wikipedia , lookup
Fetal origins hypothesis wikipedia , lookup
Minimal genome wikipedia , lookup
Tay–Sachs disease wikipedia , lookup
Gene expression programming wikipedia , lookup
Frameshift mutation wikipedia , lookup
Polymorphism (biology) wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Gene expression profiling wikipedia , lookup
Behavioural genetics wikipedia , lookup
Genetic engineering wikipedia , lookup
Hardy–Weinberg principle wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Genome evolution wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Human genetic variation wikipedia , lookup
History of genetic engineering wikipedia , lookup
Skewed X-inactivation wikipedia , lookup
Neuronal ceroid lipofuscinosis wikipedia , lookup
X-inactivation wikipedia , lookup
Medical genetics wikipedia , lookup
Genomic imprinting wikipedia , lookup
Point mutation wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Genetic drift wikipedia , lookup
Dominance (genetics) wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Designer baby wikipedia , lookup
Population genetics wikipedia , lookup
Genome (book) wikipedia , lookup
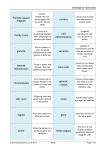
Molecular Biology & Genetics Final Exam Genetics PATTERNS OF INHERITANCE IN HUMAN DISEASE Why define patterns of inheritance: Crucial to making a diagnosis Provides critical information to families about their future Provides insight into the fundamental etiology of disease (for better treatment) 1. Autosomal Dominant Inheritance caused by genes on autosomes, so affects males and females exceptions due to sex limited diseases traits (e.g., ovarian cancer) mutant alleles dominant to wild-type, disorder manifests in heterozygote vertical transmission involving several generations exceptions can occur due to decreased penetrance or new mutations risk to each child of affected individual = 50% Penetrance: not everyone who inherits mutant gene will manifest disease at all or not until later in life (age-related penetrance) Definition: proportion of carriers who demonstrate disease New mutations: individual may be affected by an AD due to new mutation, in which case parents are disease free More common in larger genes, mutational hot spots Variable expressivity: degree to which a phenotype is expressed in an individual Most AD diseases demonstrate variable expressivity Examples: Huntington Disease Progressive degenerative neurological disease inherited in AD fashion (gene on short arm of chromosome 4) Average age of onset is after 38 years 1st human disease mapped with use of DNA linkage analysis Marfan’s Disease Highly penetrant AD, long limbs, skeletal abnormalities, aortic aneurysms Genetic Anticipation: Phenomenon in which the severity of an inherited disorder increases as it is inherited through successive generations May require that the mutation pass through a female (myotonic dystrophy) or male (Huntington) Example: in Huntington, anticipation (not transmission) requires transmission via a male (through trinucleotide expansion) Mechanisms of Dominant Disease Haploinsufficiency: usually having half the amount of a given gene product is sufficient, but in some situations this decrease results in disease; relatively unusual mechanism Increased gene dosage Promoter effects: a mutation in the promoter region may cause it to be over expressed in the wrong tissue or at the wrong time Mutations that result in abnormally increased protein activity Dominant negative mutations: many gene products act as heterodimers or homodimers, a mutant protein will interfere with the action of the normal protein Trinucleotide repeats: under certain circumstances trinucleotide repeats expand during DNA replication, which can result in abnormalities of RNA processing or an abnormal gene product Huntington CAG repeat: in 1st exon, normal number is less than 36, affected individuals have more than 36, more repeats = earlier age of onset, underlies mechanism of anticipation: CAG tract expands only during male meiosis 2. Autosomal Recessive Inheritance AR disorders or traits are caused by genes located on the autosomes, so affect males and females Disease allele recessive to wild-type allele, not evident in heterozygous state Typically confined to a single sibship = horizontal pedigree Increased rate of consanguinity in parents Importance of ethnicity: In any population that has been through population bottlenecks or has had high rates of intermarriage, certain AR diseases can be found at much higher rates (Ashkenazi Jews, Finns, French Canadians) Due to founder effect Mechanisms of AR inheritance: Much simpler than Ad Most mutations result in inactive/weakly active protein Since we are diploid, wild-type allele is usually sufficient, no phenotype manifests Thus, most mutations are recessive and exhibit no phenotype Risk calculation: risk of two heterozygotes having affected offspring = ¼ risk that a phenotypically normal offspring is a carrier = ⅔ Genetic Heterogeneity: clinically identical diseases may result from mutations in different genes, found in both AD and AR diseases (e.g., polycystic kidney disease, deafness) X LINKED AND NON-MENDELIAN PATTERNS OF INHERITANCE 1. X-linked Transmission X chromosome is large and has many genes, Y is small w/ few functional genes (none have been demonstrated to cause disease when mutated – can’t pass on infertility) Disease incidence is much greater in males than in females (carriers rarely affected) Male to male transmission is never seen All daughters of an affected male are carriers Hemophilia A: X-linked, affects boys only, results from a lack of Factor VII, residing on X chromosome, can be treated with Factor VII transfusions Example case: VR has nephew and uncle with hemophilia, so her mother is carrier, she has 50% chance of being carrier, each of her children has 25% chance of inheriting gene, her brother’s children: all females carriers, none of sons have gene X inactivation: mechanism that corrects for the potential problems related to gene dosage effects Gene dosage is important (e.g., trisomy 21), creates a potential problem given the inheritance of sex chromosomes X inactivation explains how males/females adjust for their differing number of X chromosome-associated genes Lyon Hypothesis (Mary Lyon, 1960): In somatic cells of females, only one chromosome is active (i.e., with active gene transcription), the other is condensed and inactive (barr body) Inactivation occurs early in life (morula stage) and is random (can be either maternal or paternal copy that is inactivated) Females are mosaics due to random X inactivation Provides explanation of why female carriers are occasionally affected by an X-linked disease (e.g., 8% of female heterozygotes for X-linked muscular dystrophy are affected, some women are red-green color blind) Skewed X Inactivation: if at the time of random X inactivation there are 10 cells that will ultimately give rise to an organ, the chances that all will inactivate the same X chromosome are (½)10, or about 1/1000 The observed frequency of affected females in X-linked recessive diseases has been used for rough calculations of the number of progenitor cells in the embryo that gave rise to a particular organ X Linked Dominant Inheritance Daughters of affected males will always inherit the disorder, but may be less severely affected due to random X inactivation Sons of affected males never inherit the disorder Affected females will transmit the mutation to 50% of their offspring Very rare (e.g., vitamin D resistance rickets) 2. Y-Linked Inheritance Refers to genes carried only on the Y chromosome No evidence for Y-linked disease in humans But: this is changing with new fertility techniques, it is now possible to pass on infertility 3. Non-Mendelian Inheritance Mitochondrial Inheritance: aka cytoplasmic or maternal inheritance Mitochondria have their own genome (16,710bp) mostly involved in ox phos higher rate of mutation than nuclear genome many polymorphisms between individuals and among populations Each cell (except RBCs) has hundreds of mitochondria, each mitochondrium has 5-10 copies of its own genome During mitosis a cell’s mitochondria are randomly partitioned, so a zygotes mitochondria are completely derived from mom A number of diseases result from mitochondrial mutations, and they generally involve muscles and the CNS (b/c of these tissues’ high energy demand and reliance on mitochondria for energy production) (e.g., MERRF – mitochondrial encephalopathy with ragged red fibers) All or most of an affected female’s children will be affected, none of an affected females will be affected, Variable expressivity is the rule: due to the randome proportion of diseased mitochondria that make it into a given ovum Exclusive maternal inheritance of the mitochondrial genome allows tracking of maternal lineage over long periods of time (similar studies on Y chromosome to track male lineage) Germline Mosaicism Mosaic: tissue, organ, or individual derived from a single zygote that is comprised of two populations of cells that differ in their genotype (females are mosaic due to random X inactivation) If a mutation occurs in a cell which will give rise to a subpopulation of germ cells this has implications for inheritance progenitor cell destined to become gonad acquires a mutation, then the final organ (testicle or ovary) will have a proportion of its cells carrying this mutation Even though this is a new mutation and neither parent has the disorder (so are counseled recurrence risk is low), due to germline mosaicism, recurrence risk is actually high Mutational analysis is possible in sperm, but not in ovaries Imprinting An assumption of Mendelian Inheritance is that the alleles of a given gene from both parents are expressed equally in offspring (not always the case!) Certain genes and chromosomal regions appear to be reversibly modified in parental gametes so that in offspring, maternally and paternally inherited alleles are expressed differently (5% of human genome) Imprinted alleles are inherited in a Mendelian fashion, but their expression is determined by the sex of the transmitting parent Examples: Prader-Willi: 70% of cases result from deletions of long arm of chromosome 15, always a deletion of the paternally derived chromosome 15, paternally derived allele is active while maternally derived allele has been inactivated/imprinted Only paternally derived PWS region of chromosome 15 is transcriptionally active, so if a male has inherited a chromosome 15 with a deletion in the PWS region from his mom, he is not affected, but his offspring will Angelman: 80% of cases due to deletion of nearby region of chromosome 15, always a deletion of the maternally derived chromosome 15, maternally derived allele is active while paternally derived allele has been inactivated/imprinted PWS and Angelman are oppositely imprinted Mechanisms of imprinting: Uniparental disomy: rarely, an individual will inherit both copies of a particular chromosome from only one parent If both copies of chromosome 15 are inherited from mom, PWS will result (b/c both are imprinted, it’s as if the paternal PWS region were deleted) Some cases of AS are a result of paternal uniparental disomy Must occur before fertilization Must be able to confer transcriptional silencing Must be stably transmitted through mitosis Must be reversible upon passage through the opposite parental germline (resetting) Methylation: DNA methylation of CpG dinucleotides in the promoter region of genes can be a mechanism of transcriptional silencing POPULATION GENETICS Definition: study of the distribution of alleles in populations/ethnic groups and of how the frequency of alleles and genotypes are maintained or changed Hardy-Weinberg Equilibrium: underlying principle of population genetics Ethnic groups arose from small, scattered populations early in human history that became geographically isolated, leading to genetic isolation. Over time, selection of favorable mutations in response to environmental conditions, social selection, or the chance survival of neutral mutations, led to variation in allele frequencies between ethnic groups and geographically isolated populations ALLELE/DISEASE Sickle cell anemia (BS allele of globin gene) Cystic Fibrosis Tay-Sachs disease Monotonic dystrophy ABO blood group Alcohol dehydrogenase Alpha1-antitrypsin Lactase activity (lactose intolerance) POPULATION VARIATION 1/20 in African Americans <1/200 in Hispanics 1/40-50 in European Americans Very low in Asian and African Americans 1/60 in Ashkenazi Jews <1/6,000 in other groups 1/50,000 in Europeans Non-existent in Africans 1/950 in regions of Quebec B allele common in Asians B allele absent in Native Americans Frequency of major alleles vary between populations; M1 from 0.51 to 0.98 and M2 from 0 to 0.26 1/60 in Ashkenazi Jews <1/6,000 in other groups Two major alleles – high and low activity Low activity 0.8 – 0.95 in Africans & Asians Low activity 0.17 – 0.48 in Northern Europeans Neutral allele frequencies are important markers for recent human evolution and migration/population studies Allele frequencies: Allele frequency of A is p Allele frequency of a is q p+q=1 Punnett square: all of the possible genetic outcomes for a given mating Example of double heterozygous cross Male Allele Frequencies Female Allele Frequencies p p p2 q pq q pq q2 Hardy-Weinberg derived from sum of Punnett square results for all possible matings (all genotypes) Therefore, if alleles are in Hardy-Weinberg equilibrium within a population, the equation p2 + 2pq + q2 = 1 can be used to determine the number of individuals within each genotype For more than two alleles, Hardy-Weinberg collapses to: (p + q)n = 1 with n = number of alleles Assumptions: 1. The population is large and matings are random with respect to the locus in question 2. Allele frequencies remain constant over time because: a. There is no appreciable rate of mutation b. Individuals with all genotypes are equally capable of mating and passing on their genes (i.e., no selection against any particular genotype) c. There has been no significant immigration of individuals from a population with allele frequencies very different from the endogenous population Exceptions to Random Mating: all have in common the effect of increasing the frequency of homozygous genotypes (and thus increasing the chances of recessive diseases) Stratification: when a population contains a number of subgroups that have remained, for the most part, genetically distinct Assortative mating: the choice of a mate with similar characteristics (positive assertive mating) or different characteristics (negative assortative mating) Consanguinity: selection of a related mate Exceptions to Constant Allele Frequency: usually slow and in small increments Mutation: production of new alleles Selection: works in concert with mutation or environmental changes. For example, can have selection against dominant or recessive diseases or for heterozygotes (e.g., BS allele causing sickle cell anemia) Genetic drift: fluctuation in allele frequency due to chance in small populations, usually random acting Gene flow: slow diffusion of alleles across a barrier (usually associated with migrant populations) THE GENETICS OF COMMON DISEASE Low penetrance, high frequency genes Traditional Mendelian disorders typically result from mutations that have rather high penetrance, such mutations are relatively unusual (low frequency) The genetic component of common diseases stems from the inheritance of subtle changes in genes that affect their function only slightly (low penetrance), such polymorphisms are thought to be common (high frequency) Mendelian vs. Common Disease Single Genes High Penetrance Polygenic Inheritance Low Penetrance Low frequency alleles Limited evir. Influence Tidy pedigrees Easy to discern genetic role Gene isolation tractable Limited public health impact High frequency alleles Large environmental influence Untidy pedigrees Difficult to discern genetic role Difficult to isolate genes Tremendous impact/cumulative burden Genes interact with environment to result in disease (smokers, alcoholics, TB exposure, head trauma outcome – all highly variable reactions depending on genetics) Polymorphisms: DNA sequence changes that do not destroy protein function but may alter it, are common and contribute to common diseases Underlie our susceptibility to common disease Genetic Approaches to Common Disease Twin Studies MZ twins typically share their environment and 100% of their genes, DZ twins typically share their environment and 50% of their genes, so excess concordance for a trait (disease) in MZ twins is evidence of genetic influence MZ females may be a little less familiar due to random X inactivation Heritability = genetic variance/total variance h' = V of DZ pairs – V of MZ pairs / V of DZ pairs Estimates of heritability from twin studies: (highest lowest) obesity, type II DM, schizophrenia, HTN, alcoholism, cirrhosis, atherosclerosis, type I DM Twin studies are extremely powerful, but offer no clue to precise genetic mechanisms Observational studies of family clustering Confounded by environmental exposures Offer no clue to precise genetic mechanisms Animal studies Allow elucidation of genetic factors important in common disease No guarantee that same genes are responsible for genetic variance in humand and model animal Hypothesis generated studies If biochemical information is present, one can examine polymorphisms of candidate genes to see if they influence disease (e.g., polymorphisms in angiotensinogen gene found more often in hypertensive individuals than controls) Require biochemical knowledge Sib Pair Analysis Requires no assumptions about mode of inheritance Requires no knowledge of biochemistry Based on assumption that sibs share, on average, 50% of their alleles If a marker is linked to a disease-susceptibility locus affected sibs will more often than not share that allele Requirements include genomic saturation of polymorphic markers, large wellcharacterized sets of sibs with disease in question Quantitative Trait Loci (QTL) A method that synthesizes Sib Pair analysis with the fact that many diseases and traits are quantitative By picking extremes of affected individuals (e.g., top 5%), it may be possible to decrease genetic heterogeneity and increase the power of the study Inherent weakness: immense size of human genome, isolation of genes difficult due to large # of genes in a linked region, each gene is likely to contribute little to the disease, so confirmation of gene’s influence is epidemiological Susceptibility to disease in different populations may be explained by polymorphisms in different genes An identified polymorphism will be neither necessary nor sufficient for acquisition of the disease in question Alzheimer’s Disease Death typically occurs 8-10 years post dx, treatments are inadequate Most common cause of dementia in North America and Europe 10% of people >70 have significant memory loss, >½ of those have AD Due to decreased longevity, AD represents a significant medical, economic, and social problem Genetics: 75% is sporadic (negative family history) 25% is familial (but indistinguishable from sporadic cases by clinical, pathological, or neuroanatomical means) small subset of familial cases (<5% of all AD cases) demonstrate dominant inheritance and early onset AD represents 1st disease for which we have detailed information regarding low frequency/high penetrance determining genes and high frequency/low penetrance susceptibility genes Apoprotein E (apo E) Studied for years as a lipid transport protein, but ensuing biochemical studies implicated ApoE in neuronal and glial function Sib-pair analysis suggested in 1992 that ApoE polymorphisms influence age of onset and susceptibility to AD Polymorphisms consist of single aa substitutions, fit definition of high frequency/low penetrance genes 3 polymorphisms: E2, E3, E4 6 possible genotypes E4 predisposes to AD, resulting in earlier age of onset (dose-response risk gene), but E4 allele is neither necessary nor sufficient for development of AD E2 appears protective Does ApoE mediate a life-long process which culminates in AD? (i.e., will everyone who lives to age 140 develop it?) ApoE appears to account for >50% of genetic risk (E4/E4 genotype has odds ratio of 30) Susceptibility testing for AD In classic Mendelian disorders (e.g., HD, CF, rare forms of AD) testing is straightforward with virtual 100% ppv and 100%npv (genetics is destiny) In common diseases with genes that predispose but don’t dictate, prognostication is risky at best 25 – 30% of autopsy-confirmed Ad lack E4 allele 5% of centenarians carry an E4 allele without AD predictive testing is not useful presently, would change if effective intervention were found AIDS Certain individuals seem resistant to infection with HIV in spite of dramatic exposure histories Some individuals with HIV progress rapidly to clinical AIDS, others much more slowly The mechanism of entry into the cell by HIV is via co-receptors, CD4 and CCR5 An alteration in CCR5, delta-32, is found in 1-2% of the Caucasian population Obvious implications for drug design Diabetes Mellitus Prevalence: will be highest in Asia, Africa, South America higher in men than women, increases with age prevalence increased 30% between 1990 and 1998 increased 40% ages 40-49 increased 70% ages 30-39 6% annual increase over last 5 years WHO: world incidence will more than double over next 25 years, affecting an estimated 300 million individuals Incidence: 700,000 new cases annually Impact: Leading cause of adult blindness, renal failure and non-traumatic amputations Seventh leading cause of death Approximately 17% of all health care expenditures ($105 billion annually) Type I and Type II are distinct diseases Type I Onset typically <30 Ketosis prone Absolutely insulin deficient Islet cell antibodies Other autoimmune dz HLA considerations MZ concordance <50% Type II Onset typically >30 Ketosis resistant Variable None No associatkion None >90% Genetics of Type I Genetic factors account for 25% susceptibility Environmental exposures are critical Population risk in Japan is 15x less than US, but risk for a MZ twin is similar Highest known incidence is in Finland and Sardinia HLA analysis has revealed a strong influence of one’s HLA genotype and disease susceptibility (speculation regarding an infectious trigger) Increased frequency of Class II HLA DR3 and DR4 among Caucasian Type I patients (95% of patients vs. 50% of controls) Almost 40% of general population carries high-risk alleles for DMI Identity of trigger remains elusive Non-HLA genes likely account for 50% of risk Candidate approach identified IDDM2 on 11p (polymorphic VNTR 5’ to insulin gene near regulatory region) Sib-pair analysis has identified 10 other putative loci Genetics of Type II Clash of evolutionary past and cultural present DM II represents worldwide epidemic, involving non-NA/European populations as “urbanization” occurs Pacific island of Nauru, Pima Native Americans Environmental changes on a non-evolutionary timescale (obesity, inactivity, and dietary changes are of critical importance) MZ concordance for type II approaches 100% (increase as mean weight and inactivity have increased) A number of mongenic causes for DMII have been described, are of interest, but unimportant from a public health standpoint Insulin receptor gene PPAR and other transcription factors Mutant insulin Mitochondrial mutations in a tRNA gene Mutations in glucokinase Candidate approach has not been fruitful polymorphisms found in genes that control glucose regulation, few have been found that confer risk of DM Sib Pair analysis has implicated polymorphisms in calpain-10 (CAPN10) in DM II susceptibility In calpain-like cysteine protease family Function is unclear Ubiquitously expressed Polymorphisms which reduce expression are associated with DMII susceptibility What will we do with our knowledge about the genetics of common disease? Determine an individual’s susceptibility pre-symptomatically (avoidance of environmental triggers and chemoprevention) Guide treatment (pharmacogenomics) Provide basic knowledge for new therapies GENETICS, MEDICINE, AND SOCIETY How do predictive genetic tests differ from conventional medical tests? Genetic tests affect other individuals who have not chosen to undergo testing Conventional medical tests inform us about the patient’s present condition, while genetic tests inform us about a possible future condition (adding a new dimension of uncertainty and affecting social attitudes) Our genome cannot be changed in a meaningful way (great concern over whether it should be changed if it were even possible) Genetic testing touches upon concerns related to the underlying essence of a person’s uniqueness (DNA R US)