* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Solid Waste in History
Real-time polymerase chain reaction wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
Community fingerprinting wikipedia , lookup
RNA polymerase II holoenzyme wikipedia , lookup
Molecular cloning wikipedia , lookup
Messenger RNA wikipedia , lookup
Biochemistry wikipedia , lookup
DNA supercoil wikipedia , lookup
Gene regulatory network wikipedia , lookup
Non-coding DNA wikipedia , lookup
Eukaryotic transcription wikipedia , lookup
Microbial metabolism wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Magnetotactic bacteria wikipedia , lookup
Genetic engineering wikipedia , lookup
Point mutation wikipedia , lookup
Transcriptional regulation wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Epitranscriptome wikipedia , lookup
Gene expression wikipedia , lookup
Biosynthesis wikipedia , lookup
Transformation (genetics) wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
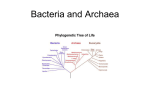
Microbial Ecology and Environmental Genomics The 2nd Week Introductions to - Principles of Microbiology - Molecular Biology of Microorganisms Basics: Microbiology The Cell (living entity) • Growth and self-reproduction • Highly organized and selectively restrict what crosses their boundaries (a lower entropy compared to their environment) • Composed of major elements (C, N, O, and S, in particular) • Self-feeding of elements, electrons, and energy Basics: Microbiology Eukaryotic cell Prokaryotic cell Basics: Microbiology Essential Cell Components • Cell membrane: a barrier between the cell and its environment (selectively transporting elements, electrons, and energy) • Cell wall: a structure member that confers rigidity to the cell and protects the membrane • Cytoplasm: most of the inside of the cell • Chromosome: stores the genetic code for the cell’s heredity and biochemical functions • Ribosomes: convert the genetic code into working catalysts that carry out the cell’s reactions. • Enzymes: biological catalysts Organism Classification Taxonomy • Science of classification • Based upon observable properties (phenotypes) including morphology and transformation • Traditional way of organism classification Phylogeny • Science of classification • Based upon evolution history (small subunit of rRNA, functional gene sequencing, genome sequencing) • New way of organism classification Basics: Microbiology Naming bacteria/archaea • Escherichia coli O157:H7 • Pseudomonas aeroginosa PA01 • Burkholderia xenovorans LB400 RULE: Genus (italic) species (italic) strain (ref. the International Code of Nomenclature of Bacteria) Species: the basic taxonomic unit Genus: population unit Basics: Microbiology Characteristic Membrane-enclosed nucleus Muramic acid in cell wall Chlorophyll-based photosynthesis Methanogenesis Reduction of S to H2S Nitrification Denitrification Nitrogen fixation Synthesis of poly-beta-hydroxylakanoate carbon storage granules Sensitivity to chloramphenicol, streptomycin, and kanamycin Rebosome sensitivity to diphtheria toxin Bacteria Archaea Eukarya Absent Present Yes No Yes Yes Yes Yes Yes Absent Absent No Yes Yes No Yes Yes Yes Present Absent Yes No No No No No No Yes No No No Yes Yes Source: Madigan, Martinko, and Parker, 1997 Phylogenetic tree of life as determined from small subunit of ribosomal RNA sequencing (C. R. Woese) -0.1 Bacteria G+ Proteo- Cyano- Eucarya Mouse Fruit fly Plant HGT -1.0 HGT -2.0 Origin of oxygenic photosynthesis 2.3 Archaea Crenarchaeota 1.0 1.5 Amito -chondriate Euryachaeota 2.1 -3.0 -4.0 Last common ancestor Origin of Earth (4.5 billion years) 3.8 Chemical evolution/ Prebiotic synthesis of biomolecules Basics: Microbiology Environmentally important microorganisms • Bacteria and Archaea (Prokaryotes): detoxification, diseasing-causing, biochemical cycles in nature • Algae (Single-celled Eukaryotes) and Cyanobacteria (Prokaryotes): water quality problem, toxin-producing. • Single celled protozoa (Eukaryotes): bacteria eater, disease-causing • Fungi (multi-cellular Eukaryotes): detoxification Prokaryotes Bacteria Are among the smallest of the entities that are generally agreed to be living. Ubiquitous (everywhere) Able to transform a great variety of inorganic and organic pollutants into harmless minerals (which is recycled back into the environment) => Beneficial to human Often cause disease or are responsible for many of the plagues of the past and for mjor sickness and misery => Threatening human health Archaea (later…) Bacteria Morphology Coccus (spherical shape) Streptococci Staphylococci Sarcina (packets of eight) Bacillus (cylindrical rod shape) Chains of bacilli Spirillum (helical shape) Bacteria Size and some number for bacteria • • • • 0.5-2 μm (width) x 1-5 μm (length) 0.5-5 μm (Diameter for Cocci) 1012 cells per gram of dry solid weight Surface area: 12m2/gram Bacteria Cell structure Cell wall: peptidoglycan (G-negatives have a higher content of lipopolysaccharide while Gpositives teichoic acids) Cytoplasmic membrane: phospholipid bilayer, semipermeable, membrane-bound electrontransport enzymes (cytochromes), selective material transport Cytoplasm: consist of water, dissolved nutrients, enzymes, proteins, and nucleic acids (RNAs and DNAs), and ribosomes (protein-RNA) Inclusion: storage for food or nutrients (e.g. PHB, fatty materials, or sulfur accumlation) DNA: chromosome, plasmid (mobile) RNA: mRNA, tRNA, rRNA Endospores (e.g. Bacillus, stress response) Capsule or slime layer: floc formation Flagella: chemotaxis, phototaxis Fimbriae and pili: attachment, involved in conjugation Bacteria (why C5H7O2N?) Chemical composition Constituent Percentage Water Dry Matter Organic C O H N Inorganic P2O5 K2O Na2O MgO CaO SO3 75 25 90 45-55 22-28 5-7 8-13 10 50 6.5 10 8.5 10 15 Macromolecular composition Macromolecule %a %b Molecules per cell Total Proteins Carbohydrates Lipids DNA RNA 100 50-60 10-15 6-8 3 15-20 100 55 7 9.1 3.1 20.5 24,610,000 2,350,000 22,000,000 2.1 255,500 NOTE a: dry weight (Rittmann and McCarty) b: dry weight, data from E.coli and S. typhimurium [Madigan, Martinko, and Parker (1997) and G. C. Neidhardt et al (1996)] E.coli dry weight for actively growing cells is about 2.8x10-13 g Trace heavy metals in enzymes molybdenum (N2 fixation) nickel (anaerobic methane production) cupper (methane oxidization) Prokaryotic Reproduction Binary fission (normal way of multiplication) Bacteria’s normal way to reproduce themselves. After reproduction, the parent cells no longer exists, and the two daughter cells normally are exact replicates (i.e., clones) of each other, both containing the same genetic information as the parent. (Joon’s question…NO AGING?) Asexual reproduction Replication: chromosome (genomic DNA) is replicated and divided into each daughter cell. Filamentous growth Norcardia species produce Replication of chromosome extensive filamentous growth Formation of long, branching, non-dividing filaments, containing multiple chromosomes. (Multicellular???) In stressed conditions, some of these species form spores (some Streptomyces and many molds) Prokaryotic Reproduction Budding division Asymmetric creation of a growing bud, on the mother cell. The bud increases in size and eventually severed from the parental cell. After division is complete, the mother cell reinitiates the process by growing another bud. Yeast and some bacteria (Caulobacter is one example) use this form of division. Sexual reproduction via conjugation Some bacteria transfer plasmid (not chromosome) into other bacteria using conjugation process (cf. Horizontal gene transfer) Conjugation requires direct contact between two cells. Conjugation results in replication of genetic information. And then multiplication can occur…. Conjugation often occurs between same species as well as between different species (even different genus levels). Conjugation Binary fission Prokaryotic Growth Prokaryotic growth curve Calculation of growth rate Growth rate: dN/dt = k * N (exponential growth) Integration: N2 = N1 * EXP[k*t] Growth rate constant: k = ln(N2/N1)/(t2-t1) here X : biomass or cell number Xo: initial biomass or cell number t2 : 2nd measurement time point t1: 1st measurement time point Example The value of growth rate possibly is influenced by the way of quantifying growth (i.e., cell number counts vs. biomass). Bacteria Energy and carbon-source classes of bacteria Phototrophs (use light as energy source) - Oxygenic phototrophs use light to convert water into O2 and H2, the electron sources. This is similar as plants do, and is dependent the type of chlorophylls. - Anoxygenic phototrophs live in the absence of O2 They use light to extract electron sources from reduced sulfur compounds (H2S), H2 or organic compounds (succinate or butyrate). One example is conversion of H2S into H2 and S. Chemotrophs (use chemicals as energy or carbon sources) - Chemoorganotrophs (organic chemicals) - Chemolithotrophs (inorganic chemicals) - Autotrohs (use inorganic carbon such as CO2 for cell synthesis) - Heterotrophs (use organic carbon for cell synthesis) Bacteria Environmental conditions for growth • Temperature • Psychrophile (-5 to 20oC) Mesophile (8 to 45 oC); Thermophile (40 to 70oC) Hyperthermophile (65 to 110 oC) Aerobes (respiration with oxygen); Anaerobes (respiration in the absence of oxygen); Aerotolerant anaerobes (can grow in the presence of oxygen but cannot use oxygen); Facultative aerobes (do both aerobic and anaerobic respiration); Microaerophiles (can grow in presence of minute quantities of oxygen molecules) • pH Typically, bacteria have a narrow pH range of for growth (6 to 8) For some species, the operating range is quite broad. Acidophilic bacteria (some chemolithotrophs oxidizing sulfur or iron for energy at highly acidic conditions.) • Oxygen Salts Halophiles (grow best under salt conditions similar to seawater, 3.5% NaCl) Extremehalophiles (live well in a saturated NaCl, 15-30%) Bacteria Characteristics of 12 phylogenic lineages of bacteria Aquifer/Hydrogenobacter: Hyperthermophilic, chemolithotrophic Thermotoga: Hyperthermophilic, chemoorganotrophic, fermentative Green nonsulfer bacteria: Thermophilic, phototrophic and nonphototrophic Deinococci Some thermophiles, some radiation resistant, some unique spirochetes Spirochetes: Unique spiral morphology Green sulfur bacteria: Strictly anaerobic, obligately anoxygenic phototrophic Bacteroides-Flavobacteria: Mixture of types, strict aerobes to strict anaerobes, some are gliding bacteria Planctomyces: Some reproduce by budding and lack peptidoglycan in cell walls, aerobic, aquatic, require dilute media Chlamydiae: Obligately intracellular parasites, many cause diseases in humans and other animals. Gram-positive bacteria: Gram-positive, many different types, unique cell-wall composition Cyanobacteria: Oxygenic phototrophic Purple bacteria (Proteobacteria): Gram-negative; many different types including anoxygenic phototrophs and nonphototrophs; aerobic, anaerobic, and facultative; chemoorganotrophic and chemolithotrophic Proteobacteria (purple bacteria) Pseudomonads (belonging to α,β, andγ groups) Pseudomonas, Commamonas, Burkholderia A broad classification of microorganisms important in organic degradation Straight or slightly curved rods with polar flagella. G-negative chemoorganotrophs that show no fermentative metabolism Sulfate Alpha: Rhodospirillum*, Rhodopseudomonas*, Rhodobacter*, Rhodomicrobium*, Rhodovulum*, Rhodopila*, Nitrobacter, Agrobacterium, Aquaspirillum, Hyphomicrobium, Acetobacter, Gluconobacter, Beijerinckia, Paracoccus, Pseudomonas (some species). Beta: Rhodocyclus*, Rhodoferax*, Rubrivivax*, Spirillum, Nitrosomonas, Sphaerotilus, Thiobacillus, Alcaligenes, Pseudomonas, Bordetella, Nesisseria, Zymomonas Gamma: Chromatium*, Thiospirillum*, other purple sulfur bacteria*, Beggiatoa, Leucothrix, Escherichia and other enteric bacteria, Legionella, Azotobacter, fluorescent Pseudomonas species, Vibrio Delta: Myxococcus, Bdellovibrio, Desulfovibrio and other sulfate-reducing bacteria, Desulfuromonas Epsilon: Thiovulum, Wolinella, Campylobacter, Helicobacter * Phototrophic representatives (SOURCE: Madigan, Martinko, and Parker, 1997) TEA Oxygen and nitrate AMD, Corrosion Major grouping of proteobacteria Archaea Archaea versus Bacteria Bacteria generally have peptidoglycan in cell walls but Archaea do not. Bacterial membrane fatty acids tend to be straight chained (ester linkages), while the archaeal membrane lipids tend to be long-chained, branched hydrocarbons (ether linkages). Bacterial RNA polymerase is of single type with a simple quaternary structure, while Archaeal RNA polymerase are of several types and structurally more complex. Meaning of studying Archaea in Biotechnolgy Methanogens (in Euryarchaea group) convert hydrogen and acetate into methane, a useful energy source. Extremophiles (Thermophiles, Halophiles, and Acidophiles) are common in Archaea =>Useful for biological treatment of industrial wastewaters that may contain extremes in salt concentration or temperature Archaea Major groups and subgroups Crenarchaeota: Desulfurococcus, Pyrodictium, Sulfolobus, Thermococcus, Thermoproteus Korarchaeota: Hyperthermophilic Archaea (have not yet been obtained in pure culture) Euryarchaeota: Archaeroglobus, Halobacterim, Halococcus, Halophilic methanogen, Methanobacterium, Methanococcus, Methanosarcina, Methanospirillu, Methanothermus, Methanopyrus, Thermoplama Eukarya Of interest in environmental biotechnology Fungi: (1) the primary decomposers in the world; (2) decompose a great variety of organic materials that tend to resist bacterial decay (decomposition of lignin, leaves, dead plants and trees, and other lignocellulosic organic debris via peroxidase pathways); (3) decomposition of dry organic matter (stabilization of sludge and refuse); (4) favor soil environment, high organic concentration, and drier and more acidic conditions compared to prokaryotes); (5) unfortunately, their detoxification is slow Algae: (1) important in surface water quality control; (2) produce organic matters using light (phytoplankton); (3) oxygenic photosynthesis is good for water quality and wastewater treatment; (4) too much algae growth cause tastes and odors in water supplies, clogging problems in water treatment plants; decreased clarity of lakes; increased sedimentation in lake; (5) a balanced population of algae is required. Protozoa: (1) common members in aerobic and anaerobic wastewater treatments; (2) also are observed in most freshwater and marine habitats; (3) feed on bacteria and small organic particulate matter (polishing effluent from wastewater treatment plants); (4) Indicate the presence of toxic materials Multicellular microscopic Eukarya: rotifers, nematodes, and other zooplankton Viruses Major characteristics Not considered to be “living” entities Replicated only when in association with a living cell Consisting of nucleic acid (DNA or RNA) surrounded by protein 15-300 nm (Smallpox 200-300 nm; Herpes simplex 100 nm; Influenza 100 nm; Adonovirus 75nm; Bacteriophase 80nm; Tobacco mosiac virus 15 x 280 nm) Bacteriophages: virus infects prokaryotes Phages are prevalent in biological wastewater treatment systems A virus infection occurs quite rapidly (within about 25 min, 200 new phases can be produced.) Infectious Diseases Microorganism Class Group Organism name Virus Viscerotropic Coxsackie virus Norwalk virus Rotavirus, Echovirus Hepatitis A virus Polio virus Campylobacter jejuni Helicobacter pylori E. Coli O157:H7 Legionella pneumophilia Salmonella typhi Shigella dysenteriae Vibrio cholerae Gambierdiscus toxicus Gonyaulax catanella Pfiesteria piscicida Giardia lamblia Entamoeba histolytica Cryptosporidium parvum Bacteria (Proteobacteria) Algae Protozoa Disease and symptoms Viscerotropic Neurotrophic Epsilon Epsilon Gamma Gamma Gamma Gamma Gamma Dinoflagellate Dinoflagellate Dinoflagellate Mastigophora Sarcodina Sporozoa Gastroenteritis Infectious hepatitis Poliomyelitis Gastroenteritis, diarrhea, etc Peptic ulcers Diarrhea, hemorrhagic colitis Respiratory illness Typhoid fever, blood in stools Dysentery, blood in stools Cholera Ciguatera fish poisoning Shellfish poisoning Memory loss, dermatitis Giardiasis, diarrhea, bloating Amebiasis, bloody stools Cryptoporidiosis, diarrhea Reading Assignments For the Current Lecture - Environmental Biotechnology; Ch.1, pp. 1-42 - Brock Biology of Microorganisms 12th; Ch.1 & Ch.2 Overview of Biology Systems Ecosystem Communities Populations (at Genus level) Cellular level Subcellular level gene (DNA) => mRNA => protein => enzyme =>function rRNA tRNA Bioremediation 2006 March 17 Park Joonhong (C) Information flow from the gene to the working enzyme catalyst Deoxyribonucleic acid (DNA) (Chromosome, plasmid) Transcription Messenger RNA (ribonucleic acids) A gene Replication Translation by the ribosome, containing ribosomal RNA Amino acid – transfer RNA Bioremediation 2006 March 17 Park Joonhong (C) Protein enzyme Unit component of nucleic acids B ase OH 5 HOCH2 H O C 4 H 3 C 1 H C H C 2 OH OH c.f.) Ribose Ribose unit unit B ase OH 5 HOCH2 H O C 4 H 3 C 1 H C H C 2 OH O HH c.f.) Riboseunit unit Deoxyribose Bioremediation 2006 March 17 Park Joonhong (C) Deoxynucleotide unit O- Ester bond formed with release of H2O Base O- P O O H 5 CH2 O C 4 H 3 Deoxyribose Glycosidic bond formed with release of H2O. C 1 H C H C 2 OH H Monophosphate deoxynucleotide Bioremediation 2006 March 17 Park Joonhong (C) Adenine (A) N O Thymine NH2 Hydrogen bond H3C N H NH N H N H H A-T compliment bond N O H H Cytosine (C) O Guanine (G) NH2 Hydrogen bond H N NH NH H N H H NH2 N H G-C compliment bond Purine Purine bases N O H H Pyrimidine bases Bioremediation 2006 March 17 Park Joonhong (C) OBase O- P Creation of a DNA polynuleotide through a phosphodiester bonds linking the 3 and 5 carbons of the deoxyribose units. O O H 5 CH2 C 4 H 3 O C 1 H C H C 2 O H Base O- P Synthesis of a DNA polynucleotide O O H 5 CH2 O C 4 H 3 Bioremediation 2006 March 17 Park Joonhong (C) C 1 H C H C 2 OH H DNA in a cell is in a double stranded form Chromosome Contain essential genes Vertical transfer of DNA Prokaryotic chromosome is circular, ds DNA Prokaryotic chromosome 2 ~11 x106 base pairs Archaea have 2 Mbps Q: Eukryotic chromosome’s characteristics? (Refer to p.85-86 in the main textbook) B-form of ds DNA c.f.) Z-form Strand 1 5’ 3’ Strand 2 3’ 5’ Plasmid Contain less essential genes But contains environmentally important genes (biodegradation, antibiotic resistance, metal reduction) Horizontal transfer of DNA via conjugation, transformation or virus transduction Prokaryotic chromosome is usually circular, ds DNA Shorter than chromosome but the length widely varies from 0.1 Mbps to couple Mpbs. Bioremediation 2006 March 17 Number of plasmid can be none, one or more… Park Joonhong (C) DNA Replication Critical Steps in DNA Replication Separted in a region (origion); Replication fork A DNA polymerase binds to one strand in the fork, and moves from base to base along both strands in the 3’ to 5’ direction. Generation of a complementary strand of DNA by the polymerase (linking the deoxyribonucleoside triphosphate complementary to the base at which the polymerase is stationed to the previous base on the new, growing chain. (leaning strand, lagged strand) Termination of replication Exonuclease that detects errors, excises the incorrect base, and replaces it with the correct one. Bioremediation 2006 March 17 Park Joonhong (C) Ribonucleic Acid (RNA) B ase OH 5 HOCH2 H O Hydrogen bond H O C 4 H 3 Uracil (U) C NH C 1 H H C 2 H N H H O Base Single stranded form (less stable than dsDNA) Messanger RNA (mRNA) Ribose unitunit c.f.) Ribose Ribosomal RNA (rRNA) Bioremediation 2006 March 17 Transfer RNA (tRNA) Park Joonhong (C) OH OH Transcription: Conversion of DNA into RNA DNA (Chromosome or plasmid) Promotor region rRNA (16S, 32S – forms ribosome, “protein factory”) Protein coding genes mRNA (translated into protein) “Junk” DNA (profound function) tRNA (shuttles for amino acid) Protein coding region in DNA => mRNA coding (open reading frame [ORF]) Non-protein coding region in DNA => rRNA coding, tRNA coding Non-coding region in DNA => “Junk” DNA Bioremediation 2006 March 17 Park Joonhong (C) Transcription: Conversion of DNA into RNA Critical Steps in Transcription A RNA polymerase binds to a promoter region (typically 35 bases ahead of where transcription begins) The dsDNA separates, and the RNA polymerase moves from base to base along one strand in its 3’ to 5’ direction. Termination of transcription: stop at the end of gene; RNA polymerase released from the DNA. Gene Expression and Regulation A RNA polymerase binds to a promoter region and produces mRNA => Gene Expression Up-regulation (Expression): the synthesis of a mRNA is increased Down-regulation (Repression): the synthesis of mRNA is reduced. Inducible versus Constitute Expression Regulation of a gene expression is highly influenced by environmental and physiological factors…(Why genomics is needed.) Bioremediation 2006 March 17 Park Joonhong (C) Translation: Conversion of mRNA into Protein Translation by the ribosome, containing rRNA (large and small subunits) mRNA Protein synthesis Amino acid – tRNA Bioremediation 2006 March 17 Park Joonhong (C) Nucleotide Sequence of a Gene 1 atgagttcag caatcaaaga agtgcaggga gcccctgtga agtgggttac caattggacg 61 ccggaggcga tccgggggtt ggtcgatcag gaaaaagggc tgcttgatcc acgcatctac 121 gccgatcaga gtctttatga gctggagctt gagcgggttt ttggtcgctc ttggctgtta 181 cttgggcacg agagtcatgt gcctgaaacc ggggacttcc tggccactta catgggcgaa 241 gatccggtgg ttatggtgcg acagaaagac aagagcatca aggtgttcct gaaccagtgc 301 cggcaccgcg gcatgcgtat ctgccgctcg gacgccggca acgccaaggc tttcacctgc 361 agctatcacg gctgggccta cgacatcgcc ggcaagctgg tgaacgtgcc gttcgagaag 421 gaagcctttt gcgacaagaa agaaggcgac tgcggctttg acaaggccga atggggcccg 481 ctccaggcac gcgtggcaac ctacaagggc ctggtctttg ccaactggga tgtgcaggcg 541 ccagacctgg agacctacct cggtgacgcc cgcccctata tggacgtcat gctggatcgc 601 acgccggccg ggactgtggc catcggcggc atgcagaagt gggtgattcc gtgcaactgg 661 aagtttgccg ccgagcagtt ctgcagtgac atgtaccacg ccggcaccac gacgcacctg 721 tccggcatcc tggcgggcat tccgccggaa atggacctct cccaggcgca gatacccacc 781 aagggcaatc agttccgggc cgcttggggc gggcacggct cgggctggta tgtcgacgag 841 ccgggctcac tcctggcggt gatgggcccc aaggtcaccc agtactggac cgagggtccg 901 gctgccgagc ttgcggaaca gcgcctgggg cacaccggca tgccggttcg acgcatggtc 961 ggccagcaca tgacgatctt cccgacctgt tcattcctgc ccaccttcaa caacatccgg 1021 atctggcacc cgcgtggtcc caatgaaatc gaggtgtggg ccttcaccct ggtcgatgcc 1081 gacgccccgg cggagatcaa ggaagaatat cgccggcaca acatccgcaa cttctccgca 1141 ggcggcgtgt ttgagcagga cgatggcgag aactgggtgg agatccagaa ggggctacgt 1201 gggtacaagg ccaagagcca gccgctcaat gcccagatgg gcctgggtcg gtcgcagacc 1261 ggtcaccctg attttcctgg caacgtcggc tacgtctacg ccgaagaagc ggcgcggggt 1321 atgtatcacc actggatgcg catgatgtcc gagcccagct gggccacgct caagccctga • bphA gene in Burkholderia xenovorans LB400 [gene index number:349602] Bioremediation 2006 March 17 Park Joonhong (C) Symbols for Amino Acids A Ala alanine R Arg Arginie B Asx aspargine S Ser Serine C Cys Cysteine T Thr Threonine D Asp Aspartic acid V Val Valine E Glu Glutamic acid W Trp Tryptophan F Phe Phenylalanine Y Tyr Tyrosine G Gly Glycine Z Glx Glutamine H His Histidine I Ile Isoleucine K Lys Lysine L Leu Leucine M Met Methionine N Asn Asparagine P Pro Proline Q Gln GlutamineBioremediation 2006 March 17 Park Joonhong (C) Standard Genetic Code UUU UUC UUA UUG Phe(F) Phe(F) Leu(L) Leu (L) UCU UCC UCA UCG Ser(S) Ser(S) Ser(S) Ser(S) UAU UAC UAA UAG Tyr (Y) Tyr (Y) Stop Stop UGU UGC UGA UGG Cys(C) Cys(C) Stop Trp(W) CUU CUC CUA CUG Leu (L) Leu (L) Leu (L) Leu (L) CCU CCC CCA CCG Pro(P) Pro(P) Pro (P) Pro (P) CAU CAC CAA CAG His (H) His (H) Gln(Q) Gln(Q) CGU CGC CGA CGG Arg (R) Arg (R) Arg (R) Arg (R) AUU AUC AUA AUG Ile (I) Ile (I) Ile (I) Met(M) ACU ACC ACA ACG Thr (T) Thr (T) Thr (T) Thr (T) AAU AAC AAA AAG Asn(N) Asn(N) Lys(K) Lys(K) AGU AGC AGA AGG Ser(S) Ser(S) Arg(R) Arg(R) GUU GUC GUA GUG Val(V) Val (V) Val (V) Val (V) GCU GCC GCA GCG Ala(A) GAU Asp(D) Ala(A) GAC Asp(D) Ala(A) GAA Glu(E) Bioremediation 2006 March 17 Ala(A) GAG Glu(E) Park Joonhong (C) GGU GGC GGA GGG Gly(G) Gly(G) Gly(G) Gly(G) Amino Acid Sequence of a Protein Methods of obtaining amino acid sequences. - Experimentally determined - Bioinformatically translated using Standard Genetic Code 1 mssaikevqg apvkwvtnwt peairglvdq ekglldpriy adqslyelel ervfgrswll 61 lgheshvpet gdflatymge dpvvmvrqkd ksikvflnqc rhrgmricrs dagnakaftc 121 syhgwaydia gklvnvpfek eafcdkkegd cgfdkaewgp lqarvatykg lvfanwdvqa 181 pdletylgda rpymdvmldr tpagtvaigg mqkwvipcnw kfaaeqfcsd myhagttthl 241 sgilagippe mdlsqaqipt kgnqfraawg ghgsgwyvde pgsllavmgp kvtqywtegp 301 aaelaeqrlg htgmpvrrmv gqhmtifptc sflptfnnir iwhprgpnei evwaftlvda 361 dapaeikeey rrhnirnfsa ggvfeqddge nwveiqkglr gykaksqpln aqmglgrsqt 421 ghpdfpgnvg yvyaeeaarg myhhwmrmms epswatlkp • BphA protein in Burkholderia xenovorans LB400 [gi:584852] Bioremediation 2006 March 17 Park Joonhong (C) Reading Assignments For the Current Lecture -Brock Biology of Microorganisms 12th; Ch.7