* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download AUG
Molecular evolution wikipedia , lookup
Transcriptional regulation wikipedia , lookup
Citric acid cycle wikipedia , lookup
Eukaryotic transcription wikipedia , lookup
Self-assembling peptide wikipedia , lookup
Protein (nutrient) wikipedia , lookup
RNA polymerase II holoenzyme wikipedia , lookup
Point mutation wikipedia , lookup
Cell-penetrating peptide wikipedia , lookup
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Polyadenylation wikipedia , lookup
Protein structure prediction wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Gene expression wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Proteolysis wikipedia , lookup
Peptide synthesis wikipedia , lookup
Non-coding RNA wikipedia , lookup
Biochemistry wikipedia , lookup
Messenger RNA wikipedia , lookup
Bottromycin wikipedia , lookup
Epitranscriptome wikipedia , lookup
Genetic code wikipedia , lookup
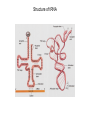
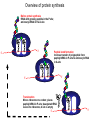
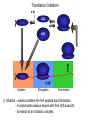
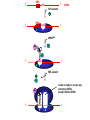
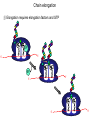
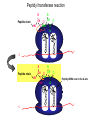
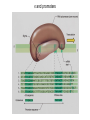
The genetic code Nucleic acids Nucleic acids Correspondence = the genetic code Amino acids Codon = triplet of three bases which encodes an amino acid 64 possible codons = 43 each of 4 nucleotides can occupy each of 3 positions in the codon Deciphering the code 61 codons encode amino acids, 3 codons do not specify amino acids Specialized codons: - for start of translation - AUG - for STOP - UAA, UAG, UGA 61 codons encode 20 amino acids - most amino acids are specified by more than one codon - degeneracy of the genetic code Transfer RNA (tRNA) is the adapter tRNA has two crucial properties: - it caarries a single amino acid to which it is covalently linked - it contains the anticodon (complementary to the codon representing its amino acid) Codon-anticodon interactions often one tRNA can recognize more than one codon tRNALys can recognize AAA or AAG 3’ 5’ CGG anticodon mRNA 5’ GCU 3’ wobble hypothesis: the pairing between codon and anticodon at the first two codon positions always follows the usual rules, but exceptional “wobbles” occur at the third position Base in first position Base recognized in third of anticodon position of codon U A/G C G A U G C/U Structure of tRNA Aminoacyl-tRNA synthetases all synthetases function by two-step mechanism: 1) activation of amino acid with ATP 2) transfer of activated amino acid to tRNA Enzyme Amino acid site ATP site R H-C-NH2 C O tRNA site O- O-O-P=O O -O-P=O O -O-P=O O Adenosine R R H-C-NH2 C O O-H OO- P=O O Adenosine H-C-NH2 C O O tRNA synthetases are responsible for the fidelity of translation Ribosome - site of protein synthesis • ribosome provides the environment for controlling the interaction between mRNA and aminoacyl-tRNA Ribosomes Bacteria Subunits rRNA Proteins 50S 23S, 5S 31 30S 16S 21 60S 28S, 5.8S, 5S 49 40S 18S 33 70S Mammals 80S The ribosome has two sites for binding charged tRNA mRNA is associated with small (30S) subunit tRNA spans both subunits amino acid end in the large subunit anticodon in the small subunit 5’ 3’ P-site = peptide site A-site = acceptor site growing peptide held by tRNA entered by aminoacyl-tRNA Ribosome movement 5’ 3’ Overview of protein synthesis Before protein synthesis tRNA with growing peptide in the P site; aminoacyl-tRNA in the A-site 3’ 5’ Peptide bond formation Involves transfer of polypeptide from peptidyl-tRNA in P-site to aminoacyl-tRNA in A-site 5’ Translocation Moves ribosome one codon; places peptidyl-tRNA in P-site; deacylated tRNA leaves the ribosome; A site is empty 5’ 3’ 3’ Translation Initiation 30S 50S Initiation Elongation Termination Initiation - reactions before the first peptide bond formation - in prokaryotes always begins with free 30S subunits - formation of an initiation complex Translation Initiation Initiation occurs at a special sequence on mRNA - ribosome binding site (RBS) or Shine-Dalgarno sequence - complementary to the 3’end of 16S rRNA 3’ end of 16S rRNA 3’ A 5’ U mRNA UCCUCCA 5’ NNNNNAGGAGGU-N5-10-AUG---- 3’ ShineDalgarno sequence Initiation codon Initiation codon - signal for initiation of translation - usually the triplet AUG (in bacteria also GUG or UUG) - AUG represents methionine Translation Initiation A special initiator tRNA starts the polypeptide chain - N-formyl-methionine tRNA - unique to bacteria - used only for initiation O NH2 O H-C-----C-O CH2 CH2 S CH3 H-C-O NH2 O H-C-----C-O CH2 CH2 S CH3 methionine N - formyl - methionine Initiation requires initiation factors - found only on 30S subunit; released when 50S joins - three factors needed for mRNA and tRNA to enter the complex 5’ RBS 3’ AUG mRNA 30S subunit IF3 5’ RBS 3’ AUG IF3 P tRNAfMet IF2 fMet IF2 5’ 3’ AUG IF3 P 50S subunit IF3 IF2 A-site is ready to accept any aminoacyl-tRNA except initiator tRNA fMet 5’ 3’ AUG P A Chain elongation Elongation requires elongation factors and GTP EF 3’ 5’ EF 3’ 5’ 5’ 3’ Peptidyl transferase reaction Peptide chain R R CH O CH O C HN C 2HN O O 3’ 5’ Peptide chain 5’ R R CH O CH O C HN C N O HO Peptidyl-tRNA now in the A-site 3’ Translocation moves the ribosome ribosome advances three nucleotides along the mRNA 5’ 3’ result - expel the uncharged tRNA from the P-site - new peptidyl-tRNA enters P-site - A-site is free for the next aminoacyl-tRNA or termination Translation termination 3 triplets not represented by a tRNA: UAG, UAA, UGA STOP codons are recognized by release factors (RF1, RF2) Release factor 5’ STOP Dissociation 5’ 3’ 3’ Antibacterial antibiotics Antibiotic Site of action Streptomycin Chloramphenicol Tetracycline Kanamycin inhibits translation initiation; binds 30S subunit inhibits elongation during translation; binds 50S inhibits translation; prevents aminoacyl tRNA binding inhibits translation; binds 30S and prevents translocation Rifamycin inhibits RNA synthesis; binds to b’ subunit of RNA polymerase Novobiocin inhibits DNA gyrase Ampicillin/Penicillin inhibits cell wall synthesis Bacteria Eukaryotic cells - mRNA transcribed and translated in the same compartment - synthesis and maturation of mRNA occur in the nucleus - transcription and translation occur simultaneously - translation occurs in the cytoplasm - mRNA is usually unstable - translated - mRNA is stable - translated for for short period of time (minutes) several hours - mRNA is usually polycistronic RBS AUG STOP RBS Intercistronic spacer - mRNA is mostly monocistronic STOP cap AAAAAAA cap AAAAAAA AUG s and promoters E. coli sigma factors recognize promoters with different consensus sequences Factor Gene Use -35 Sequence Separation -10 Sequence s70 rpoD general TGACA 16-18 bp TATAAT s32 rpoH heat shock CNCTTGAA 13-15 bp s54 rpoN nitrogen CTGGNA 6 bp CCCCATNT TTGCA