* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Document
Pathogenomics wikipedia , lookup
Neocentromere wikipedia , lookup
Copy-number variation wikipedia , lookup
Population genetics wikipedia , lookup
Skewed X-inactivation wikipedia , lookup
Dominance (genetics) wikipedia , lookup
Holliday junction wikipedia , lookup
Genetic engineering wikipedia , lookup
Y chromosome wikipedia , lookup
Essential gene wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Gene desert wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
History of genetic engineering wikipedia , lookup
Homologous recombination wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
X-inactivation wikipedia , lookup
Designer baby wikipedia , lookup
Minimal genome wikipedia , lookup
Genome evolution wikipedia , lookup
Genomic imprinting wikipedia , lookup
Ridge (biology) wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Gene expression programming wikipedia , lookup
Gene expression profiling wikipedia , lookup
Genome (book) wikipedia , lookup
Microevolution wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
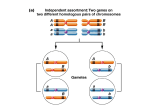
Linkage • Genes linked on the same chromosome may segregate together. Independent Assortment 2 chromosomes A a B P: AABB x aabb b Parental Gametes: AB & ab F1 A B P A b R a B R a b P Meiosis 2n = 2 One Chromosome A a No Cross Over B b Parent Cell A A a a B B b b Daughter Cells Have Parental Chromosomes Meiosis 2n = 2 One Chromosome A a With Cross Over B b Parent Cell A A a a B b B b Daughter Cells Have Recombinant Chromosomes Meiosis Prophase I A a A a B B b b If genes are linked, crossing over must occur for there to be recombination. no linkage Linkage? linkage P: AABB x aabb P: AABB x aabb F1: AaBb F1: AaBb Test cross: AaBb x aabb Test cross: AaBb x aabb 1/4: AaBb ?: AaBb 1/4: Aabb ?: Aabb 1/4: aaBb ?: aaBb 1/4: aabb ?: aabb Recombination Frequency …or Linkage Ratio: the percentage of recombinant types, – if 50%, then the genes are not linked, – if less than 50%, then linkage is observed. Linkage • Genes located on the same chromosome do not recombine, – unless crossing over occurs, • The recombination frequency gives an estimate of the distance between the genes. Recombination Frequencies • Genes that are adjacent have a recombination frequency near 0%, • Genes that are very far apart on a chromosome have a linkage ratio of 50%, • The relative distance between linked genes influences the amount of recombination observed. A B a b In this example, there is a 2/10 chance of recombination. A C a c In this example, there is a 4/10 chance of recombination. Linkage Ratio P GGWW x ggww Testcross F1: GgWw x ggww GW Gw gW gw ? ? ? ? recombinant total progeny = Linkage Ratio Linkage Ratio Units % = mu (map units) - or % = cm (centimorgan) A B a b In this example, there is a 2/10 chance of recombination. A C a c In this example, there is a 4/10 chance of recombination. cis “coupling” trans “repulsion” Study Figs 4.2, 4.3, and 4.5 Fly Crosses (white eyes, minature, yellow body) • In a white eyes x miniature cross, 900 of the 2,441 progeny were recombinant, yielding a map distance of 36.9 mu, • In a separate white eyes x yellow body cross, 11 of 2,205 progeny were recombinant, yielding a map distance of 0.5 mu, • When a miniature x yellow body cross was performed, 650 of 1706 flies were recombinant, yielding a map distance of 38 mu. Simple Mapping • white eyes x miniature = 36.9 mu, • white eyes x yellow body = 0.5 mu, • miniature x yellow body = 38 mu, 0.5 mu 36.9 mu y w m 38 mu Do We have to Learn More Mapping Techniques? • Yes, – three point mapping, • Why, – Certainty of Gene Order, – Double crossovers. Gene Order • It is often difficult to assign the order of genes based on two-point crosses due to uncertainty derived from sampling error. A x B = 37.8 mu, A x C = 0.5 mu, B x C = 37.6 mu, Double Crossovers • More than one crossover event can occur in a single tetrad between non-sister chromatids, – if recombination occurs between genes A and B 30% of the time, • (p = 0.3), • then the probability of the event occurring twice is 0.3 x 0.3 = 0.09, or nearly 10 map units. • If there is a double cross over, does recombination occur? – how does it affect our estimation of distance between genes? Why Me? Why Map? • Over 4000 human diseases have a genetic component, – knowing the protein produced at specific loci facilitates the treatment and testing, • Facilitates both classical and molecular analysis of organisms. Classical Mapping target Cross an organism with a trait of interest to homozygous mutants of known mapped genes. Then, determine if segregation is random in the F2 generation, What recombination frequency do you expect beteen the target and HY2? What recombination frequency do you expect beteen the target and TT2? • if not, then your gene is linked (close) to the known mapped gene. Three Point Testcross Triple Heterozygous (AaBbCc ) x Triple Homozygous Recessive (aabbcc) Three Point Mapping Requirements • The genotype of the organism producing the gametes must be heterozygous at all three loci, • You have to be able to deduce the genotype of the gamete by looking at the phenotype of the offspring, • You must look at enough offspring so that all crossover classes are represented. w g d Representing linked genes... P W w G g D d = WwGgDd d d = wwggdd x Testcross w w g g w g d Representing linked genes... P + w + g + d = WwGgDd d d = wwggdd x Testcross w w g g Phenotypic Classes GWgg Ddd Ddd D- G- dd W-G-DW-G-dd W-gg-D W-gg-dd wwG-DwwG-dd ww gg Ddd wwggDwwggdd # Crossovers W-G-D- 179 0 wwggdd 173 0 W-G-dd 46 1 wwggD- 52 1 wwG-D- 22 1 W-gg-dd 22 1 W-gg-D 2 2 wwG-dd 4 2 W G D w g d # W-G-D- 179 wwggdd 173 W-G-dd 46 wwggD- 52 wwG-D- 22 W-gg-dd 22 W-gg-D 2 wwG-dd 4 Parentals Recombinants 1 crossover, Region I Recombinants 1 crossover, Region II Recombinants, double crossover II I W G D w g d # W-G-D- 179 wwggdd 173 W-G-dd 46 wwggD- 52 wwG-D- 22 W-gg-dd 22 W-gg-D 2 wwG-dd 4 Total = 500 I Parentals Recombinants 1 crossover, Region I Recombinants 1 crossover, Region II Recombinants, double crossover W G D w g d Region I: 46 + 52 + 2 + 4 500 = 20.8 mu x 100 # W-G-D- 179 wwggdd 173 W-G-dd 46 wwggD- 52 wwG-D- 22 W-gg-dd 22 W-gg-D 2 wwG-dd 4 Total = 500 II Parentals Recombinants 1 crossover, Region I Recombinants 1 crossover, Region II Recombinants, double crossover 20.8 mu W G D w g d Region II: 22 + 22 + 2 + 4 500 = 10.0 mu x 100 10.0 mu W w 20.8 mu G D g d 0.1 x 0.208 = 0.0208 NO GOOD! W-gg-D wwG-dd Total = 2 4 500 Recombinants, double crossover 6/500 = 0.012 Interference …the affect a crossing over event has on a second crossing over event in an adjacent region of the chromatid, – (positive) interference: decreases the probability of a second crossing over, • most common in eukaryotes, – negative interference: increases the probability of a second crossing over. Gene Order in Three Point Crosses • Find either double cross-over phenotype, based on the recombination frequencies, • Two parental alleles, and one cross over allele will be present, • The cross over allele fits in the middle... – # A-B-C- 2001 aabbcc 1786 A-B-cc 46 aabbC- 52 aaB-cc 990 A-bb-C- 887 A-bb cc 600 aaB-C- 589 Which one is the odd one? II I A C B a c b # A-B-C- 2001 aabbcc 1786 Region I 990 + 887 + 46 + 52 A-B-cc 46 6951 aabbC- 52 = 28.4 mu aaB-cc 990 A-bb-C- 887 A-bb cc 600 aaB-C- 589 x 100 I A C B a c b # A-B-C- 2001 aabbcc 1786 Region II 600 + 589 + 46 + 52 A-B-cc 46 6951 aabbC- 52 = 18.5 mu aaB-cc 990 A-bb-C- 887 A-bb cc 600 aaB-C- 589 18.5 II mu x 100 28.4 mu A C B a c b Today • Coefficient of Confidence, • Gene mapping in humans, • Problems, problems, problems, – Be sure to at least try them before Friday.