* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Supporting Information Legends Supplementary Table S1
Point mutation wikipedia , lookup
Transcriptional regulation wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Clinical neurochemistry wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Secreted frizzled-related protein 1 wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Mitogen-activated protein kinase wikipedia , lookup
Western blot wikipedia , lookup
Homology modeling wikipedia , lookup
Bimolecular fluorescence complementation wikipedia , lookup
Biochemical cascade wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
Gene expression wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Magnesium transporter wikipedia , lookup
Gene regulatory network wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Interactome wikipedia , lookup
Proteolysis wikipedia , lookup
Signal transduction wikipedia , lookup
Paracrine signalling wikipedia , lookup
Protein–protein interaction wikipedia , lookup
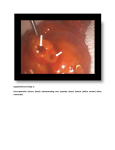
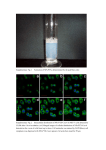
Supporting Information Legends Supplementary Table S1: Complete list of cDNAs identified by the yeast two hybrid split ubiquitin system using SlSUT2 cloned in met25pXCgate as bait. The accession number of proteins with highest homology are given and the percentage of sequence identity. In addition EST numbers from Solanum tuberosum are given. Supplementary Table S2: Quantification of interaction strength by photometric ßgalactosidase assay. StDev was calculated from three independent experiments. Strongest interaction is detectable between SlSUT2 and the BAK1-like and the MSBP1 prey protein. Quantification of the interaction between StSUT1 and StSUT4 with the PDI protein published earlier (Krügel et al., 2012) was performed as positive control of the assay. Cotransformation of the bait constructs together with the empty prey vector served as negative control. Supplementary Figure S1. Subcellular localization of SUT2 in yeast and in plants. a. StSUT2-GFP fusion protein expressed in the yeast mutant strain SuSy7 under control of the Adh1 promoter in the yeast expression vector 112A1NE (Reinders et al., 2002). b. SlSUT2YFP fusion protein expressed under control of the 35S promoter in the plant binary GATEWAY vector pK7GW2.0 in transiently expressing leaf epidermis cells of Nicotiana tabacum transiently transformed by Agrobacterium infiltration. c. The same SlSUT2-YFP fusion construct expressed in stably transformed Nicotiana tabacum plants. Note that SUT2 proteins do not efficiently reach the plasma membrane of overexpressed heterologously. Supplementary Figure S2. Yeast two hybrid split ubiquitin system a. Yeast growth on SD medium without HULT in the presence of 15 mM 3-AT of yeast strain L40ccU transformed with SlSUT2 in Cub as a bait and with the empty Nub vector as a control. Full length cDNAs of PDI, MSBP1 and BAK1-like were cloned into the Nub vector to confirm interaction with SlSUT2. b. Quantification of interaction by photometric ß-galactosidase assay. Strongest interaction with SlSUT2 was observed for MSBP1 and BAK1-like. Supplementary Figure S3. Tissue-specific expression pattern of genes encoding SlSUT2-interacting proteins as determined by qPCR. Total RNA was extracted from different flower organs and from sink and source leaves and used for qRT-PCR. Transcript accumulation of the StMSBP1 gene (a) and the StBAK1-like gene (b) is calculated using ubiquitin as internal reference gene and standard deviation is given. Note that both genes coding for SlSUT2-interacting proteins show main expression in anthers of mature flowers as previously shown for SlSUT2 (Hackel et al. 2006) indicating that interaction partners are co-expressed. Supplementary Figure S4. Sequence alignment of BAK1-like receptor kinases and BRI1 receptor kinases. The BAK1-like protein from potato (XP_006342998) shows 51% identity to BAK1 from Ricinus communis (XP_002510316) and 75% identity to a receptor kinase from Arabidopsis (At3g13690). Alignment was performed with full length amino acid sequences from BAK1 from Arabidopsis thaliana (NP_001190904), Nicotiana attenuata (ADT91693), Sacharum officinarum (ABX83681), Saccharum hybrid cultivar (AHF18212), Ricinus communis (XP_002510316), Solanum lycopersicon (XP_004235609) and with the BRI1 Sequences from Arabidopsis thaliana (NP_195650), Nicotiana benthamiana (ABO27628)and Solanum tuberosum (ABO27627) according to Clustal W using the DNASTAR Laser Gene MegAlign software version 7.1.0 Supplementary Figure S5. Co-expression analysis of At3g13690 reveals co-expression with the Arabidopsis cell elongation protein Dwarf1/DIMINUTO (At3g19820). Co-expression analysis was performed with the ATTED-II software (http://atted.jp/top_search.shtml) Supplementary Figure S6. Hypothetical model illustrating SlSUT2-brassinosteroid interaction. The model illustrates a potential sucrose efflux into the periarbuscular space from where it is retrieved by SlSUT2 back into the plant root cells. Efflux might potentially be mediated by still uncharacterized SWEET proteins which are known to act as sugar efflux carrier (Chen et al., 2010). Alternatively, sucrose is cleaved by the cell wall invertase LIN6 that is inducible by brassinosteroids and cleavage products are taken up by a monosaccharide transporter into the AM fungus. SlSUT2 forms a complex with MSBP1 and the receptor like kinase BAKL. Interaction between MSBP1 and BAKL is assumed to prevent BR signaling via the BR receptor BRI1. BR and BR signaling positively affect LIN6 activity and inhibit defense responses against biotrophic AM fungi. The model is based on experimental data in this work and on published data from tomato, Rhizophagus irregularis and Arabidopsis (Goetz et al., 2000, Song et al., 2009).