* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Ecological speciation model
Biosequestration wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Mitochondrion wikipedia , lookup
Isotopic labeling wikipedia , lookup
Basal metabolic rate wikipedia , lookup
Metalloprotein wikipedia , lookup
Fatty acid metabolism wikipedia , lookup
NADH:ubiquinone oxidoreductase (H+-translocating) wikipedia , lookup
Phosphorylation wikipedia , lookup
Glyceroneogenesis wikipedia , lookup
Photosynthetic reaction centre wikipedia , lookup
Photosynthesis wikipedia , lookup
Blood sugar level wikipedia , lookup
Electron transport chain wikipedia , lookup
Light-dependent reactions wikipedia , lookup
Lactate dehydrogenase wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Microbial metabolism wikipedia , lookup
Nicotinamide adenine dinucleotide wikipedia , lookup
Adenosine triphosphate wikipedia , lookup
Biochemistry wikipedia , lookup
Oxidative phosphorylation wikipedia , lookup
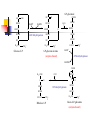
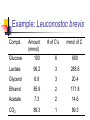
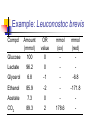
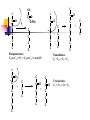
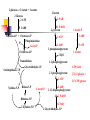
Heterofermentative Pathway Uses part of the pentose phosphate pathway Only one pyruvate is made Have a decarboxylation and C-C cleavage to give a C3 and a C2 Overview: Activation-use 1 ATP Two oxidations done to make ß-carbonyl C-C bond cleavage G3P to pyruvate like in Streptococcus Less ATP because more ox/red reactions Heterofermentative Pathway in Leuconostoc sp. Glucose ATP ADP hexokinase Glucose-6-P NADP+ G6P dehydrogenase NADPH [6-P-Gluconolactone] H2O 6-P-gluconate NADP + CO2 6PG dehydrogenase NADPH Ribulose-5-P R5P epimerase Xylulose-5-P Glyceraldehyde-3-P NAD+ PGALD dehydrogenase NADH 1,3-bisphosphoglycerate ADP PGA kinase ATP 3-phophoglycerate phosphoketolase NADPH alcohol dehydrogenase Acetaldehyde NADP+ acetaldehide dehydrogenase NADPH Acetyl-CoA Pi phosphotransacetylase CoA Acetyl-P ADP ATP Acetate Small What would happen if the organism could divert electrons away from ethanol production? amount ATP yield used 1 ATP made 2 ATP net yield= 1 ATP PGA mutase 2-phosphoglycerate enolase H2O phosphoenolpyruvate ADP pyruvate kinase ATP pyruvate Ethanol NADP + Additional oxidation/reduction reactions decrease potential ATP yield. NADH NAD + Lactate lactate dehydrogenase 6-P-gluconate HC HC OH NADP+ HCOH HOCH HCOH NADPH G6P dehydrogenase HC H 2 CO PO3= HOCH O HCOH HCOH HC HCOH H 2 CO PO3= 6-P-glucono- lactone Glucose-6-P HCOH H2O HOCH O HCOH COOH O PO3= H 2 CO NADP + (enzyme-bound) 6PG dehydrogenase NADPH COOH CO 2 H 2 COH C C O HCOH 6PG dehydrogenase O HCOH HCOH HCOH H 2 CO HCOH PO = 3 Ribulose-5-P H 2 CO PO3= 3-keto-6-P-gluconate (enzyme-bound) Mechanism of beta-decarboxylation O 1 C O- C C CO2 2 3 C C H+ O- O 1) Carbonyl accepts electrons from C-C between the 1 and 2 carbons. 2) Carbon dioxide and an enolate are formed. 3) Re-shifting of the electrons forms the keto sugar. H C C O Phosphoketolase Reaction CH 3 H 2 COH C C O + O-PO 3= HOCH O O-PO 3= HC HCOH H 2 CO Acetyl-Phosphate O HCOH PO3= H 2 CO Glyceraldehyde-3-P PO = 3 Xylulose-5-P Enzyme contains Thiamin pyrophosphate (TPP) as cofactor the function here is transketolation. E-TPP- CH OH CH 2 OH E-TPP C OH CH 2 O-PO3 = Acetyl-Phosphate Conclusion Heterofermentative organisms use a pathway with a greater number of redox reactions than Streptococcus. Make very oxidized and very reduced compounds. More NAD(P)H to be reoxidized constrains ATP synthesis, high energy intermediate used as an electron acceptor. Vitamins: essential portions of cofactors that organism can not make de novo Fermentation Analysis In order to understand how an organism makes its energy or what biochemical pathways are present, one must first know what the products of metabolism are. First Law of Thermodynamics: mass is conserved must account for all of the carbon and electrons originally present in the substrate. Fermentation analysis From this, we can then figure out the pathways and amount of ATP made. Also, inspection of the products will allow us to make predictions about the cell’s metabolism. Initially, we will look at glucose consumption in rich medium Growth factors from media supply cell carbon Most of glucose goes to products, only 5-10% incorporated into cells. In industry, one must also account for cell mass. Experimental set up Glucose added and inoculated Control: inoculated but without glucose; correct for products made from other medium components or brought in with inoculum. Take time zero and time final samples and measure Glucose and product formation. Example: Leuconostoc brevis Compd. Glucose Amount (mmol) 100 # of C’s mmol of C 6 600 Lactate 96.2 3 288.6 Glycerol 6.8 3 20.4 Ethanol 85.9 2 171.8 Acetate 7.3 2 14.6 CO2 89.3 1 89.3 Have we detected all of the products? carbon Calculate the carbon recovery by multiplying the amount detected by the number of carbon atoms for each compound, then sum up all of the carbon in the products. Carbon in glucose = 6 X 100 mmoles =600 mmoles Carbon in products = (288.6 + 20.4 + 171.8 + 14.6 + 89.3) mmoles Carbon in products = 584.7 mmoles % C recovery = (584.7 mmol/600 mmol) * 100 % C recovery = 97.4% Have detected all of the electrons? In a fermentation, electrons removed from glucose are added back to a compound derived from glucose. Thus, the ratio of oxidized products to reduced products must equal 1. Since glucose (C6H12O6) has 2 H’s for every O, products with more than 2 H’s per O have been reduced, and products with less than 2 H’s per O have been oxidized. OR value of a compound • To calculate the OR value of a compound, give a numerical score of +1 for every O and -1 for every 2 H’s. • Examples: Glucose (C6H12O6): 6O is +6, 12 H's is -6, 6-6=0 Lactate (C3H6O3): 3O is +3, 6H's is -3, 3-3=0 Acetate (C2H4O2): 2O is +2, 4H's is -2, 2-2=0 Glycerol (C3H8O3): 3O is +3, 8 H's is -4, 3-4 = -1 Ethanol (C2H6O): 1O is +1, 6 H is -3, 1-3= -2 Carbon dioxide (CO2): 2 O's = +2 Example: Leuconostoc brevis Compd Amount (mmol) Glucose 100 OR value 0 mmol (ox) - mmol (red) - Lactate 96.2 0 - - Glycerol 6.8 -1 - -6.8 Ethanol 85.9 -2 - -171.8 Acetate 7.3 0 - - CO2 89.3 2 178.6 - O/R ratio of the fermentation • Once the OR value of the compound is determined this is multiplied by the amount detected (see Table) • Calculate the O/R ratio OR ratio = |178.6/(-171.8)+(-6.8)| OR ratio = 178.6/178.6 = 1.0 Ratios close to 1 mean all of the electrons have been accounted for. C1 to C2 ratio • A common C-C cleavage reaction is C3 --> C1 + C2 usually indicating pyruvate is an intermediate. • If this occurs in your organism, then expect a C1/C2 ratio of 1. C1 = 89.3 mmoles C2 = 85.9 mmoles + 7.3 mmoles = 93.2 mmoles C1 to C2 ratio = 89.3 mmoles/ 93.2 mmoles C1 to C2 ratio = 0.96 Value is close to one so probably have pyruvate cleavage. Conclusion Fermentation balance is the first step in understanding the metabolism of an organism Must have C recovery close to 100% and an O/R ratio close to 1. C1/C2 ratio indicates pyruvate cleavage You can use the above information in the lab to determine what analyses are needed to complete the balance. What happens if an alternate electron acceptor is present in a fermentation? Electron flow dictates carbon flow and energy yield Alternate electron acceptors provide fermentative bacteria a “choice” The result will be less lactate and ethanol and more acetate and ATP are made. We will study the effect of oxygen on the metabolism of lactic acid bacteria De Felipe et al., J. Bacteriol. vol 180, p 3804, 1998 Utilization of oxygen by facultative lactic acid bacteria. Some lactic acid bacteria possess enzymes that reoxidize NADH (and NADPH) by reducing oxygen to water (Dolin’s enzymes) Oxidase NAD(P)H + H+ + O2 --> H2O2 + NAD(P)+ Peroxidase NAD(P)H + H+ + H2O2 --> 2 H2O + NAD(P)+ What happens when oxygen is present? When oxygen and Dolin’s enzymes are present, NAD(P)H is reoxidized by reducing oxygen to water rather than pyruvate to lactate or acetyl-P to ethanol. More acetate and ATP, less ethanol and lactate, are made. Make more ATP Acetate kinase acetyl-P + ADP --> acetate + ATP For every acetate made, one ATP is made by substratelevel phosphorylation by this reaction. When Dolin’s enzymes and oxygen are present, 1) acetyl-P goes to acetate and ATP rather than to ethanol, and 2) pyruvate is metabolized to acetate and CO2 rather than to lactate. CO2 Pyruvate NAD+ NADH Acetyl-CoA Acetyl-P Pi CoA Acetate ADP ATP Streptococcus sp. and Dolin’s enzymes No O2 or Dolin's enzymes With O2 and Dolin's enzymes Glucose Glucose 2 NAD+ 2 ATP net 2 NADH 2 Pyruvate 2 NADH 2 NAD+ 2 Lactate 2 NAD+ 2 ATP net 2 NADH 2 Pyruvate 2 CoA 2 NAD+ 2 CO2 2 NADH 2 Acetyl-CoA 2 Pi 2 CoA 2 O2 4 H 2O 2 Acetyl-P 2 ADP 2 ATP 2 Acetate Net of 4 ATP 4 NADH 4 NAD+ Summary If there is an alternate electron acceptor, less lactate, more acetate, CO2, and ATP Bifidobacterium sp. Bifid or 2 lobes Gram positive, curved rod found in the feces of breast-fed infants, Requires many growth factors, including N-acetylglucosamine Makes 2 lactate and 3 acetate from 2 glucose Makes high-energy intermediate by phosphoketolase reaction rather than ox/red. Schell, M. A. et. al. (2002). The genome sequence of Bifidobecterium longum reflects its adaptation to the human gastrointestinal tract. PNAS V99 N22 p14422-14427. Outline of pathway Activation: 2 glucose to 2 glucose-6-P uses 2 ATP Make 2 G3P and 3 acetyl-P from 2 glucose by transketolase, transaldolase, and phsophoketolase reactions 2 G3P to 2 lactate by reactions seen in Streptococcus 3 acetyl-P to 3 acetate and 3 ATP by acetate kinase Synthesis of 2 G-3-P and 3 C2 units from 2 glucose. Phosphoketolase : C6 -> C4 + acetyl-P Transaldolase: C6 + C4 -> C7 + C3 Transketolae: C7 + C3 -> C5 + C5 Phosphoketolase: C5 + C5 -> 2 C3 + 2 acetyl-P Net Result: 2 C6 -> 2 C3 + 3 acetyl-P Uses these enzymes to interconvert hexoses and pentoses. C C C CH 3 C O C Pi C O O-PO3 C C C C C C O C + C C C C C Phosphoketolase: C6 (or C5) + Pi -> C4 (or C3) + acetyl-P Transaldolase: C6 + C4 -> C7 + C3 C C O C + C C C C C C C C C C C C C C C O + C C O O C C C C C C C Transketolase C7 + C3 -> C5 + C5 + C C C C C C 2 glucoses -> 2 lactate + 3 acetate 2 lactate 2 Glucose 2 ATP 2 NAD+ 2 NADH 2 pyruvate 2 ADP Fructose-6-P + Fructose-6-P Phosphoketolase Pi Acetyl-P 2 ATP 2 ADP 2 phosphoenolpyruvate Erythrose-4-P 2 H2O Transaldolase Sedoheptuloase-7-P Glyceraldehyde-3-P Ribulose-5-P 2 Xylulose-5-P 3 ATP 3 Acetate ATP yield: 2 3-phosphoglycerate Ribose-5-P 3 ADP 2 2-phosphoglycerate 2 ATP Xylulose-5-P 3 Acetyl-P 2 Acetyl-P 2 ADP 2 1,3-bisphosphoglycerate 2 NADH 2 NAD+ 2 Glyceraldehyde-3-P (7-2)/2 glucose = 2.5 ATP/ glucose Summary Make acetyl-P by phosphoketolase rather than by ox/red reaction Don’t have to use acetyl-P as electron acceptor ATP yield higher than other anaerobes for this reason. Avoidance of ox/red leads to higher ATP gain.