* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Receptors as Drug Targets
Discovery and development of non-nucleoside reverse-transcriptase inhibitors wikipedia , lookup
Drug interaction wikipedia , lookup
Discovery and development of cephalosporins wikipedia , lookup
Discovery and development of neuraminidase inhibitors wikipedia , lookup
Discovery and development of tubulin inhibitors wikipedia , lookup
Discovery and development of direct thrombin inhibitors wikipedia , lookup
Discovery and development of beta-blockers wikipedia , lookup
Psychopharmacology wikipedia , lookup
Discovery and development of integrase inhibitors wikipedia , lookup
Discovery and development of TRPV1 antagonists wikipedia , lookup
Discovery and development of direct Xa inhibitors wikipedia , lookup
5-HT2C receptor agonist wikipedia , lookup
NMDA receptor wikipedia , lookup
CCR5 receptor antagonist wikipedia , lookup
Drug design wikipedia , lookup
Toxicodynamics wikipedia , lookup
5-HT3 antagonist wikipedia , lookup
Discovery and development of angiotensin receptor blockers wikipedia , lookup
Neuropsychopharmacology wikipedia , lookup
Neuropharmacology wikipedia , lookup
Cannabinoid receptor antagonist wikipedia , lookup
Discovery and development of antiandrogens wikipedia , lookup
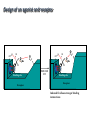
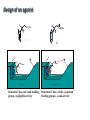
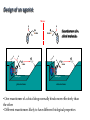
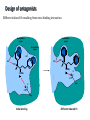
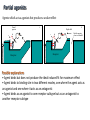
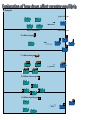
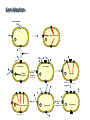
Receptors as Drug Targets Notes on Drug Design M M M RE RE R Signal transduction • Agonists are drugs designed to mimic the natural messenger • Agonists should bind and leave quickly - number of binding interactions is important • Antagonists are drugs designed to block the natural messenger • Antagonists tend to have stronger and/or more binding interactions, resulting in a different induced fit such that the receptor is not activated. Design of agonists •Agonists mimic the natural messenger of a receptor • Agonists bind reversibly to the binding site and produce the same induced fit as the natural messenger - receptor is activated • Similar intermolecular bonds formed as with natural messenger • Agonists are often similar in structure to the natural messenger • The agonist must have the correct binding groups • The binding groups must be correctly positioned to interact with complementary binding regions • The drug must have the correct shape to fit the binding site Agonist Agonist Agonist Induced fit RE RE R Signal transduction Design of a agonist and receptor Binding groups van der Waals binding region H-bond binding region O H H O2C OH NH2Me Neurotransmitter Binding site Receptor Ionic binding region Design of an agonist and receptor H O H H O O NH2Me O C 2 Receptor O NH2Me H H Binding site O C 2 H INDUCED FIT Binding site Receptor Induced fit allows stronger binding interactions Design of an agonist Ionic binding group H-bonding group HO van der Waals -bonding group NH2Me H H2N NH2Me HO NHMe HO H H H Me Hypothetical neurotransmitter NH2Me H Possible agonists with similar binding groups Compare Binding groups: • Identify important binding interactions in natural messenger • Agonists are designed to have functional groups capable of the same interactions • Usually require the same number of interactions Design of an agonist H H CH2Me NH2Me H H I II H H O O O H CH2Me 2 C H Binding site Receptor Structure I has one weak binding group - negligible activity H NH2Me O 2 C H Binding site Receptor Structure II has 2 of the 3 required binding groups - weak activity Design of an agonist No interaction OH OH H H O N H 2M e H NH M e 2 O 2C H H Binding site 2 Interactions only • Binding groups must be positioned such that they can interact with complementary binding regions at the same time • Example has three binding groups, but only two can bind simultaneously • Example will have poor activity Design of an agonist Mirror H H O H H O Enantiomers of a chiral molecule O NH2Me MeH2N H H H O O O NH2Me H Binding site 3 interactions 2 C H O NH2Me 2 C OH Binding site 2 interactions • One enantiomer of a chiral drug normally binds more effectively than the other • Different enantiomers likely to have different biological properties Design of an agonist H Steric block O H N H2 CH 3 No Fit H O Me Me Steric block O2 C Binding site • Agonist must have correct size and shape to fit binding site • Groups preventing access are called steric shields or steric blocks Design of antagonists • Antagonists bind to the binding site but fail to produce the correct induced fit receptor is not activated • Normal messenger is blocked from binding Perfect Fit (No change in shape) H H H O N O Me H Binding site H O C 2 Design of antagonists OH O C 2 Receptor binding site Extra binding regions Design of antagonists Antagonists can form binding interactions with extra binding regions neighboring the binding site for the natural messenger Hydrophobic binding region Extra hydrophobic binding region HO H-bond binding region HO H NH2Me Hypothetical neurotransmitter Ionic binding region - O O Asp Design of antagonists Induced fit resulting from binding of the normal messenger Hydrophobic region Hydrophobic region HO HO HO Induced fit HO H NH2Me H NH2Me - O O O Asp O Asp Design of antagonists Different induced fit resulting from extra binding interaction Hydrophobic region Hydrophobic region Hydrophobic region HO HO HO HO HO H H NHMe - NHMe O O - Asp O O Asp Initial binding Different induced fit Irreversible antagonists X Covalent Bond X OH OH O Irreversible antagonism • Antagonist binds irreversibly to the binding site • Different induced fit means that the receptor is not activated • Covalent bond is formed between the drug and the receptor • Messenger is blocked from the binding site • Increasing messenger concentration does not reverse antagonism • Often used to label receptors Irreversible antagonists O O HO Cl Cl N Cl Cl Propylbenzilylcholine mustard 1 Cl Irreversible binding Nu Nu Nu Nu Antagonist binding site Receptor Agonist binding site 2 Cl Receptor Allosteric antagonists Binding site unrecognisable Binding site ACTIVE SITE (open) Receptor ENZYME Allosteric binding site Induced fit (open) Receptor ENZYME Antagonist • Antagonist binds reversibly to an allosteric binding site • Intermolecular bonds formed between antagonist and binding site • Induced fit alters the shape of the receptor • Binding site is distorted and is not recognised by the messenger • Increasing messenger concentration does not reverse antagonism Antagonists by the umbrella effect • Antagonist binds reversibly to a neighbouring binding site • Intermolecular bonds formed between antagonist and binding site • Antagonist overlaps the messenger binding site • Messenger is blocked from the binding site messenger Binding site for antagonist Binding site for messenger Receptor Antagonist Receptor Partial agonists Agents which act as agonists but produce a weaker effect Partial agonist H H O 1 Slight shift H H NHM e O O O 2C H Receptor NHM e O 2C 2 Partial opening of an ion channel O H Receptor Possible explanations • Agent binds but does not produce the ideal induced fit for maximum effect • Agent binds to binding site in two different modes, one where the agent acts as an agonist and one where it acts as an antagonist • Agent binds as an agonist to one receptor subtype but as an antagonist to another receptor subtype Inverse agonists Properties shared with antagonists • Bind to receptor binding sites with a different induced fit from the normal messenger • Receptor is not activated • Normal messenger is blocked from binding to the binding site Properties not shared with antagonists • Block any inherent activity related to the receptor (e.g. GABA receptor) • Inherent activity = level of activity present in the absence of a chemical messenger • Receptors are in an equilibrium between constitutionally active and inactive forms Explanation of how drugs affect receptor equilibria A) Resting state Agonist binding site Inactive conformations B) Addition of agonist C) Addition of antagonist D) Addition of inverse agonist E) Addition of partial agonist Active conformation Desensitization • Receptors become desensititized on long term exposure to agonists • Prolonged binding of agonist leads to phosphorylation of receptor • Phosphorylated receptor changes shape and is inactivated • Dephosphorylation occurs once agonist departs 1 H Ion channel O2C (closed) O Agonist Receptor NH3 Desensitization • Receptors become desensititized on long term exposure to agonists • Prolonged binding of agonist leads to phosphorylation of receptor • Phosphorylated receptor changes shape and is inactivated • Dephosphorylation occurs once agonist departs O2C H O Agonist NH3 Receptor •Induced fit alters protein shape •Opens ion channel Desensitization • Receptors become desensititized on long term exposure to agonists • Prolonged binding of agonist leads to phosphorylation of receptor • Phosphorylated receptor changes shape and is inactivated • Dephosphorylation occurs once agonist departs O2C H O Agonist Receptor NH3 Desensitization • Receptors become desensititized on long term exposure to agonists • Prolonged binding of agonist leads to phosphorylation of receptor • Phosphorylated receptor changes shape and is inactivated • Dephosphorylation occurs once agonist departs O2C H O Agonist NH3 P Receptor • Phosphorylation alters shape • Ion channel closes • Desensitization Sensitization • Receptors become sensititized on long term exposure to antagonists • Cell synthesises more receptors to compensate for blocked receptors • Cells become more sensitive to natural messenger • Can result in tolerance and dependence • Increased doses of antagonist are required to achieve same effect (tolerance) • Cells are supersensitive to normal neurotransmitter • Causes withdrawal symptoms when antagonist withdrawn • Leads to dependence Sensitization Neurotransmitter Normal response Antagonist No response Receptor synthesis No response Receptor synthesis Sensitization Response Increase antagonist Excess response Dependence No response Stop antagonist Tolerance No response Design of an antagonist for the estrogen receptor His 524 Me OH H H Glu353 H H O Hydrophic skeleton H2O Arg394 Oestradiol • Phenol and alcohol of estradiol are important binding groups • Binding site is spacious and hydrophobic • Phenol group of estradiol is positioned in narrow slot • Orientates rest of molecule • Acts as agonist H Action of the oestrogen receptor Binding site AF-2 regions H12 Coactivator Coactivator Oestradiol DNA Oestrogen receptor Dimerisation & exposure of AF-2 regions Nuclear transcription factor Transcription Design of an antagonist for the estrogen receptor Asp351 N H Side chain His 524 O Glu353 O OH H O S Arg394 Raloxifene Raloxifene is an antagonist (anticancer agent) Phenol groups mimic phenol and alcohol of estradiol Interaction with Asp-351 is important for antagonist activity Side chain prevents receptor helix H12 folding over as lid AF-2 binding region not revealed Co-activator cannot bind Tamoxifen as an antagonist for the estrogen receptor O Me2N CH2CH3 Anticancer agent