* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Deletion Map of Chromosome 9 and p16 (CDKN2A) Gene Alterations
Epigenetics in stem-cell differentiation wikipedia , lookup
History of genetic engineering wikipedia , lookup
Gene desert wikipedia , lookup
Gene nomenclature wikipedia , lookup
Saethre–Chotzen syndrome wikipedia , lookup
Neuronal ceroid lipofuscinosis wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Gene expression profiling wikipedia , lookup
Y chromosome wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Gene therapy wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Gene expression programming wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Point mutation wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Mir-92 microRNA precursor family wikipedia , lookup
Skewed X-inactivation wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Microevolution wikipedia , lookup
Neocentromere wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Designer baby wikipedia , lookup
Genome (book) wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Oncogenomics wikipedia , lookup
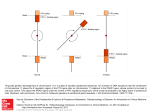
ICANCER RESEARCH57. 907-912. March I. 19971 Deletion Map of Chromosome 9 and p16 (CDKN2A) Gene Alterations in Neuroblastoma1 Junko Takita, Yasuhide Hayashi, Takashi Kohno, Naohito Yamaguchi, Ryoji Hanada, Keiko Yamamoto, and Jun Yokota2 Biology Division If. T., T. K., J. Y.J and fancer Information and Epidemiology Division fN. Y.J, National Cancer Center Research Institute, 1.1. Tsukifi 5-chome, Chuo-ku, Tokyo 104: Department of Pediatrics. Faculty of Medicine. Unis'ersitv of Tokyo. 3-I, Hongo 7-chome. Bunkyo-ku, Tokyo I 13 Ii. T., Y. HI; and Division of Hematology/Oncology. Saitama Children ‘s Medical Center, 2100, Magome. Iwatsuki 339 (R. H., K. Y.J. Japan ABSTRACT We reported previously that loss of heterozygosity (LOH) on chromo somes 2q, 9p and 18q frequently occurs in neuroblastoma and that patients with 9p LOH in the tumors showed statistically significant asso ciation with an advanced stage of the disease and poor prognosis. To determine the role of chromosome 9 loss in neuroblastoma, we performed deletion mapping of chromosome 9 in 80 cases of neuroblastoma using 11 polymorphic microsatellite markers and a restriction fragment length porymorphism marker. LOll at one or more loci on chromosome 9 was detected in 33 ofSO cases (41%). Chromosome 9p was lost in 24 of8O cases (32%), whereas chromosome 9q was lost in 18 of 80 cases (23%). There were two commonly deleted regions mapped to 9p2l between the D9S171 marker and the IFNBJ marker and 9q34—qterdistal to the D9S176 marker. In addition, patients with LOH at 9p21 but not at 9q34—qterin the tumors showed statistically significant association with poor prognosis (P = 0.023). Because the commonly deleted regions at 9p21 includes the p16 (CDKN2A) gene, the status of thepl6 gene was further examined in 80 fresh tumors and 19 cell lines of neuroblastoma. A missense mutation was detected at codon 52 in a fresh tumor. ThepI6 gene was not expressed in 13 of 19 cell lines (72%), and 5 of the 13 cell lines displayed methylation ofthe CpG island surrounding the first exon of thepl6 gene. These results suggest that the p16 gene is a candidate tumor suppressor gene for neuroblastoma, and its inactivation may contribute to the progression of neuroblastoma. INTRODUCTION Recent molecular studies have revealed that the genesis and pro gression of human cancer is largely attributed to accumulation of a series of genetic events that culminate in the transformation of a cell into a malignant clone (1). Central to this theory are the roles of oncogenes and tumor suppressor genes, the activation and inactivation of which, respectively, cause disruption of critical events in cell division and differentiation (1). The paradigm of alterations in the tumor suppressor gene is a mutation of one allele and a loss of the other allele. Reduction to homozygosity of the tumor suppressor gene can be detected as LOH3 of informative polymorphic markers in the region of the tumor suppressor gene. Thus, allelic losses are hallmarks of chromosomal regions harboring tumor suppressor genes (2). Although NB is one of the most common childhood tumors, little is known about the genetic changes that contribute to the development of tumor. It has been reported that LOH occurs frequently on at least three chromosome arms, ip, 1lq, and 14q, in NB (3—10).In addition, we demonstrated recently that three additional loci on chromosomes MATERIALS as stage whom requests for reprints 1 year. Patients or surgery 3 The abbreviations used are: IVS. In our 80 NB patients, 74 multidrug with stage I, II, or IVS were treated radiotherapy. chemotherapy consisting Patients consisting with either of vincristine surgery alone and cyclophosphamide with stage III or IV were treated of cyclophosphamide, Adriamycin, with cispla tin, and etoposide with or without surgery and radiotherapy. In 53 cases, histological data was available; thus, tumors were histologically classified as described by Ota and Shimizu (19). There were 4 cases of GNB classified as well differentiated, 8 cases of GNB classified as composite, 9 cases of GNB classified as poorly differentiated, 27 cases of NB classified as rosette-fibril lary, and 5 cases of NB classified as round cell. We also used 19 NB cell lines, should LOH, IV, and 8 as stage plus chemotherapy with or without NB1, NB9, NB16, NB19, NB39, NB69, LAN1, LAN2, LAN5, KP-N-NS, GOTO, CHP-134, IMR-32, TNB-l, TGW, SCMCN2, SCMCN3, SCMCN4, be addressed. Phone: 81-3-3542-251 and SCMCN5, for the analysis of the pitS gene alterations in NB. DNA, RNA, and Protein Extraction. DNA was isolated from tumors, normal tissues, and cell lines by proteinase K digestion and phenol/chloroform isoamyl alcohol (24:1) extraction as described previously (20). mRNA was 1, Ext. extracted from cells growing in culture using the FastTrack 2.0 mRNA 4650: Fax: 8 1-3-3542-0807. toma; NB, neuroblastoma; III, 16 as stage patients were infants under I year of age at diagnosis, and 6 patients were over The costs of publication of this article were defrayed in part by the payment of page charges. This article must therefore be hereby marked advertisement in accordance with 18 U.S.C. Section 1734 solely to indicate this fact. This work was supported in part by a Grant-in-Aid from the Ministry of Health and Welfare for the second term Comprehensive 10-Year Strategy for Cancer Control and Grants-in-Aid from the Ministry of Health and Welfare and from the Ministry of Education, Science, Sports and Culture of Japan. 2 To AND METHODS Primary Tumors and Cell Lines. Tumor samples were randomly obtained from 80 patients admitted to various institutions between May 1987 and July I993 at surgery or at autopsy. Corresponding normal tissues were available in all cases. The patients were staged according to the classification of staging in NB (18). Of the 80 cases, 21 were classified as being stage I, 27 as stage II, 8 Received 8/5/96; accepted 1/6/97. @ 2q, 9p, and l8q were deleted with high frequency in NB. Moreover, several studies have shown the correlation of genetic changes with prognosis of the patients with NB (1 1). N-myc (NMYC) oncogene amplification has been known to be an appreciable prognostic factor in an advanced stage of the disease. It is also indicated that chromo some Ip deletion frequently occurs in an advanced stage of the disease, and there may be two tumor suppressor genes on chromo some Ip associated with progression of the disease (9, 10). In addi tion, we also reported that 9p LOH was significantly associated with advanced stages of the disease and with poor prognosis (12). To determine the locus on chromosome 9 that may harbor putative tumor suppressor genes involved in the progression of NB, we per formed deletion mapping of chromosome 9 in 80 cases of NB using 11 microsatellite markers and a RFLP marker. The result indicated that there were two commonly deleted regions on chromosome 9, 9p2l and 9q34—qter, in NB, and that LOH at 9p2l was significantly associated with poor prognosis. The p16 (CDKN2A) tumor suppressor gene has been mapped to 9p2l and is inactivated in a variety of malignancies by various mechanisms, including deletion, point mu tation, and methylation of the CpG island in the 5' end of the p16 gene (13—17).Therefore, we further examined the alterations of the pitS gene in NB. Although deletions and mutations of the pi6 gene are infrequent, transcriptional silencing and DNA methylation were fre quently detected in NB cell lines. Thus, it was indicated that the p/tS gene is a candidate tumor suppressor gene involved in the progression of NB. loss of heterozygosity; SSCP, single-strand conformational GNB, isolation kit according to the manufacturer's ganglioneuroblas polymorphism. protein was extracted by lysing instructions (lnvitrogen). Cellular 1 X 106 cells with 40 p1 of lysis buffer 907 Downloaded from cancerres.aacrjournals.org on August 9, 2017. © 1997 American Association for Cancer Research. [50 mM CHROMOSOME9 DELETIONSAND p16 IN NEUROBLASTOMA HEPES-NaOH (pH 7.0), 1% NP4O, 1% sodium deoxycholate, 0. 1% SDS, 250 mM NaCl, 5 mM EDTA, 50 msi NaF, 1 mi@iDTT, 1 mist phenylmethylsulfonyl fluoride, and 50 @g/ml aprotinini. Sequencing. PCR productsselected for sequencingwere ligated with the pCRIITA-cloning vector (Invitrogen) and transformed into competent DH5a cells (Life Technologies, Inc.). Both strands were sequenced for each PCR PCR-LOHAnalysis.DNAfromtumorsandcorresponding normaltissues product from at least three independent clones. were analyzed for LOH by PCR amplification of microsatellite sequences. Microsatellite reaction markers for PCR-LOH volumes were 10 analysis were listed in Fig. I . Total @l containing 50—100 ng DNA, 10 msi Tris-HCI (pH 8.3), 50 mM KCI, 1.5 mM MgC12,250 @.LM of each deoxynucleotide triphos phate, 0.01% gelatin, 125 ng of each primer, 1.14 @sCi of [a-32P)dCTP,and 1 40%, as described unit of Taq trisomy, besides LOH by this definition. Therefore, it is possible that certain DNA polymerase (Pharmacia Biotech, Inc.). Because unequal previously (23). We could score allelic imbalance, such as amplification of alleles occurs with 35 cycles of PCR (21), PCR amplifications were performed for 35 cycles consisting of denaturation at 94°Cfor 40 s, chromosome annealing and clinical features of the disease among the patient group was examined by Fisher' s exact test. The vital status of the patients was observed until December system at 55°C for 40 s, and extension 9600 (Perkin-Elmer) as described at 72°C for 90 s in a Gene Amp PCR (12). Southern and Northern Blot Analyses. Approximately10 @.tg of purified DNA were digested with appropriate restriction enzymes and separated by electrophoresis on 0.8% agarose gel. DNA was transferred from the gel to nylon membranes ( 12). The membranes were hybridized with a PCR-generated fragment corresponding to exon 1 of the p16 gene (p16-i), a full-length pitS eDNA fragment, JNFBi, and pNB-l labeled with [a-32P]dCTP. LOH at the IFNB1 locus was examined by Southern blot analysis l.Lgof mRNA was denatured with 40% formamide/32% (12). Approximately loci scoring Statistical Analyses. 3 1, 1995. The survival LOH are trisomic rather than monosomic. Significance of the differences in various biological curves for each group of the patients were estimated by the Kaplan-Meier method, and the resulting curves were compared using the log-rank test for univariate analysis. Multivariate analysis was performed using the Cox proportional hazards model. Western Blot Analysis. Fifty jsg of protein were separatedin a 4—20% gradient SDS/polyacrylamide gel and electroblotted to Hybond-Enhanced Chemiluminescence 3 lington formaldehyde and was Heights, (ECL) nitrocellulose membrane IL). After being blocked (Amersham with 5% nonfat Corp., Ar dry milk and 0.1% electrophoresed on a 1.2% agarose gel containing 25% formaldehyde. Then mRNA was transferred to nylon filters. The filters were hybridized with p16-I and full-length eDNA probes labeled with [a-32P] dCTP and were exposed to Kodak XAR-5 film at —80°C. Prehybridization, hybridization, and posthy Tween 20 in Tris-buffered saline, membranes were incubated at 37°Cfor 2 h bridization Biotechnology). The blot was subsequently probed by the ECL Western washes were performed basically as described (20). PCR-SSCPAnalysis.All sampleswerescreenedformutationsinexons1 with the 1:400 dilution amino acids blotting 137—156 at the COOH detection confirmed analysis. RESULTS The primer sets for the pi6 gene were: exon 1, PQIS, 5'-TCTGCG were separated by electrophoresis on 6% polyacrylamide gel with 5% glycerol and without glycerol. Moreover, to better assure detection of mutations by PCR-SSCP, we used two cases of lung carcinomas with p16 mutations as positive controls. 9 4 9 16 18 23 24 28 of a rabbit polyclonal anti-p16 antibody (PharMingen), the epitope of which is unknown, and a rabbit polyclonal anti-p16 antibody for and 2 of the pi6 gene by PCR-SSCP analysis (22). Exon 1 was amplified as one fragment, whereas exon 2 was split into two fragments for PCR-SSCP GAGAGGGGGAGAGCAGGCA (sense) and PQ1A, 5'-TCTGCGGAGA GAGGGGGAGAGCAGGCA (antisense); first exon 2, PQ2AS, 5'ACAAGCTFCCTITFCCGTCATGCCG (sense) and PQ2AA, 5'-CCAG GCATCGCGCACGTCCA (antisense); and second exon 2, PQ2BS, 5'-TFC CTGGACACGCTGGTGGT (sense) and PQ2BA, 5'-TCTGAGClTFGGA AGCTCTCAG (antisense). PCR conditions for exon I of the p16 gene were 35 cycles of 50 s at 94°C, 50 5at 65°C, and 50 s at 72°C. PCR conditions for exon 2 of the pi6 gene were 35 cycles of 50 5 at 94°C,50 s at 64°C,and 50 s at 72°C. The PCR products system by staining terminus (Amersham the membrane Corp.). of the p16 protein Equal loading (Santa of protein Cruz was after detection. Frequency and Common Regions of LOH on Chromosome 9 in NB. Eighty cases of NB were examined for LOH on chromosome 9 using 11 microsatellite polymorphic markers and a RFLP DNA marker. The incidence of LOH at each locus is summarized in Fig. 1. All cases showed heterozygous genotypes in their normal tissue at one or more loci on chromosome 9, and LOH at one or more loci was detected in 33 of 80 cases (41%). LOH on chromosome 9p was detected in 24 of 80 cases (30%), whereas LOH on chromosome 9q was detected in 18 of 80 cases (23%). Five of 33 cases (15%) showed LOH at all informative loci, whereas the other 28 tumors showed partial deletions of chromosome 9 (Fig. I). Case 23 showed LOH at 29 30 39 42 60 61 79 80 8 15 33 34 58 59 70 76 7722 40 67 75 27 31 373872 I— D9S281 2135(6%) 24 23 22 L 21 21 @\D9S162 6/41(15%) 13 I\\\FNBI9/27(33%) 12 k\tNA@ 6/41(20%) 11 11 13 21.1 212 21.3 22.1 222 22.3 A I I I I \@D9s@5 2135(9%) \\ 12 @ Definition of LOH. The signal intensity of the polymorphic alleles was quantified and calculated by the scanning densitometer and data analysis system (The Discovery Series; Quantity One, pdi, NY). LOH was considered to be present if reduction rates of signal intensities in tumors were more than I I S1712137(8%) 09S186 20 1/32(9%) l—D9S152 4/37(11%) I 27 D9S195 1127(4%) V26 D9S176 9/44(20%) /34 D9S158 8/339(21%) 31 32 33 34.1 342 34.3 Fig. I . Schematic representation of deletion map of chromosome 9 in NB. The approximate locations of markers used are shown on the left and tumor numbers are shown above. ., LOH; 0, heterozygosity; and no symbol, not informative. cM distances between the markers are indicated in italics. and the number of LOH/informative cases is shown on the right of each locus name. 908 Downloaded from cancerres.aacrjournals.org on August 9, 2017. © 1997 American Association for Cancer Research. CHROMOSOME 9 DELETIONS AND p16 IN NEUROBLASTOMA the D9Si62, IFNB1, and IFNA loci, but heterozygosity was retained at the D9Si 7i locus (Fig. 2). In case 60, LOH was detected at the D9Si7i locus, but heterozygosity was retained at all loci distal to the IFNB1 locus (Fig. 2). Cases 33, 75, and 76 showed LOH at D9S/58, but heterozygosity was retained at all informative loci proximal to the D9Si76 locus (Fig. 2). The result from these five patients implicates the presence of two commonly deleted regions that are mapped between the IFNBi and D9S17J loci at chromosome 9p2l and distal to the D9S176 locus at chromosome 9q34—qter (Fig. 1). The size of genetic distance of a commonly deleted region on chromosome 9p2l is 5 cM and that on 9q34—qter is more than 34 cM. 23603376TNTNTNTN TL D9S1 62 D9SI 62 D9S1 52 D9S1 52 a IFNBI IFNB1 D9S195 D9S195 IFNA INFA D9S176 D9S176 I D9S1 71 D9S1 71 D9S1 58 D9S1 58 Fig. 2. LOH in cases showing partial or interstitial deletions of chromosomes 9. DNA was isolated from tumors (Lanes 7) and corresponding normal tissues (Lanes N) from patient 23 (Lane I), patient 60 (Lane 2), patient 33 (Lane 3), and patient 76 (Lane 4). Allelic fragments that showed LOH are indicated by arrowheads. Table I Correlation Relationship between LOH on Chromosome 9 and Clinicopath ological Findings of NB. Because age, stage, and N-myc amplifica tion are known to be associated with prognosis of patients with NB, the relationship between LOH on chromosome 9 and these clinico pathological findings was examined. There were 56 patients with stage I+II+IVS, 8 patients with stage III, and 16 patients with stage Iv. N-myc amplificationwas detected in 10 of 80 cases. Because 62 of 80 (78%) were found by a mass screening program, the ratio of infantile and early-stage patients in this study was higher than that in previous studies (5, 10). However, age, stage, and genotype of N-myc were also significantly associated with short survival of the patients in our 80 cases (P < 0.001). Therefore, clinical outcome of patients in this study was similar to that of patients in previous studies (5, 10). All patients were followed-up for at least 36 months and up to 82 months. The mean follow-up periods of patients with LOH on chromosomes 9, 9p, and 9q were 44, 46, and 47 months, respectively. The medium ranges of survival of the patients with LOH on chromosomes 9, 9p, and 9q were 50, 42, and 57 months, respectively. LOH on chromo some 9p was significantly associated with the stage of the disease (P = 0.029) and the survival of the patients (P 0.023). However, neither LOH on chromosome 9q nor LOH on chromosome 9 showed association with any of these parameters (Table 1). Ten of 24 patients with 9p LOH died within 24 months, whereas only 9 of 56 patients without 9p LOH died within this period (Fig. 3). However, significant association was not observed between 9q LOH and survival of the patients (P = 0.551; Table 2). Accordingly, LOH on chromosome 9 was not significantly associated with survival (P —0.387; Table 2). There was no correlation between 9p LOH and other clinicopatho logical findings, such as age, course of diagnosis (found by mass screening or clinical symptoms), histological types, and genotype of N-myc and lp LOH. When multivariate analysis of survival was performed by age (under 1 year versus over 1 year) and genotype of N-myc amplification (present or absent) as covariates, no significant of LOH on chromsome 9 with biological and clinical variables in neurblastoma LOH 99p21—229q34—qterAge< I yr (0.566)Stage―I, (0.478)Result of screening+ 3/18(0.871)N-myc amplification+ (0.438)Histological typeGNB 1 yr29/74 II, IVS III IV20/55 4/6 (0.293)―22/74 3/9 10/16 (0.185)12/55 —28/62 5/18(0.565)18/62 —5/10 28/70 (0.349)4/1 Well differentiated Composite Poorly differentiated NB Rosette fibrillary Round cell3/4 3/8 4/9 12/27 2/5 (0.793) 0.3872/4 Survivalb a Fisher's exact b P-value, log-rank 2/6 (0.339)14/74 2/6 3/9 9/16 (0.029)13/55 1/9 2/16 6/18(0.807)13/62 0 20/70 (0.580)1/10 1/8 4/9 7/27 3/5 (0.584) 0.0232/4 15/70 3/8 3/9 6/27 0/5 (0.412) 0.551 test. test. 909 Downloaded from cancerres.aacrjournals.org on August 9, 2017. © 1997 American Association for Cancer Research. CHROMOSOME9 DELETIONSAND p16 IN NEUROBLASTOMA mRNA was not detected in 13 of 19 cell lines (72%). The p16 protein was undetectable by Western blot analysis in the corresponding 13 cell lines (Fig. 5). % Survival 100 Methylation of the p16 Gene in NB Cell Lines. We next exam 90 80 70 60@ 50@ 40@ 9p LOH-(n=56) 30@ 20@ 10@ 0 —.------ 9p LOH+ (n=24) med whether the pi6 gene was transcriptionally silenced in NB cell lines by methylation of a CpG island in the pitS gene. DNA was digested with EC0RI and SinaI and analyzed by Southern blot hybrid ization. Unmethylated fragments of 650 and 350 bp were not detected in five cell lines that did not express piO mRNA, indicating that the critical SmaI site within exon I of all alleles was methylated in these cell lines. However, this site was not methylated in eight other cell lines in which pitS mRNA was not detected. The remaining six cell lines that express pi6 mRNA showed unmethylated 650- and 350-bp bands. There were no cell lines in which p/tS mRNA was expressed in spite of having the methylated SmaI site in exon I. P0.023 DISCUSSION 12 24 36 Monthes after diagnosis Fig. 3. Kaplan-Meier survival curves for patients with NB classified according to patients with LOH on chromosome 9p. The difference in survival times between the patients with 9p LOH and without 9p LOH was significant by log-rank test. We present here the deletion map of chromosome 9 and alterations of the pi6 gene in NB. In this study, we detected two commonly deleted regions: one was between the IFNB1 and D9Si 7/ loci at chromosome 9p2l and the other was distal to the D9Si76 locus at chromosome 9q34—qter. Recently, it has been demonstrated that both short and long arms of chromosome 9 are frequently deleted in many types of human cancers. Loss of the short arm of chromosome 9, in association was found, but patients with 9p LOH showed poorer survival than patients without 9p LOH (P 0.0486). Mutation of the p16 Gene in NB. Because the pitS gene has been mapped to 9p2l between the JFNB/ and D9S171 markers, PCR-SSCP analysis was performed to screen for p/iS mutations in 19 cell lines and 80 fresh tumors using three sets of primers flanking two exons of the pitS gene. Although SSCP variants were not detected in the 19 cell lines, a tumor-specific SSCP pattern was observed in one of 80 fresh tumors. Sequence analysis of the variant SSCP band revealed a missense mutation at codon 52 from ATG (Met) to AAG (Lys; Fig. 4). Because these patients did not show heterozygous genotypes at 9p2l, we could not determine whether LOH had occurred on chromosome 9p2l. Although several polymorphisms were reported (15), we did not detect such polymorphisms in this study. Previously, we detected polymorphisms only in 3 of 71 Japanese patients (4%) with lung cancers (15). Thus, the population with polymorphic alleles would be rare in Japanese. Expression of the p16 Gene in NB Cell Lines. To investigate the expression pattern of the pitS gene in NB, we performed Northern and Western blot analyses in 19 NB cell lines (Table 2; Fig. 5). pitS linesCell Table 2Status Fig. 4. PCR-SSCP (A) and sequence (B) analy TN 19 cases of NB ceIl geneDNAmRNAProteinMethylationTGW linep'6 IMR-32 WT — - - LAN1 LAN2 LANS NBI NB9 NBI6 NBI9 NB39 NB69 SCMCN2 SCMCN3 SCMCN4 SCMCN5 WT WT WT WT WT WT WT WT WT WT WT WT WT + + - - - + CHP-l34 TNBI GOTO -(, KP-N-NSWV' WT, wild - - - - - - — - + - - - - - + - - - - - - + + - + + - - - + - - + WT + + — WT WT WT+ + + - - - - -+ -- type. B A ses of the p!6 gene in a case of NB. DNA was of the pitS gene iii Tumor ACGT Normal ACGT isolated from a tumor (7) and corresponding normal JrG T 52Met tissue (N) of patient 4. A. abnormal conformers were detected in the 5-region of the p16 gene exon 2. B, point mutation resulting in amino acid change from methionine to lysine was detected at codon 52. _J ,@ I 52 A —Ic IT LG •@•1 T LG 910 Downloaded from cancerres.aacrjournals.org on August 9, 2017. © 1997 American Association for Cancer Research. Lys* CHROMOSOME9 DELETIONSAND p16 IN NEUROBLASTOMA 123456789 A 4.3 kb — . 0.65kb— •. 0.35kb— • B 1 * 2 3 4 56789 0.8 kb C 123456789 p16 —@ Fig. 5. Methylation ofthe 5' CpG island in thepl6 gene and expression ofthepió gene in NB cell lines. N4l7 small cell lung carcinoma cell line and A549 non-small cell lung carcinoma cell line are used as positive and negative controls, respectively. The pitS gene is expressed in N417, whereas it is homozygously deleted in A549. Lane I, N417; Lane 2, A549; Lanes 3—9,NB cell lines, TOW, IMR-32, LAN 1, NB 16, NB69, LAN2, and NB19. A, Southern blot analysis of the 5-region of exon 1 of the p16 gene. Digestion with SmaI plus EcoRI yields two small fragments (0.65 and 0.35 kb) in six cell lines (Lanes 1, N-myc amplification. Although the age of children at diagnosis is also a factor to predict the outcome of patients, 9p LOH was not correlated with the age of patients. This might be due to the small number of patients over 1 year in this study. Therefore, further studies with a large population of children over 1 year may lead to conclusive data for the correlation between age of patients and 9p LOH. Although lp LOH is also considered to correlate with poor survival, we found no association between lp LOH and 9p LOH. Because we used only two markers at lp32 for detection of Ip LOH (12), it could influence the statistical analysis. The pi6 gene, a candidate tumor suppressor gene involved in many types of human cancers (13—15),have been mapped to chromosome 9p2l between the IFNBi and D9Si7i loci, which is one of the commonly deleted regions on chromosome 9 in NB. However, no homozygous deletions have been reported in NB (32). It is also reported that there were no pitS gene mutations and no LOH at the JFNA locus close to the pi6 gene locus (32). In this study, we found no homozygous deletions in both primary tumors and cell lines, and a missense mutation was detected only in a primary tumor. Because this type of mutation has not been reported previously, we do not know if it is functionally significant. These data suggest that the pitS gene is not a target tumor suppressor gene inactivated in NB. How ever, it is possible that the p16 gene is inactivated by alternative mechanisms in most tumors, such as intronic deletions and mutations not detected by sequence analysis of exons or Southern blot analysis. Moreover, recent evidence indicated that transcriptional repression by DNA methylation of promoter and 5' regulatory sequences may be a pathway for inactivation of the pitS gene in several types of human cancers (16—17).To clarify whether the p/tS gene is inactivated in NB, we examined the status of the pi6 gene using Southern, Northern, and Western blot analyses. The pi6 gene was not expressed in most NB cell lines. Absence of the pi6 mRNA in the samples lacking the p16 protein suggests that pitS expression is likely to be regulated at the transcriptional level. Moreover, we found that hypermethylation of the 5' CpG island in the pitS gene is frequent in cell lines lacking p16 expression. Thus, it is likely that the pitS gene is inactivated mostly by 5' CpG island methylations rather than DNA alterations in NB. Similar results were also reported in several other types of human cancers (16—17).However, the mechanisms for the absence of the pi6 mRNA in the remaining cell lines is not clear. We cannot role out the 3.4. 5. 6,and7),indicating thattheSmaIsiteisunmethylated, andonelargefragment(4.3 possibility that mutation harbors in the promoter region of the pi6 kb) in two cell lines (Lanes 8 and 9), indicating that the SmaI site is methylated. B, gene with the consequence of gene inactivation. Recent studies mdi Northern blot analysis of the p16 gene. The pitS transcripts of 0.8 kb were detected in two cated that expression of not only the pi6 gene but also the pi5 gene of NB cell lines (Lanes 3 and 5). C, Western blot analysis of the p16 protein. p16 protein was detected in two of NB cell lines (Lanes 3 and 5). is suppressed by homozygous deletion, point mutation, and hyperm ethylation of the 5' CpG island of this gene in several human cancers (33, 34). Particularly, in leukemia, the methylation of the 5' CpG particular 9p2l—22, occurs in a variety of human cancers, including island in the first exon of the p/S gene is more frequent than that in melanoma (24), renal cell carcinoma (25), lung cancer (26), bladder the pi6 gene (17). Therefore, to determine the pathways to inactivate cancer (27), head and neck cancer (28), and ovarian cancer (29). Chromosome 9q, in particular 9q34—qter, is also frequently deleted in these genes and whether the p15 and/or pi6 genes are involved in the progression of NB, more detailed analysis of these genes will be several human cancers (27, 29—31).Interestingly, both of the short necessary. and long arms are deleted in some human cancers, such as bladder In conclusion, it was demonstrated here that at least two tumor cancer, ovarian cancer, esophageal carcinoma, and renal cell carci suppressor genes on chromosome 9 are involved in the genesis noma (27, 29 —3 1). Therefore, as in the case of several other types of and/or progression of NB. Particularly, the gene on chromosome cancers, at least two tumor suppressor genes on both short and long 9p is likely to be associated with progression of NB, and the pi6 arms of chromosome 9 may contribute to genesis and progression of gene is a candidate target tumor suppressor gene involved in the NB. Furthermore, we found that 9p LOH significantly correlates with progression of NB. However, because we have not examined for an advanced stage of the disease and poor prognosis of the patient, and 9q LOH did not correlate with these clinical parameters. In the present p16 methylation in primary tumors, the biological significance of p16 inactivation in the genesis and progression of NB is still study, 9p LOH was significantly associated with poor prognosis independently of N-myc amplification. Thus, it is possible that a unclear. For this reason, we are currently investigating the asso ciation ofpl6 expression and methylation in tumors with prognosis tumor suppressor gene located on chromosome 9p2l plays an impor (ant role in the progression of NB through a different pathway from of patients with NB. 911 Downloaded from cancerres.aacrjournals.org on August 9, 2017. © 1997 American Association for Cancer Research. CHROMOSOME9 DELETIONSAND p16 IN NEUROBLASTOMA ACKNOWLEDGMENTS silencing of the tumor suppressor p16/CDKN2JMTSI in human cancers. Nat. Med., 1: 686—692, 1995. 17. Herman, J. 0., Merlo, A., Mao, L., Lapidus, R. G., Issa. J-P. J., Davidson, N. E., We thank Dr. S. Nagata for providing the IFNBI cDNA probe. Sidransky, D., and Baylin. S. B. Inactivation of the CDKN2JpI6JMTSI gene is frequently associated with aberrant DNA methylation in all common human cancers. Cancer Res., 55: 4525—4530, 1995. 18. Evans, A. E., D'Angio, G. J., and Randolph. J. A proposed staging for children with REFERENCES 1. Weinberg. R. A. Tumor suppressor genes. Science (Washington DC), 254: 1138— 1146, 1991. 2. Yokota, J., and Sugimura, T. Multiple steps in carcinogenesis involving alterations of multiple tumor suppressor genes. FASEB J., 7: 920—925. 1993. 3. Fong, C. T., Dracopoli, N. C., White, P. S., Merrill, P. 1.. Griffith, R. C., Housman, D. E., and Brodeur, 0. M. Loss of heterozygosity for the short arm of chromosome I in human neuroblastomas. Proc. NatI. Acad. Sci. USA, 86: 3753—3757, 1989. 4. Suzuki. T., Yokota, J., Mugishima, H., Okabe. I., Ookuni, M., Sugimura, 1., and Terada. M. Frequent loss of heterozygosity on chromosome l4q in neuroblastoma. Cancer Res., 49: 1095—1098,1989. 5. Fong. C. T.. White, P. 5., Peterson, K., Sapienza, C., Cavenee, W. K., Kern, S. E., Vogelstein, B., Cantor, A. B., Look, A. 1., and Brodeur. 0. M. Loss of heterozygosity for chromosomes I and 14 defines subsets of advanced neuroblastoma. Cancer Res., 52: 1780—1785,1992. 6. Takayama. H.. Suzuki. 1., Mugishima, H., Fujisawa. 1., Ookuni, M., Schwab, M.. Gehring, M.. Nakamura, Y., Sugimura, T., Terada, M., and Yokota, J. Deletion map of chromosome l4q and Ip in neuroblastoma. Oncogene, 7: 1185—1 189, 1992. 7. Caron, H., van Sluis, P., van Hoeve. M.. de Kraker, J., Bras, J., Slater. R., Mannens, M., Voute, P. A.. Westerveld. A., and Versteeg. R. Allelic loss of chromosome lp36 in neuroblastoma is of preferential matemal origin and correlates with N-myc am plification. Nat. Genet., 4: 187—190,1993. 8. Srivatsan, E. S., Ying, K. L.. and Seeger, R. C. Deletion of chromosome I I and l4q sequences in neuroblastoma. Genes Chromosomes Cancer, 7: 32—37,1993. 9. Schleiermacher, G., Peter, M., Michon, J., Hugot, J-P., Vielh, P., Zucker, J-M., Magdelenat. H., Thomas, G., and Delattre, 0. Two distinct deleted regions on the short arm of chromosome I in neuroblastoma. Genes Chromosomes Cancer, 10: 275—281, 1994. 10. Takeda, 0., Homma, C., Maseki, N., Sakurai, M., Kanda, N., Schwab, M., Nakamura, Y., and Kaneko, Y. There may be two tumor suppressor genes on chromosome arm Ip closely associated with biologically distinct subtype of neuroblastoma. Genes Chromosomes Cancer, 10: 30—39.1994. I I. Tonini, G. P. Neuroblastoma: a multiple biological disease. Eur. J. Cancer, 29A: 802—804, 1993. 12. Takita, J., Hayashi, Y., Kohno, T.. Shiseki, M., Yamaguchi, N., Hanada, R., Yamamoto, K., and Yokota, J. Allelotype of neuroblastoma. Oncogene, 11: 1829— 1834, 1995. 13. Kamb, A., Gruis, N. A., Weaver-Feldhaus, J., Liu, Q., Harshman, K.. Tavtigian, S. V., Stockert, H., Day. R. S.. III. Johnson. B. E., and Skolnick, M. H. A cell cycle regulator potentially involved in genesis of many tumor types. Science (Washington DC). 264: 436—438. 1994. 14. Nobori, T., Miura, K., Wu, D. J., Lois, A., Takabayashi. K., and Carson, D. A. Deletion of the cyclin-dependent kinase-4 inhibitor gene in multiple human cancers. Nature (Lond.), 368: 753—756,1994. @ 15. Okamoto. A.. Hussain. S. P., Hagiwara, K.. Spillare, E. A., Rusin. M. R., Demertrick, D. J., Serrano, M.. Hannon, 0. J., Shiseki. M.. Zariwala, M., Xiong, Y., Beach, D. H., Yokota. J.. and Harris, C. C. Mutations in the p16@4/MTS1/CDKN2, pJ5t@45/ MTS2, and p!8 genes in primary and metastatic lung cancer. Cancer Res., 53: 382—387,1995. 16. Merlo, A., Herman, J. G., Mao, L.. Lee, D. J., Gabrielson, E., Burger, P. C., Baylin. S. B.. and Sidransky. D. 5' CpG island methylation is associated with transcriptional neuroblastoma. Cancer (Phila.), 27: 374—378,1971. 19. Ota, K., and Shimizu, K. Histological Classification and Atlas of Tumors in Infancy and Child: a Report from the Committee on the Histological Classification of 20. 21 . 22. 23. 24. 25. 26. 27. 28. 29. 30. 31. Childhood Tumors of the Japanese Pathological Society. Tokyo: Kanahara Publisher. 1980. Maniatis, T.. Fritsch, F. F.. and Sambrook, J. Molecular Cloning: A Laboratory Manual. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory. 1982. Sameshima, Y., Matsuno, Y., Hirohashi, S., Shimosato, Y., Mizuguchi. H.. Sugimura. T.. Terada, M., and Yokota, J. Alterations of the p53 gene are common and critical events for the maintenance of malignant phenotypes small-cell lung carcinoma. Oncogene. 7: 451—457,1992. Hussussian, C. J., Struewing, J. P., Goldstein, A. M., Higgins. P. A. T.. Ally, D. S., Sheahan, M. D., Clark, W. H., Jr., Tucker, M. A.. and Dracopoli, N. C. Germline p/tS mutation in familial melanoma. Nat. Genet., 8: 15—12,1994. Ookawa, K., Sakamoto, M., Hirohashi, S., Yoshida, Y., Sugimura. 1.. Terada, M., and Yokota. J. Concordant p53 and DCC alterations and allelic losses on chromosomes I 3q and 14q associated with liver metastases of colorectal carcinoma. Int. J. Cancer, 53: 382—387,1993. Cannon-Albright, L. A., Golgar, D. E.. Meyer, L. J., Green, R., MacLennan, N. G., Martin, L. J.. and Kamb, A. Assignment of a locus for familial melanoma. MLM, to chromosome 9pl3—22. Science (Washington DC), 258: 1148—1152, 1993. Paul, C., Tokino. K., Eby, Y., and Sidransky. D. Localization oftumor suppressor loci on chromosome 9 in primary human renal cell carcinoma. Cancer Res., 55: 224—227, 1995. Merlo, A., Gabrielson, E., Askin, F., and Sidransky, D. Frequent loss of chromosome 9 in human primary non-small cell lung cancer. Cancer Res., 54: 640—642, 1994. Ruppert, J. M., Tokino, K., and Sidransky, D. Evidence for two bladder cancer suppressor loci on human chromosome 9. Cancer Res., 55: 5093—5095, 1995. Riet, P., Nawroz, H.. Hruban, R. H., Corio, R., Tokino, K., Koch, W., and Sidransky, D. Frequent loss of chromosome 9p2l—22early in head and neck cancer progression. Cancer Res., 54: 1156—1 158, 1994. Schultz, D. C., Vanderveer, L., Buetow, K. H.. Boente, M. P., Ozols, R. F., Hamilton, T. C., and Godwin, A. K. Characterization of chromosome 9 in human ovarian neoplasia identifies frequent genetic imbalance on 9q and rare alterations involving 9p, including CDKN2. Cancer Res., 55: 2150—2157, 1995. Caims, P., Tokino, K., Eby, Y., and Sidransky, D. Localization of tumor suppressor loci on chromosome 9 in primary human renal cell carcinoma. Cancer Res., 55: 224—227, 1995. Miura, N., Okita, K., Furukawa, Y., Matsuno S., and Nakamura, Y. Deletion mapping in squamous cell carcinomas of theesophagusdefinesa regioncontaining a tumor suppressor gene within a 4-centimorgan interval of the distal long arm of chromosome 9. Cancer Res., 55: 1828—1830,1995. 32. Beltinger. C. P.. White. P. S., Sulman, E. P.. Mans. J. M.. and Brodeur, G. M. No CDKN2 mutation in neuroblastomas. Cancer Res., 55: 2053—2055,1995. 33. Herman, J. G., Jen, J.. Merlo, A.. and Baylin, S. B. Hypermethylation-associated inactivation indicated tumor suppressor role for @j5@@@4ttI Cancer Res., 56: 722—727.1996. 34. Jen. J., Harper. J. W.. Bigner, S. H., Bigner. D. D., Papadopoulos. N.. Markowitz, S., Willson, J. K. V.. Kinzler, K. W., and Vogelstein. B. Deletion of p/iS and p/S gene in braintumors.CancerRes.,54: 6353—6358, 1995. 912 Downloaded from cancerres.aacrjournals.org on August 9, 2017. © 1997 American Association for Cancer Research. Deletion Map of Chromosome 9 and p16 (CDKN2A) Gene Alterations in Neuroblastoma Junko Takita, Yasuhide Hayashi, Takashi Kohno, et al. Cancer Res 1997;57:907-912. Updated version E-mail alerts Reprints and Subscriptions Permissions Access the most recent version of this article at: http://cancerres.aacrjournals.org/content/57/5/907 Sign up to receive free email-alerts related to this article or journal. To order reprints of this article or to subscribe to the journal, contact the AACR Publications Department at [email protected]. To request permission to re-use all or part of this article, contact the AACR Publications Department at [email protected]. Downloaded from cancerres.aacrjournals.org on August 9, 2017. © 1997 American Association for Cancer Research.