* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Molecular Cloning
Agarose gel electrophoresis wikipedia , lookup
Comparative genomic hybridization wikipedia , lookup
Maurice Wilkins wikipedia , lookup
List of types of proteins wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Transcriptional regulation wikipedia , lookup
Gene expression wikipedia , lookup
Gel electrophoresis of nucleic acids wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Molecular evolution wikipedia , lookup
Community fingerprinting wikipedia , lookup
Non-coding DNA wikipedia , lookup
Expression vector wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
DNA supercoil wikipedia , lookup
DNA vaccination wikipedia , lookup
Transformation (genetics) wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
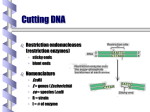
Lecture 13 Molecular Cloning • • • • Recombinant DNA technology depends on the ability to produce large numbers of identical DNA molecules (clones). Clones are typically generated by placing a DNA fragment of interest into a vector DNA molecule, which can replicate in a host cell. When a single vector containing a single DNA fragment is introduced into a host cell, large numbers of this fragment are reproduced along with the vector. Two commonly used vectors for cloning are E. coli plasmid vectors and bacteriophage λ vectors DNA restriction cutting, plasmid, phage, DNA ligation, antibiotic selection, cDNA, cDNA Library, expression vector Ligase reaction: opposite to restriction enzyme, requires 5’-P (from DNA2) and 3’-OH (from DNA1). Ligase mechanism: DNA ligase forms activated ligase-AMP (from ATP or NAD), AMP links to 5’-P, 3’-OH attacks forming new phosphodiester bond, seal! Ligation of the sticky ends: two DNA molecules, cleaved with EcoRI and ligate to form recombinant molecules Formation of cohesive ends Addition and cleavage of a chemically synthesized linker. Producing sticky ends with adaptors. Plasmid and cloning vector Plasmid: circular, ds, extrachromosomal self-replicating DNA molecule occuring naturally in bacteria, yeast and higher eukaryotic cells, either parasitic or symbiotic, ~kb to 100 kb, at least one to the daughter cell (drug-resistance, transfer genes encoding proteins forming macromolecular tube or pilus). Cloning vector: engineered plasmids with reduced size (~3 kb), containing only ori, drugresistance gene, cloning site. Ampr encodes β-lactamase which inactivates ampicillin. Basic cloning cut both of the vector and insert with BamH1, vector treated with alkaline phosphatase (prevent resealing), ligase put insert into the vector, two nicks will be sealed in vivo. Recombinant DNA The plasmid and the foreign DNA are cut by a restriction endonuclease (EcoRI in this example) producing intermediates with sticky and complementary ends. Those two intermediates recombine by basepairing and are linked by the action of DNA ligase. A new plasmid containing the foreign DNA as an insert is obtained. A few mismatches occur, producing an undesirable recombinant. 1973 PNAS 70, 3240 First recombinant DNA produced: Stanley Cohen and Herbert Boyer cut 2 plasmids with EcoRI, ligate and screen for E.coli clones that are resistant to both antibiotics since they harbor the recombinant plasmids. Patent 4,237,224, 1980 http://www.dnai.org/text/mediashowcase/index2.html?id=1186 pBR322 plasmid Host containing pBR322 with insert at different restriction sites can be selected. There are many sites can be used. Cloning at the PstI site Make construct, transform bacteria, screen for tetracycline-resistant but ampicillin-sensitive clones using replica plating (make a replica plate treating with ampicillin, sensitive clones are recovered from the original plate). Polylinkers Chemically synthesized polylinker containing one copy of several different restriction sites is introduced into the vector to facilitate directional cloning. MCS: multiple cloning site MCS is inserted into a gene encoding the N-terminal part of β-galactosidase. Clones harboring the vector plus an insert remain white. pUC series: derived from pBR322 (40% deleted), the MCS is in the lacZ’ (Nterminal of β-galactosidase, the host contains the C-terminus portion, MCS starts after the 7th codon, retains the ORF), so no insert blue (with X-gal), with insert white. Molecular cloning The general procedure for cloning with plasmid vectors: plasmid with Ampr and insert is transformed into host with CaCl2 or other methods, grow in ampicillin plate, surviving cells form colony. Plasmid cloning permits isolation of DNA fragments from complex mixtures: Each colony is derived from a single cell containing the same plasmid. Cloning strategy Process by which a plasmid is used to import recombinant DNA into a host cell for cloning. In DNA cloning, a DNA fragment that contains a gene of interest is inserted into a cloning vector or plasmid. The plasmid carrying genes for antibiotic resistance, and a DNA strand, which contains the gene of interest, are both cut with the same restriction endonuclease. The plasmid is opened up and the gene is freed from its parent DNA strand. They have complementary "sticky ends." The opened plasmid and the freed gene are mixed with DNA ligase, which reforms the two pieces as recombinant DNA. This recombinant DNA stew is allowed to transform a bacterial culture, which is then exposed to antibiotics. All the cells except those which have been encoded by the plasmid DNA recombinant are killed, leaving a cell culture containing the desired recombinant DNA. DNA cloning allows a copy of any specific part of a DNA (or RNA) sequence to be selected among many others and produced in an unlimited amount. This technique is the first stage of most of the genetic engineering experiments: production of DNA libraries, PCR, DNA sequencing, et al. Molecular cloning Molecular cloning DNA Libraries in λ phage and other cloning vectors • Cloning all of the genomic DNA of higher organisms into plasmid vectors is not practical due to the relatively low transformation efficiency of E. coli and the small number of transformed colonies that can be grown on a typical culture plate • Cloning vectors derived from bacteriophage do not suffer from such limitations • A collection of clones that includes all the DNA sequences of a given species is called a genomic library • A genomic library can be screened for clones containing a sequence of interest (i) EM of bacteriophage λ virion. (ii) Simplified view of bacteriophage genome:about 60 genes, replaceable region not essential and can be replaced by foreign DNA up to 25 kb. Large segment of the 48kb DNA of the λ phage are not essential for productive infection and can be replaced with inserts. (iii)Assembly of bacteriophage λ virion: during late stage of λ infection concatomers are formed, Nu 1 and A protein push 1 copy of λ from concatomer into the head, then tail added and they are ready to go to the war. cloning λ phage vector Charon phage 4 takes 20 kb, remove the stuffer, ligate the insert, mix with the in vitro packaging extract, infect cells, the insert has to be between 12 kb and 20 kb. Alternative infection modes for λ phage: lytic or lysogenic pathway A set of lambda (or plasmid) clones that collectively contain every DNA sequence in the genome of a particular organism. Nearly complete genomic libraries of higher organisms can be prepared by lambda cloning. Genomic Library (i) A set of lambda (or plasmid) clones that collectively contain every DNA sequence in the genome of a particular organism. Nearly complete genomic libraries of higher organisms can be prepared by lambda cloning. (ii) Construction of a genomic library: Sau3A and BamH1 generate complementary sticky ends. (iii) Plaque hybridization: selection of positive clones, filter replicates the dish, plaque containing phage DNA denatured by base, block the filter with nonspecific DNA or protein, hybridize with specific gene probe, autoradiography to select the target .gene. (iv) The most common approach to identifying a specific clone involves screening a library by hybridization with radioactively labeled DNA or RNA probes. Identification of a specific clone from a l phage library by membrane hybridization. Identifying, analyzing, and sequencing cloned DNA Cosmid vector Larger DNA fragments up to 45 kb can be cloned in cosmids and other vectors, cosmid has cohesive ends for packaging and plasmid ori for replication as plasmids in bacteria. cosmids cannot replicate as phages but they are still infectious. M13 Phage M13 Phage: a filamentous virus 900 nm long and 9 nm wide, single strand 6.4kb circle DNA protected by 2710 identical proteins, gets into E. coli via sex pilus, replicate to RF, only (+) is packed into virus particle. Useful for sequencing. Complementary DNA DNA (cDNA) libraries are prepared from isolated mRNAs Complementary oligo-dT column to purify mRNA Complementary DNA (cDNA) libraries Formation of cDNA duplex: RT, alkali digestion, oligo(dG) tailing, oligo(dC) primer Preparation of a cDNA library Making a cDNA Library Making a cDNA Library: Reverse Transcriptase, RNase H (degrade RNA in RNA-DNA hybrid) , DNA polymerase I (using RNA as primer and perform nick translation), Terminal deoxynucleotidyl transferase (adding dCTP to the cDNA duplex, and dGTP to the vector), Ligase and pol I in the host perform ligation, RNA removal, and sealing. Use RT-PCR to clone a single cDNA if the sequence of mRNA is known. Phagemid pBS or pBluescript, MCS inserted into lacZ’, ori of the ss phage f1, T3 and T7 phage RNA polymerase promoters. Expression vector The expression vector contains a fragment of the E. coli chromosome containing the lac promoter and the neighboring lacZ gene. In the presence of the lactose analog IPTG, RNA polnormally transcribes the lacZ gene, producing lacZ mRNA, which is translated to the encoded protein, β-galactosidase. (i) Forming fusion protein in plasmid vector: pUC and pBS vectors place inserted DNA under the control of the lac promoter. If the DNA is in frame with the lacZ’ gene, a fusion protein will be expressed. E. coli expression systems can produce full-length proteins: Producing high levels of proteins from cloned cDNAs. Many proteins are normally expressed at very low concentrations within cells, which makes isolation of sufficient amounts for analysis difficult. To overcome this problem, DNA expression vectors can be used to produce large amounts of full length proteins. Lac promoter. The lacZ gene can be cut out of the expression vector with restriction enzymes and replaced by the G-CSF cDNA. After transformation, IPTG will induce the expression of G-CSF protein. Two-step system : T7 RNA polymerase and T7 late promoter Even larger amounts of a desired protein can be expressed with a two-step system, 10-70% of the total protein synthesized by these cells after IPTG is the protein of interest. Forming fusion protein in phage vector: λg11with lac control region followed by lacZ gene, cloning sites are located within the lacZ gene, direct detection of protein with antibody. Degenerate probe: screen for the correct cDNA or genomic clone from the library.