* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download metabolic pathways - MPG Systems Biology Forum
Magnesium in biology wikipedia , lookup
Lactate dehydrogenase wikipedia , lookup
Fatty acid metabolism wikipedia , lookup
Mitochondrion wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Biosynthesis wikipedia , lookup
Photosynthesis wikipedia , lookup
Metabolomics wikipedia , lookup
Electron transport chain wikipedia , lookup
Light-dependent reactions wikipedia , lookup
Biochemical cascade wikipedia , lookup
Microbial metabolism wikipedia , lookup
Pharmacometabolomics wikipedia , lookup
Basal metabolic rate wikipedia , lookup
Nicotinamide adenine dinucleotide wikipedia , lookup
Biochemistry wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Oxidative phosphorylation wikipedia , lookup
Citric acid cycle wikipedia , lookup
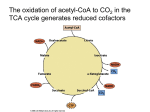
Elucidating the Roadmap of Metabolism by Pathway Analysis Stefan Schuster Friedrich Schiller University Jena Dept. of Bioinformatics Topics of this talk: Introduction • Metabolism is bridge between genotype and phenotype • Networks of metabolic reactions are complex due to their size and the presence of bimolecular reactions • Many kinetic parameters unknown. Maximal velocities depend on enzyme concentrations „Roadmap“ as a metaphor Roads form simple graph Metabolism is hypergraph Technologically relevant metabolic syntheses: • • • • • • Antibiotics by fungi Ethanol by yeast Amino acids by bacteria Dyes Perfumes etc. etc. Theoretical Methods • • • • • • Dynamic Simulation Stability and bifurcation analyses Metabolic Control Analysis (MCA) Metabolic Pathway Analysis Metabolic Flux Analysis (MFA) Optimization, Evolutionary Game Theory • and others Theoretical Methods • • • • • • Dynamic Simulation Stability and bifurcation analyses Metabolic Control Analysis (MCA) Metabolic Pathway Analysis Metabolic Flux Analysis (MFA) Optimization, Evolutionary Game Theory • and others Metabolic Pathway Analysis (or Metabolic Network Analysis) • Decomposition of the network into the smallest functional entities (metabolic pathways) • Does not require knowledge of kinetic parameters!! • Uses stoichiometric coefficients and reversibility/irreversibility of reactions History of pathway analysis • „Direct mechanisms“ in chemistry (Milner 1964, Happel & Sellers 1982) • Clarke 1980 „extreme currents“ • Seressiotis & Bailey 1986 „biochemical pathways“ • Leiser & Blum 1987 „fundamental modes“ • Mavrovouniotis et al. 1990 „biochemical pathways“ • Fell (1990) „linearly independent basis vectors“ • Schuster & Hilgetag 1994 „elementary flux modes“ • Liao et al. 1996 „basic reaction modes“ • Schilling, Letscher and Palsson 2000 „extreme pathways“ Mathematical background Steady-state condition NV(S) = 0 If the kinetic parameters were known, this could be solved for S. If not, one can try to solve it for V. The equation system is linear in V. However, usually there is a manifold of solutions. Mathematically: kernel (null-space) of N. Spanned by basis vectors. These are not unique. non-elementary flux mode elementary flux modes S. Schuster et al.: J. Biol. Syst. 2 (1994) 165-182; Trends Biotechnol. 17 (1999) 53-60; Nature Biotechnol. 18 (2000) 326-332 An elementary mode is a minimal set of enzymes that can operate at steady state with all irreversible reactions used in the appropriate direction All flux distributions in the living cell are non-negative linear combinations of elementary modes Related concept: Extreme pathway (C.H. Schilling, D. Letscher and B.O. Palsson, J. theor. Biol. 203 (2000) 229) - distinction between internal and exchange reactions, all internal reversible reactions are split up into forward and reverse steps Mathematical background (2) Steady-state condition NV = 0 Sign restriction for irreversible fluxes: Virr 0 This represents a linear equation/inequality system. Solution is a convex region. All edges correspond to elementary modes. In addition, there may be elementary modes in the interior. Geometrical interpretation Elementary modes correspond to generating vectors (edges) of a convex polyhedral cone (= pyramid) in flux space (if all modes are irreversible) Rate 3 Rate 2 generating vectors Rate of enzyme 1 Software for computing elementary modes EMPATH (in SmallTalk) - J. Woods METATOOL (in C++) - Th. Pfeiffer, F. Moldenhauer, A. von Kamp Included in GEPASI - P. Mendes and JARNAC - H. Sauro part of METAFLUX (in MAPLE) - K. Mauch part of FluxAnalyzer (in MATLAB) - S. Klamt part of ScrumPy (in Python) - M. Poolman Alternative algorithm in MATLAB – C. Wagner (Bern) On-line computation: pHpMetatool - H. Höpfner, M. Lange http://pgrc-03.ipk-gatersleben.de/tools/phpMetatool/index.php Biochemical Applications: 1. Can sugars be produced from lipids? • Known in biochemistry for a long time that many bacteria and plants can produce sugars from lipids (via C2 units) while animals cannot Glucose AcCoA is linked with glucose by a chain of reactions. However, no elementary mode realizes this conversion in the absence of the glyoxylate shunt. CO2 PEP Pyr AcCoA Cit Oxac CO2 IsoCit Mal CO2 OG Fum Succ SucCoA CO2 Glucose Elementary mode representing conversion of AcCoA into glucose. It requires the glyoxylate shunt. CO2 PEP AcCoA Pyr Cit Oxac CO2 Mal Mas Gly Icl IsoCit OG Fum Succ SucCoA CO2 CO2 The glyoxylate shunt is present in green plants and many bacteria (e.g. E. coli). This example shows that a description by usual graphs in the sense of graph theory is insufficient… S. Schuster, D.A. Fell: Modelling and simulating metabolic networks. In: Bioinformatics: From Genomes to Therapies (T. Lengauer, ed.) Wiley-VCH, Weinheim, in press. A successful theoretical prediction Glucose PEP Pyr Oxac Red elementary mode: Usual TCA cycle Green elementary mode: Catabolic pathway predicted in Liao et al. (1996) and Schuster et al. (1999). Experimental hints in Wick et al. (2001). Experimental proof in: E. Fischer and U. Sauer: CO2 A novel metabolic cycle catalyzes AcCoA glucose oxidation and anaplerosis in hungry Escherichia coli, J. Biol. Chem. 278 (2003) Cit 46446–46451 CO2 Mal IsoCit Gly OG Fum Succ SucCoA CO2 CO2 2. Crassulacean Acid Metabolism (CAM) • Variant of photosynthesis employed by a range of plants (e.g. cacti) as an adaptation to arid conditions • To reduce water loss, stomata are closed during daytime • At nighttime, PEP + CO2 oxaloacetate malate • At daytime, malate pyruvate (or PEP) + CO2 carbohydrates CAM metabolism during daytime Pi hexose Pi starch 7 5 RBP 9 TP TP Pi 2 Pi PEP 3 PEP 11 Pi oxac CO 2 mal 1 cytosol 8 12 4 CO CO Pi 6 Pi 2 pyr 10 pyr chloroplast 2 Elementary modes A) B) P hexose P CO RBP TP TP P P i PEP P CO mal i i P CO P i pyr chloroplast Hexose synthesis via malic enzyme as occurring in Agavaceae and Dracaenaceae Dracaena starch i RBP 2 TP TP P P i PEP P oxac i 2 i CO 2 PEP pyr cytosol P CO P hexose starch i 2 oxac i CO mal i PEP i P i P 2 pyr cytosol 2 i pyr chloroplast Starch synthesis via malic enzyme as occurring in Cactaceae and Crassulacea Ferocactus C) P hexose P CO i RBP TP TP P P i PEP P CO mal i i P CO P starch i RBP 2 i pyr chloroplast Simultaneous starch and hexose synthesis via malic enzyme as occurring in: Clusia minor TP TP P P i PEP P oxac i 2 i 2 PEP pyr cytosol CO P hexose starch P 2 oxac D) i CO mal 2 i PEP i P i P 2 pyr cytosol CO pyr i chloroplast Hexose synthesis via PEPCK as occurring in Clusia rosea and in: Ananus comosus = pineapple F) E) P hexose P RBP P oxac CO mal P CO i P pyr Pi chloroplast Starch synthesis via PEPCK as occurring in Asclepadiaceae Caralluma hexagona i PEP 2 oxac i 2 starch RBP TP PEP pyr cytosol CO 2 Pi i PEP Pi Pi TP P 2 hexose starch i TP CO i CO mal Pi cytosol TP Pi PEP P i 2 pyr CO 2 pyr Pi chloroplast Simultaneous starch and hexose synthesis via PEPCK as occurring in: Aloe vera „Pure“ pathways • In a review by Christopher and Holtum (1996), only cases A), B), D), and E) were given as “pure” functionalities. F) was considered as a superposition, and C) was not mentioned. • However, F) is an elementary mode as well, although it produces two products. It does not use the triose phosphate transporter • The systematic overview provided by elementary modes enables one to look for missing examples. Case C) is indeed realized in Clusia minor (Borland et al, 1994). • Interestingly, (almost) pure elementary modes are realized here, although this should reduce robustness S. Schuster, D.A. Fell: Modelling and simulating metabolic networks. In: Bioinformatics: From Genomes to Therapies (T. Lengauer, ed.) Wiley-VCH, Weinheim, in press. 3. Adenine and adenosine salvage pathways • Human erythrocytes cannot synthesize nucleotide phosphates de novo • They can recycle nucleotides to give nucleotide phosphates • In particular, they can recycle adenine and adenosine, but not hypoxanthine S. Schuster, D. Kenanov: Adenine and adenosine salvage pathways in erythrocytes and the role of S-adenosylhomocysteine hydrolase – A theoretical study using elementary flux modes. FEBS J. 272 (2005) 5278-5290. membrane DPGM GLCim GLCext GLC HK PGI G6P G6PDH GL6PDH ADP 2GSH GSSGR RU5P R5P NADH ADP GSSG TKI S7P TA F6P X5P GA3P PYRext LACtrans LAC LACext PRM ATP HGPRT ADPRT ADENINE AMP AMPDA SAHH2 ApK S-AdoHcy MetAcc Acc MT SAM SAMext ATP AMP ADP SAHH1 HCY AK ADP NUC ADA NUC ADO + AMP R1P PRPP HYPX IMP PRPP 3'KetoRibose HCY K PYRtrans E4P R5P ATP leak PYR NAD GA3P ADP + ATP NADH LDH Na K ADP EN PEP PK PRPPsyn NaK ATPase K PGM 2PG GSHox + + 3PG ATP DHAP leak + NAD PGK 1,3 DPG TKII Xu5PE Na GAPDH GA3P TPI NADPH R5PI CO2 ALD FDP NADP GL6P PGLase GO6P + PFK ATP ATP ADP Na F6P 2,3DPG DPGase INO HXtrans PNPase HYPXext Question • As salvage pathways use enzymes consuming ATP as well as enzymes producing ATP, it is not easy to see whether a net synthesis of ATP is possible. • Invest ATP to gain ATP - Bootstrapping like Baron Münchhausen? Goal: • Analyse theoretically how many salvage pathways exist • Which enzymes involves each of these and in what flux proportions (i.e. relative fluxes) • Compute the net overall stoichiometry of ATP anabolism • Medical impact of enzyme deficiencies External metabolites • Adenine or adenosine depending on which salvage is analysed • glucose, lactate, CO2, ATP • hypoxanthine, sodium and potassium outside the cell Adenine as a source • • • • 153 elementary modes 4 of these produce ATP ATP per adenine yield: 1:1 ATP per glucose yields: 3:10, 2:7, 1:4, 1:6 membrane DPGM GLCim GLCext GLC HK PGI G6P G6PDH GL6PDH ADP 2GSH GSSGR RU5P R5P NADH ADP GSSG TKI S7P TA F6P X5P GA3P PYRext LACtrans LAC LACext PRM ATP HGPRT ADPRT ADENINE AMP AMPDA SAHH2 ApK S-AdoHcy MetAcc Acc MT SAM ATP AMP ADP INO HXtrans PNPase HCY AK ADP NUC ADA NUC ADO + R1P PRPP HYPX IMP PRPP 3'KetoRibose HCY K PYRtrans E4P R5P ATP leak PYR NAD GA3P ADP + ATP NADH LDH Na K ADP EN PEP PK PRPPsyn NaK ATPase K PGM 2PG GSHox + + 3PG ATP DHAP leak + NAD PGK 1,3 DPG TKII Xu5PE Na GAPDH GA3P TPI NADPH R5PI CO2 ALD FDP NADP GL6P PGLase GO6P + PFK ATP ATP ADP Na F6P 2,3DPG DPGase AMP SAHH1 SAMext Elementary mode with highest ATP:glucose yield (3:10) HYPXext membrane DPGM GLCim GLCext GLC HK PGI G6P G6PDH GL6PDH ADP 2GSH GSSGR RU5P R5P NADH ADP GSSG TKI S7P TA F6P X5P GA3P PYRext LACtrans LAC LACext PRM ATP HGPRT ADPRT ADENINE AMP AMPDA SAHH2 ApK S-AdoHcy MetAcc Acc MT SAM ATP AMP ADP INO HCY AK ADP NUC ADA NUC ADO + R1P PRPP HYPX IMP PRPP 3'KetoRibose HCY K PYRtrans E4P R5P ATP leak PYR NAD GA3P ADP + ATP NADH LDH Na K ADP EN PEP PK PRPPsyn NaK ATPase K PGM 2PG GSHox + + 3PG ATP DHAP leak + NAD PGK 1,3 DPG TKII Xu5PE Na GAPDH GA3P TPI NADPH R5PI CO2 ALD FDP NADP GL6P PGLase GO6P + PFK ATP ATP ADP Na F6P 2,3DPG DPGase AMP SAHH1 SAMext Elementary mode with lowest ATP:glucose yield (1:6) HXtrans PNPase HYPXext Adenosine as a source • 97 elementary modes • 12 of these produce ATP • ATP per adenosine yield: 1:1, 2:3, 5:8, 8:17, 2:5, 5:14, and 1:4. • ATP per glucose yields: 1:1, 5:6, 2:3, 5:9, 1:3, 1:6, and: 1:0!! • When ATP/glucose = 1:0, then all adenosine is used as exclusive energy source membrane DPGM GLCim GLCext GLC HK PGI G6P G6PDH GL6PDH ADP 2GSH GSSGR RU5P R5P NADH ADP GSSG TKI S7P TA F6P X5P GA3P PYRext LACtrans LAC LACext PRM ATP HGPRT ADPRT ADENINE AMP AMPDA SAHH2 ApK S-AdoHcy MetAcc Acc MT SAM ATP AMP ADP INO HXtrans HYPXext PNPase HCY AK ADP NUC ADA NUC ADO + R1P PRPP HYPX IMP PRPP 3'KetoRibose HCY K PYRtrans E4P R5P ATP leak PYR NAD GA3P ADP + ATP NADH LDH Na K ADP EN PEP PK PRPPsyn NaK ATPase K PGM 2PG GSHox + + 3PG ATP DHAP leak + NAD PGK 1,3 DPG TKII Xu5PE Na GAPDH GA3P TPI NADPH R5PI CO2 ALD FDP NADP GL6P PGLase GO6P + PFK ATP ATP ADP Na F6P 2,3DPG DPGase AMP SAHH1 SAMext One elementary mode using adenosine as energy and nucleotide source 2.3-Diphosphoglycerate bypass • Neither for adenine nor adenosine salvage, any elementary mode involves the 2.3DPG bypass • ATP balance would not be positive anymore Molar investment ratio moles of ATP consumed moles of ATP produced - moles of ATP consumed In glycolysis: 2 =1 4-2 In elementary mode 1 of adenine salvage, this ratio is 18:(20-18) = 9:1. S. Schuster, D. Kenanov: FEBS J. 272 (2005) 5278-5290. Maximization of tryptophan:glucose yield Model of 65 reactions in the central metabolism of E. coli. 26 elementary modes. 2 modes with highest tryptophan: glucose yield: 0.451. PEP Pyr S. Schuster, T. Dandekar, D.A. Fell, Trends Biotechnol. 17 (1999) 53 Glc 233 G6P Anthr 3PG PrpP GAP 105 Trp Conclusions • Elementary modes are an appropriate concept to describe biochemical pathways • Information about network structure can be used to derive far-reaching conclusions about performance of metabolism • Elementary modes reflect specific characteristics of metabolic networks such as steady-state mass flow, thermodynamic constraints and and the systemic interactions (Systems Biology) Conclusions (2) • It can be tested whether connected routes can function at steady state • A complete list of potential pathways can be generated. Thereafter, experimental search for realized pathways. • Pathway analysis is well-suited for computing maximal and submaximal molar yields Cooperations •Steffen Klamt, Jörg Stelling, Ernst Dieter Gilles (MPI Magdeburg) •Thomas Dandekar (U Würzburg) •David Fell (Brookes U Oxford) •Thomas Pfeiffer, Sebastian Bonhoeffer (ETH Zürich) •Thomas Wilhelm (IMB Jena) •Reinhart Heinrich, Thomas Höfer (HU Berlin) •Marko Marhl (U Maribor, Slovenia) •Hans Westerhoff (VU Amsterdam) • and others •Acknowledgement to DFG and BMBF for financial support Current group members • • • • • • • Dr. Axel von Kamp Dr. Ina Weiß Dimitar Kenanov Jörn Behre Beate Knoke Ralf Bortfeldt Gunter Neumann