* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download gene expression analysis of chondrocyte mechanical response by
Non-coding RNA wikipedia , lookup
Epitranscriptome wikipedia , lookup
List of types of proteins wikipedia , lookup
Magnesium transporter wikipedia , lookup
Secreted frizzled-related protein 1 wikipedia , lookup
Protein moonlighting wikipedia , lookup
Protein adsorption wikipedia , lookup
Genome evolution wikipedia , lookup
Transcriptional regulation wikipedia , lookup
Genomic imprinting wikipedia , lookup
Community fingerprinting wikipedia , lookup
Western blot wikipedia , lookup
Proteolysis wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Gene regulatory network wikipedia , lookup
Ridge (biology) wikipedia , lookup
Signal transduction wikipedia , lookup
Expression vector wikipedia , lookup
Molecular evolution wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Gene expression wikipedia , lookup
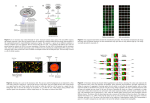
GENE EXPRESSION ANALYSIS OF CHONDROCYTE MECHANICAL RESPONSE BY LARGE SCALE DNA MICROARRAY +Kanbe, K; +Inoue, K; *Xiang, C; **Wei, L; **Chen, Q +Tokyo Women’s Medical University/Daini Hospital, Tokyo, Japan *NIH/NIMH, **Brown Medical School/Rhode Island Hospital Introduction: Mechanical stress-induced matrix deformation plays a fundamental role in regulating cellular activities. Abnormal mechanical loading is a hallmark for osteoarthritis development. Previously, we have shown that mechanical stress plays an important role in regulating chondrocyte proliferation and differentiation. We also identified several mechanoresponsive genes in cartilage, such as Indian hedgehog, type X collagen, and matrilin-1, which happen to be up regulated in OA cartilage. In this study, we test the hypothesis that mechanoresponsive genes in cartilage play an important role for transducing mechanical signals to chondrocytes. To test this hypothesis, we identified mechanoresponsive genes in cartilage by large scale DNA microarray analysis. Conventional studies of gene expression and regulation have been restricted to analysis of individual genes. In contrast, DNA microarray technology provides a format for simultaneous measurement of the expression level of thousands of genes in a single hybridization assay. With DNA microarray, DNA probes representing thousands of cDNA clones are arrayed onto glass slides and hybridized with fluorescence-labeled cDNA derived from chondrocyte mRNA. Our study was designed to determine the change of gene expression profile in mouse chondrocytes subjected to mechanical stress induced deformation. We determined the alteration of gene expression in response to mechanical stress with cDNA microarray containing 12,000 sequenceverified mouse genes, about a third of total mouse genome. Methods: 3D chondrocytes culture system: Primary chondrocytes were isolated from rib cages of new born mice by digestion with collagenase D (3 mg/ml) for 6 hours. Chondrocytes were collected and seeded onto culture plates in DMEM contained 10% FCS and 1% of penicillinstreptomycin. After overnight incubation, chondrocytes were suspended by trypsin digestion. One-hundred micro liters of cell suspension containing 1 million chondrocytes were applied to 2x2x0.25-cm Gelform sponges presoaked with Hanks’ balanced salt solution (HBSS). After overnight incubation, the sponges were subjected to cyclic deformation with a Bio-Stretch device to induce 5% elongation for 15 min/h at 1 Hz for periods of 2 and 4 days for microarray. Chondrocyte growth analysis: Collagen sponges containing chondrocytes were digested with 0.03% collagenase, followed by collection of chondrocytes by centrifugation. Chondrocytes were counted with a hemacytometer. The viability of cells was confirmed by trypan blue exclusion assay. Isolation of mRNA: Total RNA from chondrocytes cultured in sponges was extracted with RNeasy mini kit. Quantification of mRNA including type X collagen and Indian hedgehog (Ihh) was performed by real-time quantitative reverse transcriptase (RT)-PCR with a Perkin-Elmer ABI Prism 7700 sequence detection system. DNA Microarray: Five micro grams of total RNA were reverse transcribed in the presence of Aminoallyl dUTP. cDNA from stretched chondrocytes was coupled with Cy5, while cDNA from non-stretched chondrocytes was coupled with Cy3 as reference. After hybridization, labeled slides were scanned with GenePix 400A. Microarray data were analyzed by IPLab software. Results: Identified mechanoresponsive genes consist of candidate components of mechanotransduction pathways including extracellular matrix protein, ion channels, cytoskeleton, and nuclear transcriptional factors. About 1 percent of the genes alter their expression significantly in response to mechanical stress. The amplitude of the change of gene expression by mechanical stress is modest, with most changes below 3fold. Biomechanically up-regulated genes after 2 days of mechanical loading include those encoding extracellular matrix such as perlecan, HA binding protein, osteopontin and follistatin, those encoding cytoskeletal proteins such as beta spectrin2 and vimentin, those encoding cytoplasmic proteins such as elongation factor 2 and cholesterol 24-hydroxylase, and those encoding nuclear proteins including SOX-1 (Table 1). Downregulated genes in response to mechanical stress include those encoding membrane proteins ATPase, cytoplasmic protein retinol binding protein 1 and nuclear protein CLK2 (protein kinase). After 4 days of cyclic deformation, up-regulated chondrocyte genes including those encoding membrane proteins such as Pak3 binding protein, those encoding cytoplasmic protein such as ubiquitin specific protease 25, and those encoding nuclear proteins such as transcription factor CA150. Down regulated genes include collagen alpha 2 (VI) chain precursor in the matrix, NMDA1 in the membrane, and CLOCK in the nucleus. Discussion: Our microarray data not only confirmed mechanosensitive genes identified previously, such as osteopontin and glutamate receptor NMDA1, but also suggested unexpected genes, such as those in retinoic acid signaling and circadian clock regulation. Since this is one of the first analyses of chondrocyte mechanical response using large scale DNA microarray, validation of the microarray data at three different levels was carried out. First, known mechanoresponsive genes such as matrilin-1 and Indian hedgehog were confirmed to be up-regulated by mechanical stress in mRNA samples for microarray by real-time RT-PCR. Thus, these RNA samples can be used for microarray analysis. Second, mechanoresponsive genes identified by microarray, such as osteopontin, is verified by real-time RT-PCR from the same RNA samples. Therefore, microarray analysis is accurate for quantifying mRNA changes. Third, mechanoresponsive genes identified previously, such as osteopontin and NMDA1, are confirmed by our microarray analysis. Therefore, our microarray analysis is reliable and can be used to identify new mechanoresponsive genes. Interestingly, we identified a group of extracellular matrix proteins in cartilage to be mechanoresponsive molecules. They include perlecan, osteopontin, HA binding protein and type VI collagen. Among them, osteopontin and type VI collagen are known to be upregulated in OA cartilage. Perlecan is also implicated to play a mechanical role during cartilage growth and differentiation as its mutation causes severe chondrodysplasia with dyssegmental ossification. In conclusion, identification of mechanoresponsive genes in cartilage may help to better understand the mechanism underlying abnormal activation of genes in OA and other pathological cartilage that experiences abnormal mechanical loading. Location Description matrix perlecan 1.947 matrix HA binding protein 1.898 matrix osteopontin sexreted follistain 1.799 cytoskeleton B-spectrin 2 1.855 cytoskeleton vimentin 1.847 cytoplasm proteasome ru93 1.774 cytoplasm (3158)ribosomal protein 1.764 cytoplasm thiolase peroxisomal 1.764 cytoplasm elomgation factor 2 1.731 cytoplasm ribosomal protein L3 1.723 cytoplasm ribosomal protein L18 1.715 cytoplasm Laminin receptor 1.712 cytoplasm aldehyde reductase 1.708 ER membrane cholesterol 24-hydroxylase 1.891 nuclear tax interaction protein 1.811 nuclear SOX-1 1.721 nuclear YY1 interaction protein 1.717 50th Annual Meeting of the Orthopaedic Research Society Poster No: 0818 Ratio 1.85