* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Human Insulin-Receptor Gene
Epigenetics of neurodegenerative diseases wikipedia , lookup
Gene therapy wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Gene expression profiling wikipedia , lookup
Oncogenomics wikipedia , lookup
Saethre–Chotzen syndrome wikipedia , lookup
Genetic engineering wikipedia , lookup
Genome (book) wikipedia , lookup
Gene expression programming wikipedia , lookup
Neuronal ceroid lipofuscinosis wikipedia , lookup
Gene nomenclature wikipedia , lookup
Frameshift mutation wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Helitron (biology) wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Designer baby wikipedia , lookup
Microevolution wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Point mutation wikipedia , lookup
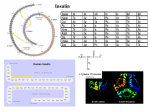
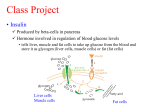
Perspectives in Diabetes Human Insulin-Receptor Gene SUSUMU SEINO, MITSUKO SEINO, AND GRAEME I. BELL The human insulin-receptor (hlNSR) gene spans a region of >120,000 base pairs (bp) on the short arm of chromosome 19. It is comprised of 22 exons or coding regions that vary in size from 36 to >2500 bp. To a large degree, the introns appear to divide the hlNSR gene into segments that encode structural and/or functional elements of the hlNSR protein. The exonintron organization of the hlNSR gene provides a clue to the evolutionary history of this gene and suggests that it is a mosaic constructed from protein-coding regions recruited from other genes. Eight mutations in the hlNSR gene that result in expression of structurally abnormal proteins have been described. These mutations are associated with insulin resistance and provide insight into the role of the hlNSR gene in the development of diabetes mellitus. Diabetes 39:129-33, 1990 nsulin exerts a wide variety of effects on responsive cells, resulting in increased glycogen, lipid, and protein synthesis (1-6). These effects include stimulation of glucose, amino acid and ion uptake, enhanced tyrosine and serine phosphorylation of cellular proteins, modulation of enzymatic activity of regulatory enzymes by dephosphorylation, and stimulation or repression of transcription of specific genes. In addition, insulin promotes cell growth. Although the cellular responses to insulin are diverse and in many instances tissue or cell specific, all are mediated by the insulin receptor (INSR). The human lNSR (hlNSR) is an integral membrane protein present on the surface of all cells. The surface concentration From the Howard Hughes Medical Institute ana the Departments of Medicine. Biochemistry, and Molecular Biology, The University of Chicago, Chicago, llllnois Address c:orrespondence and reprint requests to Susumu Seino, MD, DMS. Howard Hughes Med~calInstitute, The University of Chicago, 5841 South Maryland Adenue, Box 391, Chlcago, IL 60637 Received for publication 30 May 1989 ana accepted in revised form 9 August 1989. DIABETES, VOL. 39, FEBRUARY 1990 of receptors varies widely from as few as 40 on circulating erythrocytes to >200,000 on adipocytes and hepatocytes. The mature hlNSR is a heterotetramer of two a-subunits of 719 or 731 amino acids* and two 620-amino acid p-subunits (7,8). The insulin-binding a-subunit and the membranespanning protein tyrosine kinase p-subunit are generated by proteolytic processing of a common single-chain precursor. Cleavage of the proreceptor occurs during ~ntracellular transfer from the endoplasmic reticulum to the plasma membrane. After cleavage of the proreceptor, the nascent a- and p-subunits are glycosylated on asparagine (N linked) and possibly serinelthreonine residues (0linked). Thus, the aand p-subunits have apparent M, of 135,000 and 95,000, respectively, which are much greater than those predicted from their sequences: the a- and p-subunits have predicted M, of 82,000 or 84,000 and 70,000, respectively. In the absence of the a-subunit, the endogenous tyrosine kinase of the p-subunit is constitutively active, suggesting that one function of the a-subunit is to repress the activity of the tyrosine kinase (9). Insulin binding to the a-subunit probably induces a conformational change in the receptor that relieves this repression. The identification of genetic defects in the hlNSR that are associated with extreme insulin resistance and glucose intolerance has provided new insight into the possible role of the hlNSR in the development of non-insulin-dependent diabetes mellitus (NIDDM) (10-16). It is appropriate to consider the contribution of genetic variation in the hlNSR gene in relation to the development of the common forms of NIDDM. Is there a subpopulation of diabetic patients who have mutations in the hlNSR that impair its function but less severely than those associated with extreme insulin-resistant syndromes? We are in a position to address this important question 'The numbering of the amino acid res~duesof the hlNSR is for a-subunit and precursor of 731 and 1355 amno acids, respectively The four basic amino acids at the proreceptor cleavage site are presumed to be removed from the COOH-terminal of the a-subunit during processing of the proreceptor 129 EXON-INTRON ORGANIZATION OF hlNSR GENE The hlNSR gene is located on the distal short arm of human chromosome 19 in the region of bands p l 3 . 3 4 p13.2 (17). The low-density lipoprotein (LDL)-receptor gene is also in this region of chromosome 19 and is located toward the centromere and -10-15 x lo6 base pairs (bp) from the hlNSR gene (18). Most of the hlNSR gene has been isolated as a series of overlapping DNA fragments in the bacteriophage-A (19). These fragments span a region of >130,000 bp, which includes both the gene and flanking regions (Fig. 1). We have sequenced -13,000 bp of the hlNSR, including the 5'-flanking promoter region, each exon, and part of each intron (19, unpublished observations). The gene is composed of 22 exons and 21 introns. The exons range in size from 36 bp (exon 11) to >2500 bp (exon 22, most of which codes for the 3'-untranslated region of hlNSR mRNA), whereas the 21 introns that separate the exons vary in size from -500 to >25,000 bp (19). All of the introns interrupt protein-coding regions of the gene. The actual size of the hlNSR gene is uncertain because parts of introns 2. 3, 9, and 10 were not isolated, but it must be >120,000 bp. MullerWieland et al. (20) isolated a fragment of the hlNSR gene that contains both exons 1 and 2, and their data indicate that intron 1 is -25,000 bp. The distribution of the exons is rather striking. Exons 1-1 1, which encode the a-subunit of the receptor, are dispersed over >90,000 bp. In contrast, exons 12-22, which encode the p-subunit, are located together in a region of -30,000 bp. The promoter region of the hlNSR gene has also been characterized, and its features are reviewed in Seino et al. (19). The exon organizationof the hlNSR gene appears to reflect the structural organization of the protein, because many of the exons code for structural or functional modules of the protein (Fig. 1). Exons 1-3 code for the signal peptide, part of the insulin-binding region (21), and a very cysteine-rich domain of the protein that may also interact with insulin (22), respectively. The region encoded by exons 2-5 is also homologous to the corresponding region of the epidermal growth factor-receptor gene (19). Residues encoded by exon 6 appear to be located at the interface between adjacent a-subunits in the heterotetrameric form of the hlNSR Mutations Exon Protein domains and contribute to the cooperative site-site binding interactions of the hlNSR (23). Exon 11 is the smallest exon, only 36 bp in size, and alternative splicing of this exon results in the synthesis of hlNSR proteins having a-subunits with different COOH-terminal sequences (7,8,24). Exon 12 codes for the tetrabasic amino acid sequence Arg-Lys-Arg-Arg, which is the site in the proreceptor molecule cleaved by the proteolytic processing enzyme, thereby generating the aand p-subunits. Exon 12 also encodes the NH,-terminal of the p-subunit; 20% of the amino acids in this region of the p-subunit are either serine or threonine, suggesting that the NH,-terminal of the p-subunit may represent a region of Olinked oligosaccharide modification. Exon 15 codes for the membrane-spanning region of the p-subunit. Thus, exons 1-14 encode the extracellular region of the hlNSR, whereas exons 16-22 encode the region that is localized within the cell. Exon 16 codes for a 22-amino acid segment that separates the transmembrane and tyrosine kinase domains. Exons 17-21 code for the protein tyrosine kinase; the amino acids involved in ATP binding by the kinase, residues 10031008 (Gly-Gln-Gly-Ser-Phe-Gly)and 1030 (Lys), are all encoded by exon 17. The tyrosine residues that are autophosphorylated by the kinaseon insulin binding are encoded by exons 20 (Tyr 1158, 1162, and 1163) and 22 (Tyr 1328 and 1334). Exon 22 also codes for the highly charged COOH-terminal of the p-subunit. The data suggest that exons encode functional/structuraI domains of the hlNSR protein and that the hlNSR gene, like the LDL-receptor gene (25,26), is a mosaic constructed from exons recruited from other sources. ALTERNATIVE SPLICING GENERATES AMINO ACID-SEQUENCE ISOFORMS OF hlNSR The a- and (3-subunits are generated by proteolytic processing of a precursor of 1370 or 1382 amino acids, which includes a 27-amino acid signal peptide in addition to the 1343 or 1355 proreceptor molecule (7,8). The difference in size of the a-subunit and its precursors is due to tissuespecific, possibly developmentally regulated, alternative splicing of exon 11, which codes for a 12-amino acid segment at the COOH-terminal of the a-subunit (24). Brain and 0 ./I+ -B+wt+*~ I 2 Signal peptide Insulin binding 00 /' 3 Cysteine rich 4 5 6 7 8 9 10 11 a-subunit association spliced exon - L - V 1 12 13 a -subunit 14 15 16 17 18192021 22 - I J a- and @-subunit association EGF receptor homology \ 0 processing site kinase ~ransmbmbrane Receptor recycling (?) \ 0 0 \ - @-subunit FIG. 1. Map of human insulin-receptor (hlNSR) gene. Relative locations and sizes of exons (m) and introns are indicated. Oblique lines, regions of nonsense; -, deletion. introns 2, 3, 9, and 11 that have not been cloned. Location of mutatlons in hlNSR gene are indicated: 0, mlssense; 0, Table 1 describes mutations in greater detail. Domains of hlNSR protein encoded by each exon or group of exons are below gene map. kb, Kllobase. DIABETES. VOL. 39. FEBRUARY 1990 - spleen express almost exclusively the 719-amino acid asubunit, whereas other tissues, including placenta, liver, kidney, and adipose tissue, express a-subunits of both 719 and 731 amino acids. It is unknown whether both types of asubunit are expressed in an individual cell, thereby generating a mixed population of receptors, including proteins having two a-subunits of either 719 or 731 amino acids and hybrid molecules with one a-subunit of each type. However, because all cultured cell lines that have been examined express only a single type of a-subunit, we suspect that individual cells express only one form of a-subunit, i.e., of either 719 or 731 amino acids. The biological consequences of this diversity of hlNSR isoforms is unknown. Transfection of cultured cells with cloned genes encoding recombinant hlNSR protein representing both hlNSR isoforms has indicated that each is biologically active and capable of binding insulin, stimulating autophosphorylation, and mediating postreceptor responses (27-29). However, preliminary studies suggest that the hlNSR isoform with an a-subunit of 731 amino acids may have reduced affinity for insulin (30). Additional studies of the splicing of hlNSR mRNA might account for some of the observed heterogeneity in size of the protein expressed in different tissues. GENETIC VARIATION IN hlNSR GENE Genetic disorders of extreme hormone resistance provide a unique opportunity to examine the functional consequences of expression of an abnormal receptor. Extreme insulin resistance is associated with three syndromes: the type A syndrome of insulin resistance and acanthosis n~gricans,leprechaun~srn,and the Rabson-Mendenhall syndrome (31). Recent studies have identified eight different mutations in one or both of the INSR alleles of individuals with these syndromes (Fig. 1; Table 1). These mutations affect both the biosynthesis of the receptor and its biochemical properties and include four mutations in the a-subunit (1 patient is a compound heterozygote and has different mutations in each of the parental INSR alleles; 10-12), one at the proreceptor processing site (13), and three in the p-subunit (14-16). Two of the a-subunit mutations (Pro'" and appear to alter the structure of the protein such that posttranslational pro- - cessing and transport of the receptor to the plasma membrane is delayed, thereby resulting in reduced numbers of receptors on the cell surface (1 1,12). Individuals expressing both a normal allele and the Pro2" mutation have mild insulin resistance. Subject 3 is a compound heterozygote expressing two different a-subunit mutations (10; Table 1). There is a nonsense mutation in the codon for amino acid 672 of the paternally derived allele, which results in the synthesis of a truncated 671-amino acid fragment of the a-subunit; such a molecule would be expected to be secreted from the cell and to be biologically inactive. This patient's maternally derived hlNSR allele has a missense mutation (GIu4=O)that results in the expression of a protein with qualitative abnormalities in insulin binding, including increased stability of the insulin-INSR complex. Such a mutation might impair the release of insulin in the acidic endosome after internalization of the insulin-hlNSR complex and thereby affect hlNSR recycling. Cells from the patient's mother, which express both normal and mutant (GIuJ60)INSRs, also exhibited qualitative abnormalities in insulin binding similar to the patient; however, the mother was neither insulin resistant nor diabetic. By contrast, the father, who expresses both the normal INSR protein and the truncated molecule, was moderately insulin resistant with impaired glucose tolerance, although not allele in the paovertly diabetic. It is unclear why the GIu~~O tient is less effective than a normal allele in complementing the nonsense mutation. A mutation at the proreceptor processing site (Ser735)results in the expression of fully glycosylated but uncleaved proreceptors on the cell surface (13). This mutant INSR is capable of binding insulin and undergoing insulin-stimulated autophosphorylation, albeit weakly and at high insulin concentrations, and provides an elegant demonstration that the hlNSR acquires normal insulin affinity and sensitivity only after proteolytic separation of the a- and p-subunits. Three mutations in the p-subunit that represent postbinding defects in insulin action have been described, including two missense mutations (Va11°08and SerlZoo;14,15) and a deletion involving exons 17-22 that results in the expression of a receptor lacking the tyrosine kinase domain (16). All are TABLE 1 Mutations at human insulin-receptor locus associated with insulin resistance in subjects 1-7 Ethnic origin Biochemical properties of mutant protein Leu 233 -+ Pro Phe 382 + Val Lys 460 -, Glu Gln 672 -t AM Dutch Venezuelan American Delayed transport to cell surface Delayed transport to cell surface Qualitative abnormalities in insulin binding Truncated a-subunit 11 10 Arg 735 Japanese Impaired proreceptor processing 13 Gly 1008 -, Val Trp 1200 -, Ser Japanese American 15 14 > 10-kb deletion Japanese Decreased tyrosine kinase activity Decreased receptor affinity and autophosphorylation Decreased tyrosine kinase activity Subject Mutation Proreceptor processing site 4 P-Subunit 5 6 - Ser Exon Ref. 12 16 Subjects 1 2, and 4 are homozygous for indicated mutation. Subject 3 is a compound heterozygote having inherited different mutations from each parent. Subjects 5-7 are heterozygous and express both mutant and normal insulin-receptor proteins. AM, amber (TAG) translation termination codon; kb, Kilobase. DIABETES, 'JOL 39, FEBRUARY 1990 131 associated with decreased or defective tyrosine kinase activity or autophosphorylation. The mutation at residue 1008 is of a conserved glycine residue that is part of the ATP binding site of all protein kinases. The codominant expression of normal lNSRs and receptors having mutations in the tyrosine kinase domain appears to result in suppression of the function of the normal protein. The dominant negative character of these mutations may be due to the fact that the hlNSR is a heterotetramer and that heterotetramers formed from two mutant subunits and mixed heterotetramers having one normal and one mutant subunit are inactive or have reduced activity. As a consequence, only 25% of the receptors would possess full biological activity. In addition to mutations in the hlNSR gene that are associated with insulin resistance, population-association studies suggest that there is also genetic variation in the region of the hlNSR gene that increases (32) or decreases (33) the risk for NIDDM. The increased risk could be due to the expression of abnormal receptors that suppress the function of the normal receptors as suggested above. The decreased risk associated with some hlNSR haplotypes might be due to genetic variation that increases expression of the hlNSR and results in increased numbers of receptors per cell. Along with nucleotide substitutions that result in the expression of an abnormal protein, silent substitutions in the coding region of the hlNSR gene that alter the sequence of the gene and mRNA but rot the protein have been described (8,10,19t). Several restriction-fragment-length polymorphisms have also been reported at the hlNSR locus (3342) and all appear to be located within introns. NIDDM suggested that the INSR gene was not the major susceptibility locus for NIDDM, at least in this population (46), the INSR gene may still contribute to the development of NlDDM in a small but measurable subpopulation of patients. The isolation and characterization of the hlNSR gene and recent technical developments provide a strategy for examining the contribution of this gene to the development of NlDDM by amplifyingindividual exons of the hlNSR gene with the polymerase chain reaction (PCR) and then sequencing the amplified DNA. Where should we look for hlNSR mutations? Because mutations in the tyrosine kinase domain of the hlNSR are associated with severe insulin res~stance in the heterozygous state, whereas heterozygosity for mutations in the a-subunit appears to be associated with lesssevere insulin resistance, the NlDDM subjects should first be classified as to the degree of insulin resistance. In the most-resistant subjects, PCR amplification and sequencing of exons 17-21, which code for the tyrosine kinase domain of the hlNSR, should be the first priority. In the less-resistant patients, the analysis should begin with the amplification and sequencing of exons 2 and 3, which appear to be involved in insulin binding. Examination of the sequences of selected regions of the hlNSR gene in 50-100 well-characterized NlDDM subjects could provide valuable insight into the contribution of this gene to the development of this disorder. ACKNOWLEDGMENTS The studies from our laboratory were supported by the Howard Hughes Medical Institute, the Diabetes Research and Training Center of The University of Chicago (DK-20595), and a gift from Sandoz Research Institute. CONCLUSION The causes of NlDDM are unclear. Although impaired p-cell function and insulin resistance are both features of NIDDM, REFERENCES 1. Kahn CR, Current concepts of the molecular mechanism of insulin action controversy exists as to which defect is primary. Studies of Annu Rev Med 36:429-51,1985 2. Czech MP: The nature and regulation of the insulin receptor Annu Rev patients expressing abnormal insulin (43-45) or INSR proPhysiol 47357-81, 1985 teins (10-1 6) have indicated that mutations in either of these 3. Rosen OM: After insulin binds. Science 237.1452-58.1987 genes can be associated with NIDDM. In fact, insulin gene 4. Goldfine ID: The insulin receptor. molecular biology and transmembrane signalling. Endocr Rev 8235-55, 1987 mutations represent a paradigm for (3-cell defects that can 5. Czech MP, Klariund JK. Yagaloff KA, Bradford AP, Lewis RE. Insulin recause diabetes. Similarly, INSR mutations are an example ceptor signalling: activation of multiple serine kinases. J Biol Chem 2631 1017-20. 1988 of a defect in a res~onsivecell that can contribute to the 6. Kahn CR. White MF: The insul~nreceptor and the molecular mechan~sm development of this'disorder. of insulin action. J Clin Invest 82:115156,1988 In contrast to insulin gene mutations, which are extremely 7. Ullrich A, Bell JR. Chen EY, Herrera R, Petruzzelli LM, Dull TJ, Gray A, Coussens L, Liao Y-C Tsubokawa M, Mason A, Seeburg PH, Grunfeld rare, mutations in the hlNSR gene may be more common. C, Rosen OM, Ramachandran J: Human insulin receptor and its reiat~onM ~ because ~ heterozygous ~ individuals ~ ~ who express ~ ~ ship to,the tyros~nekinase famlly of oncogenes. Nature (Lond) 313:756both normal and mutant receptors may exhibit mild insulin 61.1985 resistance, e.g., parents of subjects 1 and 3 (Table 1; 10,12), 8. Eblna Y, Ellis L, Jarnagin K, Edery M, Graf L, Clauser E, Ou J-H, Mas~arz F, Kan YW, Goldfine ID, Roth RA, Rutter WJ. The human insul~nreceptor it seems reasonable to consider that codominant expression cDNA: the structural basis for hormone activated transmernbrane signalling. Cell 40.747-58,1985 of normal and abnormal hlNSR proteins may contribute to 9. Ellis L. Morgan DO, Clauser E, Roth RA, Rutter WJ: A membrane-anchored the development of NIDDM. Although linkage studies of 20 cytoplasmic domain of the human insulin receptor mediates a constiluAmerican Black families in which at least two siblings had tively elevated insulin-independent uptake of 2-deoxyglucose. Mol Endocrinol 1 :I 5-24, 1987 10. Kadowaki T, Bevins CL. Cama A. Ojamaa K, Marcus-Samuels B, KadotComparlson of hlNSR cDNA and genornic sequences (19)suggested that residue 421 of the a-subunit could be either Ile or Thr and that residue 465 could be Gln or Lys; i.e., these sites were polymorphic. We have reexamined the sequence of the kidney hlNSR cDNA described in ref. 22 by hybr~dization with allele-specific oligonucleotides, and it appears to encode Ile 421 and Gln 465 rather than Thr 121 and Lys 465 as suggested in Table 2 of ref. 12 We have also examined the hlNSR alleles of 3 unrelated individuals by sequence or hybridlzation with allele-specific oligonucleotides, and a l encode proteins having Ile 421 and Gln 465.The polymorphism of amino acids 421 and 465 still requires confirmation. 132 waki H, Be~tzL, McKeon C, Taylor SI: Two mutant alleles of the insulin receptor gene in a patient with extreme insulin resistance. Science 240:787-90,1988 11 Accili D. Frap~erC, Mosthaf L, McKeon C, Elbeln S, Permutt MA Ramos E. Lander E, Ullrich A, Taylor SI: A mutation in the insul~nreceptor gene that impairs transportof the receptor tothe plasma membrane and causes insul~nresistant diabetes. EMBO J 8:2509-17,1989 12. Klinkhamer MP, Groen NA, van der Zon GCM, Lindhout D, Sandkuyl LA, Krans HMJ. Moller \F/. Maassen JA A leucine-lo-proline mutatton in the insulin receptor in a family with insulin resistance. EMBO J 8:2503-507. 1989 DIABETES, VOL 39, FEBRUARY 1990 13. Yoshimasa Y, Seino S, Whittaker J, Kakehi A, Kuzuya H, lmura H, Bell GI, Steiner DF: Insulin resistant diabetes due to a point mutationthat prevents insulin receptor processing. Science 240:784-87, 1988 14. Moller DE. Flier JS: Detection of an alteration in the insulin receptor gene in a oatient with insulin resistance. acanthosis nioricans. and the oolvcystik ovary syndrome (type A 'insulin resista&e) N Engl J ' ~ i d 319:1526-29, 1988 15. Odawara M, Kadowaki T. Yamamoto A, Shibasaki Y, Tobe K, Accili D, Bevlns C. Mikami Y, Matsuura N, Akanuma Y, Takaku F, Taylor SI, Kasuga M. human diabetes associated with a mutation in the tyrosine kinase domain of the insulin receptor Science 245:66-68, 1989 16. Taira M, Taira M, Hashimoto N, Sh~madaF, Suzuki Y Kanatsuka A. Nakamura F, Ebina Y, Tatibana M. Makino H, Yoshida S: Human diabetes assoclared wlth a deletion of the tyrosine kinase domain of the insulin receptor. Science 245:63-66, 1989 17. Yang..Feng TL, Franke U, Ullrich A: Gene for human insulin receptor. localization to a site on chromosome 19 involved in pre-8-cell leukemia Science 228:728-31, 1985 18. Shaw DJ, Meredith AL. Brook JD, Sarfarazi M. Harley HG, Huson SM, Bell GI, Harper PS: Linkage relationships of the insulin receptor gene with the cc~mplementcomponent 3, LDL receptor, apolipoprotein C2 and myotonic dystrophy loci on chromosome 19. Hum Genet 74:267-69, 1986 19. Seino S, Seino M. Nishi S. Bell GI Structure of the human insulin receptor gene and characterization of its promoter Proc NaH Acad Sci USA 86:114-18, 1989 20. Muller.Wieland D, Taub R, Tewari DS, Kriauciunas KM, Sethu S, Reddy K, Katin CR. Insulin-receptor gene and its expression in patients with insulin resistance. Diabetes 38:31-38, 1989 21 DeMeyts P, Gu J-L, Shymko RM, Bell G, Whittaker J: Identification of an insulin receptor binding domain complementary to the cooperative site of insulin (Abstract). Diabetolog~a31:484A, 1988 22. Yip CC:. Hsu H, Patel RG, Hawley DM, Maddux BA, Goldine ID: Localization of the insulin bindina site to cysteine rich domain of the a subun~t. Blochem Biophys Res Commun 157.321-29, 1988 23. Kadowaki H, Kadowaki T, Marcus-Samuels B. Cama A, Rovira A, Taylor SI: Site-directed mutagenesis of position 460 in the alpha-subunit of the ~ n s ~ lreceptor in alters cooperative site-site interactions in insulin binding (Abstracl). Diabetes 38 (Suppl. 2) 2A, 1989 24. Seino S Bell GI: Alternative splicing of human insulin receptor messenger RNA. Biochem Biophys Res Commun 159:312-16, 1989 25. Brown Ids, Goldstein JL: A receptor-medlated pathway for cholesterol homeostasis. Science 232:34-47. 1986 26. Russell DW, Esser V, Hobbs HH: Molecular basis of familial hypercholesterolemia. Arteriosclerosis 9 (Suppl. 1). 18-13, 1989 27. Ebina Y, Edery M, Ellis L, Standring D, Beaudoin J, Roth RA Rulter WJ. Express.on of a functional human ~nsulinreceptor from a cloned cDNA in Chinese hamster ovary cells. Proc Natl Acad Sci USA 82:8014-18, 1985 28 Chou CK, Dull TJ. Russell DS, Gherzi R, Lebwohl D. Ullrich A, Rosen OM: Human insulin receptors mutated at the ATP-binding site lack protein tyrosine kinase activity and fail to mediate postreceptor effects of insulin J Biol Chem 262: 1842-47, 1987 DIABETES, VOL. 39, FEBRUARY 1990 29. Whittaker J, Okamoto AK, Thys R, Bell GI, Steiner DF, Hofmann CA Highlevel expression of human insulin receptor cDNA in mouse NlH3T3 cells. Proc Nail Acad Sci USA 84:5239-41. 1987 30 McClain D, Mosthaf L, Ullrich A: Properties of the two naturally occurring alternative forms of the insulin receptor (Abstract). D~abetes38 (Suppl. 2):lA. 1989 31 Taylor SI: Receptor defects in patients wlth extreme insulin resistance Diabetes Metab Rev 1: 171-202, 1985 32. McClain DA, Henry RR, Ullrich A, Olefsky JM: Restriction-fragment-length polymorphism in insulin-receptor gene and insulin reslstance in NIDDM D~abetes37:1071-75, 1988 33. Xiang K-S, Cox NJ, Sanz N. Huang P, Karam JH, Bell GI Insulin-receptor and apolipoprotein genes contribute to development of NIDDM in ~ h i n e s e Americans. Diabetes 38:17-23, 1989 34. Shaw DJ. Bell GI: Rsal polymorphism at the insulin receptor locus (INSR) on chromosome 19. Nucleic Acids Res 13:8661. 1985 35 Elbein SC, Corsetti L, Ullrich A, Permutt MA: Multiple restriction fragment length polymorphisms at the insulin receptor locus: a highly informative marker for linkage analysis. Proc Natl Acad Sci USA 83:5223-27. 1986 36. Sanna MA, Bell GI, Cao A, P~rastuM: Three RFLPsfor the insulin receptor gene INSR: EcoRI, Pst I, Hindlll. Nucleic Acids Res 14:6776, 1986 37. Takeda J, Seino Y, Fukumoto H, Koh G, lmura H, Bell GI. Pvull polymorphic sites in the human insulin receptor gene INSR. Nucleic AcldsRes 14:6777, 1986 38 Patel P, O'Rahilly S, Ullrich A, Turner RC. Wainscoat J S A new Sst 1 RFLP assocated with human insulin receptor locus. Nucle~cAcids Res 16:5700, 1988 39 Cox NJ, Spielman RS. Kahn CR, Muller-Wieland D, Kriauciunas KM, Taub R. Four RFLPs of the human insulin receptor gene: Pstl, Kpnl, Rsal (2 RFLPs). Nucle~cAcids Res 16:8204, 1988 40. Accili D, Elbein S, McKeon C. Taylor SI: A new EcoRl polymorphism for the insulin receptor gene. Nucleic Acids Res 17:821, 1989 41. Sten-Linder M. Stern I, Bell GI. Luthman H: Human insulin receptor RFLPs detected by Hlndlll and Dra I Nucleic Acids Res 17:1277, 1989 42 Elbein SC: Molecular and clinlcal character~zationof an insertional polymorphism of the insulin-receptor gene Diabetes 38:737-43, 1989 43. Haneda M, Polonsky KS, Bergenstal RM, Jaspan JB, Shoelson SE, Blix PM, Chan SJ, Kwok SCM. Wishner WB, Zeidler A. Olefsky JM, Freidenberg G, Tager HS, Steiner DF, Rubenstein AH: Famil~alhyperinsulinemia due to a structurally abnormal insulin: def~n~tion of an emerging new clinical syndrome. N Engi J M e d 310:1288-94, 1984 44. Nanjo K, Sanke T, Miyano M, Okai K, Sowa R. Kondo M, Nishimura S. Iwo K, Miyamura K, Given BD, Chan SJ, Tager HS, Steiner DF, Rubenstein AH: Diabetes due to secretion of a structurally abnormal insulin (insulin Wakayama): clin~caland functional characteristics 01 [LeuA3]insulin.J Ciin Invest 77.514-19, 1986 45. Nanjo K, Miyano M, Kondo M, Sanke T, Nishimura S, Miyamura K, lnouye K, Given BD, Chan SJ, Polonsky KS. Tager HS, Steiner DF, Rubenstein AH. Insulin Wakayama: familial mutant insulin syndrome in Japan. Diabetologfa 30:87-92, 1987 46. Cox NJ, Epstein PA, Spielman RS: Linkage studies of NlDDM and the ~nsulinand insulin-receptor genes Dlabetes 38:653-58, 1989