* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download From DNA to Protein
Proteolysis wikipedia , lookup
RNA interference wikipedia , lookup
RNA silencing wikipedia , lookup
Biochemistry wikipedia , lookup
Transformation (genetics) wikipedia , lookup
Polyadenylation wikipedia , lookup
Real-time polymerase chain reaction wikipedia , lookup
Community fingerprinting wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Eukaryotic transcription wikipedia , lookup
RNA polymerase II holoenzyme wikipedia , lookup
Gene regulatory network wikipedia , lookup
Non-coding DNA wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Messenger RNA wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Genetic code wikipedia , lookup
Transcriptional regulation wikipedia , lookup
Point mutation wikipedia , lookup
Gene expression wikipedia , lookup
Biosynthesis wikipedia , lookup
Silencer (genetics) wikipedia , lookup
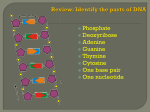
From DNA to Protein ... Chapters 10 & 11 Overview • Review of DNA & RNA • Transcription & Translation • Gene Mutations • Controls over Genes DNA: A Review Holds: Genetic information Protein-building instructions Double-helix of nucleotide bases with sugar-phosphate backbone Bases held together by H-bonds: – A always pairs with T – G always pairs with C So what is a gene? Segment of DNA molecule Carries instructions for 1 polypeptide chain Bases grouped in triplets that code for specific amino acid Variations in arrangement of bases lets cells make all proteins needed Exons = protein-coding base sequences Introns = non-coding, repetitive sequences (genome scrapyard of ready-to-use DNA segments & small RNA molecules) Both transcribed but introns removed before mRNA reaches cytoplasm RNA: A Review Similar to DNA, except: – Single-stranded – Uracil replaces thymine • Adenine pairs with uracil Decodes DNA & acts as messenger Types of RNA: mRNA Messenger RNA Carries protein-building instructions from gene to ribosome “Half-DNA” Types of RNA: rRNA Ribosomal RNA One of components of ribosomes With tRNA, translate protein-building instructions carried by mRNA Ribosomes 2 subunits of rRNA & structural proteins Have 2 tRNA binding sites Come together as whole functional ribosome during translation Ribosomes of prokaryotes and eukaryotes are similar in function but different in composition Certain antibiotics (e.g. tetracycline, streptomycin) inactivate prokaryotic ribosomes but don’t affect eukaryotic ribosomes Types of RNA: tRNA Transfer RNA 45 different types With rRNA, translate proteinbuilding instructions carried by mRNA Has anti-codon head: = 3-base sequence complementary to codon on mRNA transcript Anti-codon head is complementary to amino acid it carries 45 tRNAs exist in eukaryotic cells Codon-anticodon pairing has “wiggle room” for 3rd base of codon e.g. AUU, AUC, AUA (isoleucine) use same tRNA The Genetic Code The rules that link codons in RNA with the corresponding amino acids in proteins Bases read 3 at a time = codon 64 codons that code for 20 amino acids Some amino acids have ≥ 1 codon (↓ transcription & translation errors) AUG = methionine = START UAA, UAG, UGA = STOP Transcription & Translation Process that turns sequence of nucleotide bases in genes into sequence of amino acids in proteins transcription DNA translation RNA protein DNA base sequence acts as template to make RNA Occurs in eukaryotic nucleus RNA moves into cytoplasm Amino acids join to become polypeptides (proteins) Transcription DNA gene’s base sequence → complementary mRNA base sequence First step in protein synthesis Sequence of nucleotides bases on DNA strand exposed Becomes template for RNA to be built from A, C, G, T Transcription factor binds to promoter (START) base sequence on DNA Promoter determines where mRNA synthesis begins & which DNA strand is template RNA polymerase binds to promoter (unwinds 16-18 bps of DNA helix) RNA polymerase moves along protein-coding gene region RNA polymerase unwinds DNA in front & rewinds behind as mRNA elongates Incoming RNA nucleotides bind with complementary bases on template strand e.g. (AGC) on DNA → (UCG) on mRNA Creates complementary sequence from DNA base sequence template mRNA is released at end of gene region (STOP) Is actually pre-mRNA because has intron junk mRNA modified before leaving nucleus = introns cut out & exons respliced to form functional mRNA mRNA associates with proteins & leaves nucleus = is now ready for protein synthesis mRNA enters cytoplasm = location of pool of tRNA & free amino acids Protein synthesis (translation) begins Translation mRNA base sequence → amino acids → proteins mRNA transcript enters ribosome Codons translated into polypeptide chain Initiation of Translation mRNA binds to small ribosomal unit Initiator tRNA binds to start codon (AUG) (this tRNA carries Met & has anticodon UAC) Large ribosomal subunit binds to small subunit to form functional ribosome Initiator tRNA fits into P site of ribosome (P site holds growing polypeptide) A site lies vacant for the next amino-acidcarrying tRNA Elongation of Translation Chain of polypeptides is synthesized as mRNA passes between ribosomal subunits tRNAs transfers amino acids from cytosol to ribosome Elongation is a 3-step process 1. Codon recognition: Anti-codon of incoming amino-acidcarrying tRNA pairs with mRNA codon in A site Amino acids bind to mRNA in order dictated by template of codons 2. Peptide bond formation: Polypeptide separates from tRNA in P site & attaches to amino acid carried by tRNA in A site Peptide bond catalyzed by rRNA in large ribosomal subunit 3. Translocation: P site tRNA leaves ribosome Ribosome moves tRNA in A site (with attached polypeptide) to P site (mRNA moves along too) Next mRNA codon is brought into A site Elongation begins over again for next addition Polyribosomes Once mRNA passes through ribosome, may become attached to multiple other ribosomes in row Allows many copies of same protein to be made quickly & simultaneously Termination of Translation mRNA STOP codon enters ribosome (no tRNA has complementary anticodon) Release factors bind to ribosome & detach mRNA & polypeptide chain Ribosome separates back into 2 subunits Proteins either: – Join pool of free proteins in cytoplasm – Enter RER to be modified for transport Summary of Transcription & Translation Genetic info → protein synthesis Via info transfer of complementary base pairing Phe Gly Arg Phe Gene Mutations Most mutations are spontaneous & occur during DNA replication DNA polymerases & ligases (proofreaders) catch most errors but not all Bases can be substituted, inserted, deleted Effects on protein structure & function depend on how mRNA sequence is changed Point Mutations a.k.a. base substitution Single nucleotide replaced with different nucleotide Can be harmless if still codes for same amino acid Can be harmful or even fatal (wrong amino acid can alter protein function or even code for STOP) a. Missense mutation Substitution alters codon so that it codes for different amino acid Usually changes protein function (good / bad / neutral effects) GCA-UUC-GUC ala - phe - val GCA-UUA-GUC ala - leu - val b. Nonsense mutation Substitution alters codon so that it codes for STOP signal Results in premature termination of translation Shortened protein is usually non-functional GCA-UAU-GUC ala - tyr - val GCA-UAG-GUC ala - STOP c. Silent mutation Substitution occurs in 3rd base of mRNA codon New codon codes for same amino acid (does not affect protein function) GCA-UUC-GUC ala - phe - val GCA-UUU-GUC ala - phe - val Frameshift Mutations 1 or more base inserted or deleted Deletion or insertion shifts 3-base reading window Protein is generally useless = extensive missense & eventually nonsense Mutagens Some mutations are not spontaneous Ionizing radiation (e.g. x-rays) = break up chromosomes & deposit free radicals in cells Non-ionizing radiation (e.g. UV radiation) = changes base-pairing properties due to thymine sensitivity When are mutations good? If occur in somatic (body) cells, only affects individual (not heritable) If occur in gametes (sex cells), may be heritable – Can result in harmful, beneficial, or neutral effects on individual’s survival – Adaptation or elimination? Cell Differentiation Body cells differ in composition, structure, & function Each cell type undergoes selective gene expression = determines which tissues & organs develop How Are Genes Regulated? Differentiated cells contain all genes BUT Cells only express genes necessary for their specialized functions Human genome = 25,000 – 30,000 genes Most transcribed only in certain cells at certain times (default state = off) Some transcribed in all cells because encode proteins / RNA that are essential for life = housekeeping genes Animal development is directed by cascades of gene expression & cell-to-cell signalling Homeotic gene = master control gene that regulates all other genes Gene Control How fast & when genes will be transcribed & translated Whether gene products are switched on or silenced = Controls over what kinds & how much of each protein are in a cell Regulatory elements respond to concentration changes & chemical signals in environment e.g. DNAs, RNAs, polypeptide chains, proteins Both negative & positive controls exist Promoters & Enhancers Promoters: – Short base sequences in DNA – Regulatory proteins control transcription of specific genes Enhancers: – Binding sites where promoters increase transcription rates Controls Before Transcription Access to genes – Blocked vs. open How genes are transcribed – Sequences can be rearranged or multiplied • Allows rapid & simultaneous production of gene products Control of Transcript Processing Frequency of transcription How genes are transcribed – Sequences can be rearranged or multiplied • Allows rapid & simultaneous production of gene products Control of Translation Rate of translation How many times translation can occur on a particular mRNA Controls After Translation Proteins & protein synthesis molecules can be: Activated Inhibited Stabilized Modified Degraded Animal Gene Controls: X Chromosome Inactivation 1 of 2 copies of X chromosome in female mammals is inactivated Condenses so can’t be transcribed = Barr body So that female (XX) doesn’t have twice as many X chromosome gene products as male (XY) = Dosage compensation Which X chromosome is inactivated is random in any given cell – Some cells & descendants will express genes from maternal X chromosome – Other cells & descendants will express genes from paternal X chromosome Plant Gene Controls: ABC Model 3 sets of genes determine how specialized parts of flower develop in predictable pattern In cells at tip of forming flower, different sets of genes activated to form sepals, petals, sexual structures Up to now, we have been largely focused on eukaryotic cells. What about prokaryotic cells? Prokaryotic Gene Control Primarily by changes in transcription rate (depends on environmental conditions e.g. nutrient availability, etc.) When growth & reproduction conditions are optimum, cells rapidly transcribe growth enzymes & nutrient-absorbing genes e.g. E. coli & the lactose operon Gut of human mammals Set of 3 genes produces lactosemetabolizing enzymes In front of genes is promoter & operator = operon (controls expression of > 1 gene at a time) Negative Control of the Lactose Operon in E. coli Without lactose: – Repressor binds to operators – Twists DNA region so that RNA polymerase can’t bind = no transcription occurs With lactose: – E. coli converts lactose to allolactose – Binds to repressor & changes its shape so can’t bind to operators – Twisted DNA unwinds, RNA polymerase binds, & protein synthesis of lactose-metabolizing enzymes begins Bacteria divide via binary fission = genetically-identical offspring Can increase genetic variation by transferring DNA between different bacterial cells = 3 mechanisms a. Transformation Take up DNA from surroundings e.g. from dead cells in the environment b. Transduction Transfer genes via phage (DNA stowaway) Phage = virus that infects bacteria c. Conjugation Mating & DNA transfer between 2 bacterial cells Conjugation is enabled by the F factor F factor can exist as a plasmid = small, circular DNA R plasmids carry genes that destroy antibiotics = confers antibiotic resistance Widespread use of antibiotics has resulted in antibiotic-resistant strains of “superbugs” Regardless of how DNA is transferred: When new DNA enters bacterial cell, parts integrate into existing chromosome Part of donated DNA replaces part of original DNA = recombinant chromosome Viruses “Genes in a box” Nucleic acid contained within a capsid Not living = can only reproduce within host cells Some viruses contain RNA = flu, cold, measles, mumps, AIDS, polio Some viruses contain DNA = hepatitis, chicken pox, herpes Vaccines may prevent these viruses, but very few effective anti-viral drugs (kill both host & viral cells) Amount of damage caused by virus depends on: • Immune response • Self-repair capabilities of affected tissue e.g. recover from colds quickly because of rapid regeneration of respiratory tract tissues e.g. poliovirus causes permanent damage because affects non-dividing nerve cells Viruses arise from: Mutations e.g. new strains of flu viruses Contact between species e.g. HIV transmitted from chimps to humans Spread from isolated populations e.g. HIV spread from small region of Africa to worldwide distribution Some viruses carry cancer-causing genes = oncogenes Proto-oncogene = normal gene that has potential to mutate into oncogene Tumor-suppressor genes = inhibit cell division (if mutate, cell may end up dividing multiple times & forming cancerous tumour) Carcinogen = cancer-causing agent that alters DNA e.g. X-rays, UV radiation, tobacco, etc. Prolonged exposure to carcinogens can cause activation of oncogenes & inactivation of tumor-suppressor genes Carcinogens also promote cell division = can lead to cancerous tumors Combo of virus & carcinogen may increase risk of cancer Animation of transcription: http://vcell.ndsu.nodak.edu/animations/transcription/movie.htm Animation of translation: http://vcell.ndsu.nodak.edu/animations/translation/movie.htm