* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download The International Tomato Sequencing Project and Related
Transposable element wikipedia , lookup
Molecular Inversion Probe wikipedia , lookup
Pharmacogenomics wikipedia , lookup
Copy-number variation wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Segmental Duplication on the Human Y Chromosome wikipedia , lookup
Neocentromere wikipedia , lookup
Human genetic variation wikipedia , lookup
Designer baby wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Genetic engineering wikipedia , lookup
Non-coding DNA wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
Minimal genome wikipedia , lookup
DNA sequencing wikipedia , lookup
Microevolution wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Human genome wikipedia , lookup
Genome (book) wikipedia , lookup
History of genetic engineering wikipedia , lookup
Public health genomics wikipedia , lookup
Pathogenomics wikipedia , lookup
Genome editing wikipedia , lookup
Genome evolution wikipedia , lookup
Metagenomics wikipedia , lookup
Exome sequencing wikipedia , lookup
Whole genome sequencing wikipedia , lookup
Human Genome Project wikipedia , lookup
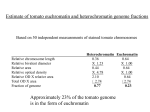
The US Contribution to the International Tomato Genome Sequencing Project Overview of Presentation Background on the International Solanaceae Initiative (SOL) and the International Tomato Genome Sequencing Project Sequencing strategy Resources available SOL Genomics Network (sgn.cornell.edu) Details about resources Informatics pipelines Educational outreach activities An International Workshop to Discuss Sequencing of the Tomato Genome: Feasibility, Benefits and Strategy November 3, 2003, Washington D.C * funded in part by the National Science Foundation On November 3, 2003 an international meeting was held in Washington DC which was attended by 70 scientists from 11 countries. The outcome was the creation of a 10 year vision for research in the family Solanaceae referred to as “ The International Solanaceae Genome Project or SOL”. SOL, which includes sequencing the tomato genome, will create a worldwide research and informational infrastructure in which a systems biology approach can be taken to address key questions in biology and agriculture for which the Solanaceae are ideally suited For details, see: http://sgn.cornell.edu/solanaceae-project/ The SOL Vision Potato Eggplant Petunia Coffee* Pepper Tomato reference genome sequence Understanding Diversification & Adaptation Nicotiana Arabidopsis and other genomes Exploring the Role of Natural Diversity in the Genetic Improvement of Crops * Coffee is closely related to the Solanaceae and has a similar genome size and chromosome karyotype -- a comparative map of coffee with solanaceous species is part of the SOL project Objectives of Tomato Sequencing Project Produce a contiguous sequence of the gene rich, euchromatic arms of each of the 12 tomato chromosomes. Groups from 10 countries are partners in the project Our group is sequencing 3 of the chromosomes, the remaining 9 are each being sequenced by a group in a different country. Process and annotate this sequence in a manner consistent and compatible with similar data from Arabidopsis, rice and other plant species. Create an international bioinformatics portal for comparative Solanaceae genomics which can store, process, and make available to the public the sequence data and derived information from this project and associated genomics activities in other solanaceous plants. Tomato Euchromatin Gene Space Sequencing Strategy The tomato genome contains approximately 950 Mb of DNA of which 23% is euchromatin. Peterson et al., 1996, Genome 39:77-82 The majority of tomato genes reside in the euchromatin. Gene rich and repeat poor Approximately 85% of the tomato genes supported by available BAC (Bacterial Artificial Chromosome) sequence data available from BACs isolated on the basis of target genes Organization of tomato genome & impact on sequencing strategy telomere euchromatin 162 bp subtelomeric repeat centromere A telomere structure pericentric heterochromatin euchromatin pericentric pericentric heterochromatin heterochromatin BAC hybridization in euchromatin C 7 bp telomeric repeat B BAC hybridization US Project Initiated in September 2004 Chromosomes 1, 10, 11 Funding from NSF Plant Genome Research Program DNA sequencing is sub-contracted to a high-capacity sequencer Distribution of materials to sequencing partners Coordination of international efforts Bioinformatics portal SOL Genomics Network (SGN) sgn.cornell.edu Jim Giovannoni PI, BTI • Overall operation of project • Interactions among co PIs • Generation of BAC libraries • Clone distribution to international project members • Clone handling & storage • Computational analysis of regulatory domains Steve Tanksley Co-PI, Cornell • Selection of seed BACs and extension BACs for sequencing. • Overgo anchoring of genetic markers. • Genetic mapping of BACs • Comparative mapping Lukas Mueller Co-PI, Cornell • Bioinformatics • Interaction with sequencing center • BAC assembly • Annotation • Data integration with other countries • Training Stephen Stack Co-PI, CSU • Distal/proximal BAC anchoring • FISH for gap estimates • Heterochromatin BAC identification of sequencing • International coordination for in situ research Joyce Van Eck Co-PI, BTI • Day to day coordination/ operations of project. • Planning and running teleconferencing of co PIs. • Assist in preparing annual reports and conference presentations. • Educational outreach activities Outline of Approach Sequencing is following a BAC-by-BAC strategy. Starting point for sequencing is approximately 1000 "seed” BACs individually anchored to a high density genetic map. Each sequenced anchor BAC serves as a seed from which to radiate out into the minimum tiling path. Especially interested in BACs located as close as possible to telomeres and euchromatin/heterochromatin borders. Fluorescence In Situ Hybridization (FISH) is being utilized for BAC localization. To steer sequencing activities into the euchromatin and away from the heterochromatin Resources Available High density genetic map Physical map Accounts for 20% of the genome sequence Fingerprint Contigs (FPC) Developed from genetic markers Integrate the genetic with the physical map Seed BACs (Bacterial Artificial Chromosomes) BAC libraries and corresponding hybridization filters BAC end sequences (~ 400,000) various types of molecular markers Overgo probes (overlapping oligonucleotide probes) Solanum lycopersicum x S. pennellii F2 population (Tanksley et al. 1992, 132:1141-1160) Assemble the BAC collection into contigs Rod Wing and Wellcome Trust Sanger Institute FISH (Fluorescence In Situ Hybridization) Future Resource Fosmid Library Use for filling small gap intervals Made from sheared genomic DNA Average insert size of 40 kb (~12x physical coverage) End-sequence 400,000 clones Selection and Verification of Seed BACs Selection Choose two seed BACs (>100kb) that are well within the euchromatic region Only one needs to be confirmed to move ahead Verification (at least one method should be chosen) Verify marker-BAC association by sequencing with marker-specific primers Rehybridizing BAC clones using overgo probes PCR amplification of genetic markers from the BAC clones Methods To Verify Locations of Seed BACs Map BACs in tomato Introgression Lines (ILs) CAPS markers Fluorescence In Situ Hybridization (FISH) Steve Stack’s lab, Colorado State University US and countries not set up to do FISH Countries doing FISH China The Netherlands France sent a participant to Stack lab to learn FISH. FISH Image BAC Libraries DNA from Heinz 1706 Library Total # name/enzyme of clones Cloning vector Average insert size (kb) # of BAC end sequences HindIII 129,024 pBeloBAC11 117 188,130 MboI 50,688 pEC BAC1 135 112,507 EcoRI 75,000 pIndigoBAC-5 95 - 100 101,375 Seed BACs for each chromosome are distributed to each respective country sequencing that chromosome euchromatin euchromatin Pachytene chromosome FISH seed BAC Genetic markers anchored via OverGo hybridizatrions Seed BACs (solid) anchored to genetic map and pachytene chromosomes via FISH; bridging clones (dashed) in MTP identified through combination of BAC end sequence database and FPC Genetic map Informatics Pipelines SOL Genomics Network (sgn.cornell.edu) BAC registry database Project members can log in to upload BAC information. Project-wide Sequencing Quality Control (QC) Implemented various global QC checks Functional and Structural Annotation The International Tomato Annotation Group (ITAG) Formed at a meeting in Ghent, Belgium (October, 2006) Established an annotation protocol for the tomato genome. Summary of Tomato Genome Annotation Pipeline Repeat Content in Annotated BACs on Chromosome 1 GBrowse on SGN Euchromatic BAC Heterochromatic BAC Tomato FISH Map on SGN -Represents FISH analyses done at several labs involved in the project Indicates euchromatin Indicates heterochromatin; this darker blue at the ends represents the telomeres FISH localized BACs Tomato FISH Map Outreach Bioinformatics Summer Internship SOL Genomics Network Undergraduates and high school students Each student has her/his own project Housing provided The Solanaceae Family goes to School Geared towards kindergarten - 5th grade Elementary schools Afterschool programs Others Presentations to various groups High school teacher workshop 2005 and 2006 Bioinformatics Summer Interns The Solanaceae Family goes to School SOL Newsletter -bimonthly -sent by e-mail -list of ~400 members worldwide -also posted as pdf on SGN (sgn.cornell.edu) -Send e-mail to [email protected] to be added to list Acknowledgements Boyce Thompson Institute Colorado State University Julia Vrebalov Ruth White Lorinda Anderson Suzanne Rogers Song-Bin Chang Cornell University Yimin Xu Nancy Eanetta Rob Buels Beth Skwarecki Marty Kreuter Naama Menda John Binns Chenwei Lin SeqWright Agencourt Bioscience SymBio Funding NSF Plant Genome Research Program