* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download The Citric acid cycle
Deoxyribozyme wikipedia , lookup
Lipid signaling wikipedia , lookup
Peptide synthesis wikipedia , lookup
Proteolysis wikipedia , lookup
Enzyme inhibitor wikipedia , lookup
Microbial metabolism wikipedia , lookup
Electron transport chain wikipedia , lookup
Mitochondrion wikipedia , lookup
Metalloprotein wikipedia , lookup
Adenosine triphosphate wikipedia , lookup
Catalytic triad wikipedia , lookup
Basal metabolic rate wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Butyric acid wikipedia , lookup
Metabolic network modelling wikipedia , lookup
Biosynthesis wikipedia , lookup
Biochemistry wikipedia , lookup
Oxidative phosphorylation wikipedia , lookup
Glyceroneogenesis wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Lactate dehydrogenase wikipedia , lookup
NADH:ubiquinone oxidoreductase (H+-translocating) wikipedia , lookup
Nicotinamide adenine dinucleotide wikipedia , lookup
Fatty acid synthesis wikipedia , lookup
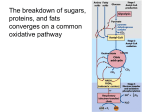
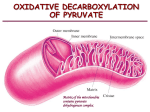
The Citric acid cycle 4/16/2003 The Citric acid cycle It is called the Krebs cycle or the tricarboxylic and is the “hub” of the metabolic system. It accounts for the majority of carbohydrate, fatty acid and amino acid oxidation. It also accounts for a majority of the generation of these compounds and others as well. Amphibolic - acts both catabolically and anabolically 3NAD+ + FAD + GDP + Pi + acetyl-CoA 3NADH + FADH + GTP + CoA + 2CO2 History By 1930 it was established that the addition of lactate, acetate succinate, malate, a-ketoglutaric acid (dicarboxylic acids) and citrate and isocitrate (tricarboxylic acids) when added to muscle mince that they stimulated oxygen consumption and release of CO2 1935Albert Szent-Gyorgyi showed that Succinate Fumarate Malate Oxaloacetate Carl Martius and Franz Knoop showed Citrate succinate cis-aconitate fumarate isocitrate malate a ketoglutarate oxaloacetate Martius and Knoop showed that pyruvate and oxaloacetate could form citrate non-enzymatically by the addition of peroxide under basic conditions. Krebs showed that succinate is formed from fumarate, malate or oxaloacetate. This is interesting since it was shown that the other way worked as well!! Pyruvate can form citrate enzymatically Pyruvate + oxaloacetate citrate + CO2 The interconversion rates of the intermediates was fast enough to support respiration rates. Overview The citric acid cycle enzymes are found in the matrix of the mitochondria Substrates have to flow across the outer and inner parts of the mitochondria Nathan Kaplan and Fritz Lipmann discovered Coenzyme A and Ochoa and Lynen showed that acetylCoA was intermediate from pyruvate to citrate. CoA acts as a carrier of acetyl groups Acetyl-CoA is a “high energy” compound: The DG' for the hydrolysis of its thioester is -31.5 kJ• mol-1 making it greater than the hydrolysis of ATP Pyruvate dehydrogenase converts pyruvate to acetyl-CoA and CO2 Pyruvate dehydrogenase A multienzyme complexes are groups of non covalently associated enzymes that catalyze two or more sequential steps in a metabolic pathway. Molecular weight of 4,600,000 Da E. coli Pyruvate dehydrogenase -- yeast E1 24 60 dihydrolipoyl transacetylase --E2 24 60 dihydrolipoyl dehydrogenase--E3 12 12 24 E2 subunits 24 E1 orange 12 E3 Red a and b together EM based image of the core E2 from yeast pyruvate dh 60 subunits associated as 20 cone-shaped trimers that are verticies of a dodecahedron Why such a complex set of enzymes? 1 Enzymatic reactions rates are limited by diffusion, with shorter distance between subunits a enzyme can almost direct the substrate from one subunit (catalytic site) to another. 2. Channeling metabolic intermediates between successive enzymes minimizes side reactions 3. The reactions of a multienzyme complex can be coordinately controlled Covalent modification of eukaryotic pyruvate dehydrogenase The five reactions of the pyruvate dehydrogenase multi enzyme complex The enzyme requires five coenzymes and five reactions Pyruvate + CoA + NAD+ acetyl-CoA + CO2 + NADH The Coenzymes and prosthetic groups of pyruvate dehydrogenase Cofactor Location Function Thiamine pyrophosphate Bound to E1 Decarboxylates pyruvate Lipoic acid Covalently linked to a Lys on E2 (lipoamide) Accepts hydroxyethyl carbanion from TPP CoenzymeA Substrate for E2 FAD (flavin) Bound to E3 NADH Substrate for E3 Accepts acetyl group from lipoamide reduced by lipoamide reduced by FADH2 Domain structure of dihydrolipoyl transacetylase E2 Pyruvate dehydrogenase 1. Pyruvate dh decarboxylates pyruvate using a TPP cofactor forming hydroxyethyl-TPP. 2 The hydroxyethyl group is transferred to the oxidized lipoamide on E2 to form Acetyl dihydrolipoamide-E2 3 E2 catalyzes the transfer of the acetyl groups to CoA yielding acetyl-CoA and reduced dihydrolipoamide-E2 4 Dihydrolipoyl dh E3 reoxidizes dihydrolipoamide-E2 and itself becomes reduced as FADH2 is formed 5 Reduced E3 is reoxidized by NAD+ to form FAD and NADH The enzymes SH groups are reoxidized by the FAD and the electrons are transferred to NADH HO C C C C HO CH3 C + CH3 O S S S N S N S N R1 H3C R1 H3C R1 H3 C S CH3 S HS + HS E2 E2 O CoA SH CH3 E2 HS S O + HS HS + CoA S Ch3 E2 E2 FAD FAD S SH SH S + + S HS S HS E2 E2 NAD+ FAD SH SH FADH2 NADH + H+ FAD S S S S O CH3 S O HS S E2 O S E2 S S SH E2 S E2 S E2 CH3 S CH3 O S HS E2 SH E2 S S CH3 S E2 Arsenite or organic arsenical compound inhibition O- S HS OH O- As + As OH S HS R R