* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download ppt
Epigenetics of human development wikipedia , lookup
Adaptive evolution in the human genome wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Human genome wikipedia , lookup
Pathogenomics wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Metagenomics wikipedia , lookup
Non-coding DNA wikipedia , lookup
Gene expression profiling wikipedia , lookup
Oncogenomics wikipedia , lookup
Expanded genetic code wikipedia , lookup
Group selection wikipedia , lookup
Protein moonlighting wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Koinophilia wikipedia , lookup
Population genetics wikipedia , lookup
Minimal genome wikipedia , lookup
Frameshift mutation wikipedia , lookup
Helitron (biology) wikipedia , lookup
Genome evolution wikipedia , lookup
Microevolution wikipedia , lookup
Genetic code wikipedia , lookup
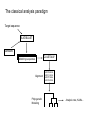
Genome evolution: a sequence-centric approach Lecture 10: Selection in protein coding genes (Probability, Calculus/Matrix theory, some graph theory, some statistics) Simple Tree Models HMMs and variants PhyloHMM,DBN Context-aware MM Factor Graphs DP Sampling Variational apx. LBP EM Generalized EM (optimize free energy) Probabilistic models Genome structure Inference Mutations Parameter estimation Population Inferring Selection Today refs: See Gruer/Li chapter 3/4, and papers we cite Tree of life Genome Size Elements of genome structure Elements of genomic information Models for populations Drift Selection and fixation Draft Protein genes: codes and structure Degenerate code Recombination easier? 1 2 3 3’ utr codons 5’ utr Conformation Epistasis: fitness correlation between two remote loci Domains Intron/exons Identifying protein coding genes From mRNAs Spliced ESTs : short low quality fragments that are easier to get Identifying protein coding genes Computationally identify protein coding genes using probabilistic models is a common genome annotation approach Comparative/Conservation based methods are becoming increasingly popular Questions on protein function and evolution Identification: • Identify protein coding genes – Not yet resolved for new species, but with new technology this question may be less important than before Structure/Function: • Define functional domains – Highly important for understanding protein function • Which parts of the proteins are “important” (e.g., catalytic?) – Difficult since structural modeling is hard and context dependent Evolution • Identify places and times where a new protein feature emerged – Positive selection • • Understand mutation/selection through codon degeneracy Understanding processes of duplication and diversitification The classical analysis paradigm Target sequence BLAT/BLAST Genbank CLUSTALW Matching sequences Alignment Phylogenetic Modeling ACGTACAGA ACGT--CAGA ACGTTCAGA ACGTACGGA Analysis: rate, Ka/Ks… Two words on sequence searches • • A huge database (1011 nt and growing) Need to efficiently find homologous sequence – Homologous – given a distance metric – Matches may include gaps • All practical approaches use some kind of hashing/indexing Query Break to k-mers index Huge Database AAAAA -> positions AAAAC -> positions … … Use genomic hits as starting points Dynamic programming to extend them DB BLAST – the workhorse of genbank for more than a decade BLAT – even simpler and faster, UCSC browser Query Clustalw and multiple alignment • ClustalW is the semi-standard multiple alignment algorithm when sequences are relatively diverged and there are many of them ClustalW Compute pairwise sequence distances (using pairwise alignment) Build a guide-tree: approximating the phylogenetic relations between the sequences “Progressive” alignment on the guide tree • S1 S2 S4 S3 Distance Matrix S5 Neighbor Joining Dist(s1,s2) = best pair align Guide tree is based on pairwise analysis! From the leafs upwards: Align two children given their “profiles” Several heuristics for gaps Other methods are used to multi-align entire genomes, especially when one well annotated model genome is compared to several similar species. Think of using one genome as a “scaffold” for all other sequences. Nucleotide substitution models • For nucleotides, fewer parameters are needed: A a a a C a G a T Jukes-Kantor (JK) • A a b b C a G b T Kimura But this is ignoring all we know on the properties of amino-acids! Simple phylogenetic modeling: PAM/BLOSSOM62 • Given a multiple alignment (of protein coding DNA) we can convert the DNA to proteins. We can then try to model the phylogenetic relations between the proteins using a fixed rate matrix Q, some phylogeney T and branch lengths ti When modeling hundreds/thousands amino acid sequences, we cannot learn from the data the rate matrix (20x20 parameters!) AND the branch lengths AND the phylogeny. Based on surveys of high quality aligned proteins, Margaret Dayhoff and colleuges generated the famous PAM (Point Accepted mutations): PAM1 is for 1% substitution probability. Using conserved aligned blocks, Henikoff and Henikoff generated the BLOSUM family of matrices. Henikoff approach improved analysis of distantly related proteins, and is based on more sequence (lots of conserved blocks), but filtering away highly conserved positions (BLOSUM62 filter anything that is more than 62% conserved) Universal amino-acid substitution rates? “We compared sets of orthologous proteins encoded by triplets of closely related genomes from 15 taxa representing all three domains of life (Bacteria, Archaea and Eukaryota), and used phylogenies to polarize amino acid substitutions. Cys, Met, His, Ser and Phe accrue in at least 14 taxa, whereas Pro, Ala, Glu and Gly are consistently lost. The same nine amino acids are currently accrued or lost in human proteins, as shown by analysis of nonsynonymous single-nucleotide polymorphisms. All amino acids with declining frequencies are thought to be among the first incorporated into the genetic code; conversely, all amino acids with increasing frequencies, except Ser, were probably recruited late. Thus, expansion of initially under-represented amino acids, which began over 3,400 million years ago, apparently continues to this day. “ Jordan et al., Nature 2005 Phylogenetic modeling: ML • Given a rate matrix and an alignment, a phylogenetic tree and branch lengths are estimated to maximize the likelihood. • Frequently the ML gene tree is NOT the ML/fossil based species tree – due to border line statistics – Or due to real lateral effects/duplications problems • Adding rate variation may be critical for elucidating protein function! • Finding the ML model is notoriously difficult: – Estimating the ML branch lengths given a tree can be done using EM – Estimating the optimal tree topology can be done in several ways, some using sampling techniques we learned ML Phylogeny using sampling • Develop a metropolis algorithm to sample from Tree topologies and branch lengths • See how easy it is to use Sampling: • Define a proposal distribution T,t -> T’,t’ • You accept with P ~ P(T’,t’|S)/P(T,t’|S) = Pr(S|T’,t)/Pr(S|T,t) • Collect samples after some burn in period to compute relevant features probabilities • Sampling can be used to ask different question on the tree: • What is the probability of a set of species being mono-phyletic • Chance of particular branch point • And of course, what is the optimal (MAP) Tree • We can tune branch lengths further using EM. • Critical to the success of the Metropolis algorithm is the proposal distribution we are using – Using something completely random is not a good idea (we will be stuck in relatively good positions for a long time) – Too small changes are also problematic Analysis: rate variation • • • If our ML model include rate variation, we can use the inferred rates to annotate the protein Same can be done by constructing a conservation profile, even if the model is simplistic. Shown here are example from Tal Pupko’s work on the Rate4Site and ConSurf programs How prevalent are compensatory mutations? PDB structures Homology modelling Pairs of interacting residues 3-Alignments Rat Mouse Human Find pairs of mutations in interacting residues (DRIP) Coupled: occurring in the same lineage Uncoupled: occurring in different lineages Choi et al, Nat Genet 2005 Synonymous vs. non synonymous mutations • • • The structure of the genetic code is a powerful tool in characterizing evolutionary trends This is because at degenerate positions of codon we can deterministically distinguish between mutations that change the protein and mutation that do not This provide us with a powerful “internal control” – we are comparing two different types of evolutionary events at the same loci, so all sources of variation in the mutational process are not affecting us. Given aligned proteins, we can count: MA – number of non-synonymous changes Ms – number of synonymous changes We then want to estimate: • Ka – rate of non-synonymous mutations (per syn site) • Ks – rate of synonymous mutations (per syn site) • Estimate V(Ka), V(Ks) • Comparing Ka and Ks can provide evolutionary insights: – – – Ka/Ks<<1: negative selection may be purging protein modifying mutations Ks/Ka>>1: positive selection may help acquiring a new function (statistics using, e.g., T-test) Chi-square test Non-Syn Syn Change MA MS No Change NA-MA NS-MS Average number of sites From dN/dS to Ka and Ks • • Classically, we start by looking at noncoding sequence and a one-parameter substitution model In a first approximation K = d/N = p (N being the number of loci), but this ignore multiple mutations • Taking into account the model: (L is the number of loci) 3 4 K ln( 1 p) 4 3 If we assume NA,NS non synonymous/synonymous sites: V (K ) p p2 4 L 1 p 3 2 3 4(dS ) K S ln( 1 ) 4 3N S 3 4(dN ) K A ln( 1 ) 4 3N A A more realistic approach: • classify sites to 0-fold, 2-fold and 4-fold degenerate (consider isoleucine a two fold site) . • For each type one count the number of syn/non-syn transitions or transversions • Express the mean KA, and KS as a function of these counts (Kimura, see Graur/Li book) • How would you do it using methods we learned in the course? Codon bias • • Different codons appears in significantly different frequencies, which is not expected assuming neutrality Bias is measured in several ways, most popular is the codon adaptation index: 1 Xi CAI iL1 X i ,max • L Codon frequency divided by the frequency of the synonymous codon with maximal frequency Possible sources of bias: – Selection for translational efficiency given different tRNA abudnances • Highly expressed genes tend to have stronger codon adaptation indices – Sequence context mutational effects • E.g. CpGs are highly mutable – Selection for low insertion/deletion potential • Weak selection for codon bias should be stronger for genomes with larger effective population size. In some cases this is true Positive selection in humans vs chimp Kn vs Ks Significantly enriched functions/tissues Testis genes: P<0.0001 Immunity genes, Gematogenesis, Olfaction P<1e-5 Inhibition of apoptosis P<0.005 Sensory perception P<0.02 Nielsen et al., 2005 PloS Biol Looking at trends for families of genes Example Mcdonald-Krietman test • • • chimp Works by comparing Ka/Ks divergence between species and Ka/Ks diversity among species populations Negative selection should make the divergence Ka/Ks smaller than the diversity Ka/Ks Positive selection should drive the opposite effect human Busstamente et al, Nature 2005 Codon volatility • Volatility is the number/fraction of adjacent non-synonymous codons • Genes under positive selection may have increased volatility • Think about the distance from the stationary codon distribution • No need to align!! Plotkin et al, Nature 2004 Using extensive polymorphisms and haplotype data, the most recent examples of positive selection: he analysis reveals more than 300 strong candidate regions. Focusing on the strongest 22 regions, we develop a heuristic for scrutinizing these regions to identify candidate targets of selection. In a complementary analysis, we identify 26 nonsynonymous, coding, single nucleotide polymorphisms showing regional evidence of positive selection. Examination of these candidates highlights three cases in which two genes in a common biological process have apparently undergone positive selection in the same population:LARGE and DMD, both related to infection by the Lassa virus3, in West Africa;SLC24A5 and SLC45A2, both involved in skin pigmentation4, 5, in Europe; and EDAR and EDA2R, both involved in development of hair follicles6, in Asia. Sabeti et al, Nature 2007 Time resolution of different positive selection methods Sabeti et al, Science 2005 Molecular clocks and lineage acceleration • How universal is the rate of the evolutionary process? • Mutations may depend on the number of cell division and thus in the length of generation • Mutations depends on the genomic machinery to prevent them (Lynch?) • Mutations may also depend on the environment • The molecular clock (MC) hypothesis state that evolution is working in a similar rate for all lineages Relative rate test: O KOA – KOB = 0 ? Test: KCA – KCB A B C Different molecular clocks in apes and primates Kim et al., 2006 PLoS genet The real challenges ahead • • • • • A model for protein evolution that include multiple selective effects Context-dependent mutations with different molecular clocks Selection for codon bias Epistasis (interaction between positions) Possible additional roles for exon’s DNA We did not talk about it today, but: • Protein coding genes are the outcome of complex duplication/loss process • how different proteins interact and how it affect their evolution?