* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Phenotype
Vectors in gene therapy wikipedia , lookup
Whole genome sequencing wikipedia , lookup
Heritability of IQ wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Gene expression programming wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Human genome wikipedia , lookup
Pharmacogenomics wikipedia , lookup
Dominance (genetics) wikipedia , lookup
Population genetics wikipedia , lookup
Pathogenomics wikipedia , lookup
Metagenomics wikipedia , lookup
Genetic engineering wikipedia , lookup
Human genetic variation wikipedia , lookup
Non-coding DNA wikipedia , lookup
Genome (book) wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Primary transcript wikipedia , lookup
Human Genome Project wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
History of genetic engineering wikipedia , lookup
Designer baby wikipedia , lookup
Helitron (biology) wikipedia , lookup
Genome evolution wikipedia , lookup
Genome editing wikipedia , lookup
Public health genomics wikipedia , lookup
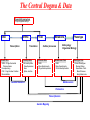
The Central Dogma & Data
Protein-DNA binding Data
Chip-chip protein arrays
DNA
Protei
n
mRNA
Translation
Transcription
Genetic Data
SNPs – Single Nucleotide
Polymorphisms
Re-sequencing
CNV - Copy Number Variation
Microsatellites
Transcript Data
Micro-array data
Gene Expression
Exon
Splice Junction
Metabolite
Cellular processes
Proteomic Data
NMR
Mass Spectrometry
2D-gel electrophoresis
Embryology
Organismal Biology
Metabonomic Data
NMR
Mass Spectrometry
2D-Gel electrophoresis
Metabonomics
Genetical Genomics
Proteomics
Transcriptomics
Genetic Mapping
Phenotype
Phenotypic Data
Clinical Phenotypes
Disease Status
Quantitative Traits
Blood Pressure
Body Mass Index
Structure of Integrative Genomics
DNA
Classes
mRNA
Protei
n
Metabolite
Phenotype
Parts
Concepts
GF Mapping
Models: Networks
Physical models:
Phenomenological models:
Unobservered/able
Hidden Structures/ Processes
Knowledge:
Evolution:
Externally Derived Constraints on which Models are acceptable
Cells in Ontogeny
Individuals/Sequences in a Population
Analysis: Data + Models + Inference
Functional Explanation
Model Selection
Species
G: Genomes
A diploid genome:
Key challenge: Making a single molecule observable!!
Classical Solution (70s): Many
De Novo Sequencing: Halted extensions or degradation
extension
degradation
80s: From one to many: PCR – Polymerase Chain Reaction
00s: Re-sequencing: Hybridisation to complete genomes
Future Solution: One is enough!!
Observing the behavior of the polymerase
Passing DNA through millipores registering changes in current
G: Assembly and Hybridisation
Target genome
3*109 bp
(unobservable)
Reads
3-400 bp
(observable)
Contigs
Contigs and Contig Sizes as function of Genome Size (G), Read Size (L) and overlap (Ø):
{A,C}
Complementary or almost
complementary strings allow
interrogation.
probe
{T,G}
Lander & Waterman, 1988 Statistical Analysis of Random Clone
Fingerprinting
Sufficient overlap allows concatenation
T - Transcriptomics
Measures transcript levels
averaging of a set of cells.
P – Proteomics, M – Metabonomics & F - Phenomics
P uses Mass Spectrometry and 2D gel electrophoresis of degraded peptides and Protein
Arrays using immuno-recognition of complete proteins
M uses Mass Spectrometry and 2D gel electrophoresis of metabolites
F: The set of all phenotypes.
Hard to define
Focus on Clinical Traits
Behavioural Traits hard to observe
Concepts
GF Mapping
Physical models:
Models: Networks
Phenomenological models:
Hidden Structures/ Processes
Knowledge:
Evolution:
Unobservered/able
Externally Derived Constraints on which Models are acceptable
Cells in Ontogeny
Individuals/Sequences in a Population
Species
G F
• Mechanistically predicting relationships between different data types is very difficult
• Empirical mappings are important
• Functions from Genome to Phenotype stands out in importance
G is the most abundant data form - heritable and precise. F is of greatest interest.
DNA
mRNA
Protei
n
Metabolite
Phenotype
“Zero”-knowledge mapping: dominance,
recessive, interactions, penetrance, QTL,.
Mapping with knowledge: weighting
interactions according to co-occurence in
pathways.
Model based mapping:
genomesystemphenotype
Height
Weight
Disease
status
Intelligence
……….
Environment
The General Problem is Enormous
Set of Genotypes:
1
3* 107
• Diploid Genome
• In 1 individual, 3* 107 positions could segregate.
• In the complete human population 5*108 might segregate.
• Thus there could be 2500.000.00 possible genotypes
Partial Solution: Only consider functions dependent on few positions
• Causative for the trait
Classical Definitions:
• Single Locus
• Multiple Loci
Dominance
Recessive
Additive
Heterotic
Epistasis: The effect of one locus depends on the state of another
Quantitative Trait Loci (QTL). For instance sum of functions for positions plus error
term.
X (G )
i
i
i causative positions
Genotype and Phenotype Covariation: Gene Mapping
Sampling Genotypes and Phenotypes
Decay of local dependency
Time
Reich et al. (2001)
Genetype -->Phenotype Function
Dominant/Recessive
Penetrance
A set of characters.
Binary decision (0,1).
Spurious Occurrence
Quantitative Character.
Heterogeneity
genotype
Genotype Phenotype
phenotype
Result:The Mapping Function
Pedigree Analysis & Association Mapping
Association Mapping:
Pedigree Analysis:
D
M
r
M
Pedigree known
Few meiosis (max 100s)
2N generations
D
r
Resolution: cMorgans (Mbases)
Pedigree unknown
Many meiosis (>104)
Resolution: 10-5 Morgans (Kbases)
Adapted from McVean and others
Heritability: Inheritance in bags, not strings.
The Phenotype is the sum of a series of factors, simplest independently genetic and
environmental factors: F= G + E
Relatives share a calculatable fraction of factors, the rest is drawn from the background
population.
This allows calculation of relative effect of genetics and environment
Heritability is defined as the relative
contribution to the variance of the genetic
factors: G2 / F2
Parents:
Has been been defined for 2 characters
simultaneously to define common factors
Siblings:
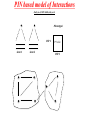
PIN based model of Interactions
Emily et al, 2009 & Rhzetsky et al.
Phenotype i
SNP 1
Gene 1
Gene 2
3*3 table
SNP 2
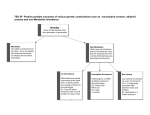
Summary of this lecture
Data
G - genetic variation
Concepts
GF Mapping
T - transcript levels
Models: Networks
P - protein concentrations
Hidden Structures/ Processes
M - metabolite concentrations
Knowledge
F – phenotype/phenome
Evolution
GF Mapping
General Function Enormous
Used for Disease Gene Finding
Can Include Biological Knowlede