* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download The relationship between amino acid sequences and protein folds.
Gene expression wikipedia , lookup
Genetic code wikipedia , lookup
Biosynthesis wikipedia , lookup
Biochemical cascade wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Transcriptional regulation wikipedia , lookup
Biochemistry wikipedia , lookup
Interactome wikipedia , lookup
Western blot wikipedia , lookup
Point mutation wikipedia , lookup
Metalloprotein wikipedia , lookup
Proteolysis wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Clinical neurochemistry wikipedia , lookup
Ligand binding assay wikipedia , lookup
Homology modeling wikipedia , lookup
Paracrine signalling wikipedia , lookup
Protein structure prediction wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Signal transduction wikipedia , lookup
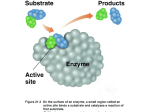
Protein Structure and Function Instructor: ! ! James Omichinski! ! Contact Information:! ! E-mail: ! ! [email protected]! Section #1: The relationship between amino acid sequences and protein folds. Correlation between sequence and protein fold. There are two competing models: Model 1: !The Local Model" •! fold specificity is defined by a few critical residues. •! this model is supported by by misfolding mutations associated with certain diseases such as cystic fibrosis. Model 2: !The Global Model" •! the entire sequence of the protein contributes equally to the fold. •! this model is supported by mutation studies that show most mutations at any position have no measurable impact on protein function. The case of the IgFF fold. If one examines the members of the immunoglobulin fold family (IgFF): •! Many members of the IgFF are distally related proteins or evolutionarily unrelated proteins •! Despite their common fold there is low sequence identity between subgroups suggesting both convergent and divergent evolution. •! It seems the common connection for all IgFF members are residues that form a common hydrophobic core. •! This supports the global model. Reference: Halby et al, Prot. Engineering 12, 563-571 (1999) The Common Core for IgFF Proteins. The conserved core region is sheet I (green) and sheet II (blue) of 52 IgFF members with less than 55% sequence identify. ! ! There are wide variations in the loop (red) regions of IgFF members. These regions have developed as part of the evolutionary process.! ! Relation Between IgFF Sequence and Structure Zone I! Zone II! Zone I: High identity and similar structures.! Zone II: Low identity and similar structures.! Sequence Identity, Homology and Fold •! Amino acid homology is a better criteria then amino acid identities. •! General rule is that sequences that share 30% sequence homology will fold similarly. •! In the immunoglobulin fold family (IgFF) there can be <10% identity. •! In IgFF the residues constituting the core are concentrated in a small number of conserved positions that define the fold. Reference: Wood and Pearson, J. Mol. Biol.291, 977-995 (1999) Protein G as a Model for Studying Cores •! Protein G contains two types of domains (GA and GB). •! GA binds to human serum albumin and has a 3-! fold . •! GB binds to Fc region of IgG and has a 4! + " fold. •! It is possible to make two proteins, a GA like domain and a GB like domain that share 77% identity in 56 amino acids (GA77 and GB77). •! GA77 and GB77 have different folds and different functions despite having 77% identity. Reference: P.A. Alexander et al., Proc Natl Acad Sci (2009), V106:21149-54 GA77 versus GB77 GB77 GA77 Reference: P.A. Alexander et al., Proc Natl Acad Sci (2009), V106:21149-54 Interconverting GA77 and GB77 •! The goal was to interconvert the two proteins with the fewest number of mutations. •! The conversion should change both the structure and the function to be considered successful. •! It turns out that it is possible to do this with a single point mutation. Reference: P.A. Alexander et al., Proc Natl Acad Sci (2009), V106:21149-54 A point mutant converts GA77 to a GB77 Reference: P.A. Alexander et al., Proc Natl Acad Sci (2009), V106:21149-54 Section #2: Intrinsically disordered proteins. The case of acidic activation domains Intrinsically disordered domains in proteins •! Many disordered segments fold on binding (coupled folding and binding). •! It is now clear that the occurrence of disordered regions is surprisingly common in functional proteins. •! These regions can be highly conserved between species in both amino acid composition and sequence. •! These regions are often characterized by a low content of hydrophobic amino acids and a high concentration of polar or charged amino acids. •! They are particularly abundant in transcription factors, signaling proteins, autoinhibitory domains and viral proteins. •! The classic example of functional unstructured region is the transactivation domain (TAD) of transcription factors. Liu et al., PNAS USA 106: 19819-23 (2009). PIC, HATs, remodeling and mediator Role of activators in transcription •! Activators function through the recruitment of general transcription factors, the mediator complex, histone acetyl transferases and the chromatin remodeling proteins. •! Activators function by participating in a series of protein/protein interactions through their transactivation domain (TAD). •! TADs are characterized by a high percentage of a particular amino acid and are generally thought to be unstructured in the unbound state. •! The tumour suppressor protein p53 and the Herpes Virion Simplex protein 16 (VP16) are two of the most potent activators known and they are acidic-rich. PIC, HATs, remodeling and mediator TFIIH and TADs TFIIH Core TFIIH CAK Complex •! TFIIH is recruited to the PIC through interactions with p62 /Tfb1 (human/yeast) and the acidic carboxyl-terminal domain of TFIIE!. •! Several acidic TADs interacts directly with the p62/Tfb1 subunit of TFIIH and this interaction correlates with their ability to activate initiation and elongation. •! The interaction of p62/Tfb1 with p53 and VP16 requires the aminoterminal Pleckstrin Homology (PH) domain of p62/Tfb1. p53 •! p53 is a potent transcriptional activator. •! It induces expression of many genes whose products mediate cellgrowth arrest and apoptosis, thus blocking cell transformation and tumor formation. •! Mutations in p53 leading to inactivation of its tumor suppressor function leads to the development of about 50% of human cancers. •! A large percentage of the critical mutations occur within the DNA binding domain of p53. •! Amino terminal 100 amino acids contains a extremely potent activation domain that is highly acidic and unstructured. Schematic of the functional domains of p53 Proline N-terminal Transactivation Rich Domain Domain TAD PP DNA Binding Domain Tetradimerization domain DBD TET REG C-terminal Regulatory Domain P P P P P P S6 S8 S15 T18 S20 TAD1 MDM2 P P S33 S37 P S46 T55 P T81 TAD2 Tfb1/p62 •! The transcriptional activation function of p53 is associated with its highly acidic TAD, which can be divided into two independent subdomains, TAD1 (residues 1-40) and TAD2 (residues 40-83). Structure of the Tfb1/p53 complex The structure of Tfb1 in the complex presents a typical PH fold. The p53 TAD2 in the complex is unstructured except for a short amphipathic !-helix involving residues 47-55. Di Lello et al., Molecular Cell, V22, pg731, 2006. Details of the Tfb1/p53 interface N C Three hydrophobic residues I50, W53 and F54 of p53 form the interface with Tfb1. (Di Lello et al., Mol. Cell, 2006) Schematic of the functional domains of VP16 Protein/protein Interaction Domain C-terminal Transactivation Domain TAD Core •! VP16 is composed of two functional domains: A core regulatory domain and a TAD (residues 412-490). •! The acidic TAD can be divided into two independent subdomains, VP16N (residues 412-455) and VP16C (residues 456-490). (pI= 3.43) VP16N VP16C Structure of Tfb1/VP16C VP16C N "7# "6# C "5# N C Tfb1 Like p53 TAD2, VP16C in complex with Tfb1 is unstructured except for a 9-residue ! helix between residues 472-480. Langlois et al., J. Am. Chem. Soc., 130: 10596, 2008 Comparison of Tfb1/p53 and Tfb1/VP16 c p53 C M478! C W53 VP16 F54 I50 F479! N F475! N - F475, M478 and F479 of VP16 in the Tfb1/VP16 complex participate in many of the same interactions as I50, W53 and F54 of p53 in the Tfb1/p53 complex. Langlois et al., J. Am. Chem. Soc., 130: 10596, 2008 Comparison of Tfb1/p53 and Tfb1/VP16 VP16 p53 T480! T55 E51 E476! D472! S46 •! E476 and T480 of VP16 in the Tfb1/VP16 complex participate in the same interactions as E51 and T55 of p53 in the Tfb1/p53 complex. However, S46 of p53 corresponds to A471 of VP16. Langlois et al., J. Am. Chem. Soc., 130: 10596, 2008 Summary •! The viral activator VP16 can mimic many of the properties of the mammalian activator p53 in complex with Tfb1/p62.! •! Hydrophobic and acidic residues play key roles in defining the interface between acidic TADs and Tfb1/p62.! ! •! Select activators like p53 have evolved so that phosphorylation events play a key role in their regulation. This is not the case for the viral activator VP16. Why disordered regions? •! Allow protein to bind to multiple partners using lower affinity interactions! •! Enhance probability of posttranslational modifications such as phosphorylation, acetylation and methylation.! ! •! Enhance turn over rates of proteins, turn on and turn off theory. Section #3: Membrane Proteins. Types of transmembrane receptors •! 7-transmembrane segment (7-TMS) receptors- contain a seven transmembrane (helical) segment, an extracellular ligand binding site and and intracellular recognition site for a GTP-binding protein. •! Single-transmembrane (1-TMS) catalytic receptors- contain a single transmembrane segment, an extracellular ligand domain and intracellular catalytic domain (tyrosine kinase or guanylyl cyclase) •! Oligomeric ion channels- contain more than one subunit each of which contains transmembrane segments, they are ion-ligated so binding of ligand opens the ion channel. Structures of transmembrane proteins •! The ones solved to date are minimal since it is very hard to mimic the membrane using NMR or X-ray techniques. •! Two types of structures have been observed so far, either all ! helical or " barrels. •! You need 20-25 residues in an ! helical sequence to span the thickness (30Å) of the lipid bilayer. •! You need only 7-9 residues in a " conformation to span the lipid bilayer since it is a more extended conformation. •! Hydrophathic plots can be used to predict ! helical transmembrane domains but not " barrels. Bicontinuous lipidic cubic phase •! This system was developed to help obtain better crystals of membrane proteins. •! Once inserted into this continuous 3D curved lipid bilayer, membrane proteins diffuse laterally to nucleate and eventually to form well-ordered crystals. •! Used to crystallize bacteriorhodopsin and rhodopsin. Rhodopsin •! It is a G-protein-coupled receptor (GPCR) •! Rhodopsins are largest subfamily (90%) •! They are light activated and turn on a signaling pathway that leads to vision. •! Composed of protein opsin and 11-cis retinal through K296. •! Absorption of photon changes 11-cis form to all-trans-retinal. •! The process is transient as trans-retinal is hydrolyzed and dissociates from the opsin to be replaced by newly synthesized 11cis retinal. Schematic of a 7-TMS receptor Structure of Rhodopsin Crystals were generated in micelles! ! Helices are arranged differently then! in Bacteriorhodpsin (bR).! ! The extramembrane regions are larger! and more ordered in Rhodopsin then in ! bR which demonstrates the functional ! difference.! ! Reference: Palczewski et al, Science 289 739-745! (2000)! The retinal binding pocket A269 and F261 are responsible for absorption differences! between red and green pigments.! "2 Adrenergic Receptor ("2AR) •! It is a G-protein-coupled receptor (GPCR) •! The ! and " andrenergic receptors differ in tissue localization, ligand specificity, G-protein coupling and downstream effector mechanisms •! Genetic modifications of andrenergic receptors are associated with a variety of human diseases including asthma, hypertension and heart failure. •! "2AR#s are found in smooth muscles throughout the body. •! "2AR agonists are used in treatment of asthma and they are of considerable interest to the pharmaceutical industry. •! It differs from rhodopsin in that the ligand is freely diffusible and not covalently bound. "2 Adrenergic Receptor ("2AR) "2AR Crystal Structure •! There have been several crystal structures GPCR over the last five years that are based on some interesting tricks to help cyrstallize membrane proteins. •! The first one solved was the structure of the "2AR (Cherezov et al, Science V318, pg 1258, 2007) and two different techniques were used. •! In the higher resolution (2.4 Å) structure, the third intracellular loop was replaced by inserting the sequence for T4-lysozyme). •! In the lower resolution (3.4/3.7 Å) structure, the "2AR was crystallized in complex with a Fab that binds the third intracellular loop. •! In both structures, the "2AR was crystallized in the presence of the partial inverse agonist carazolol (Inactive conformation). •! These structures have led to several other structures of GPCRs using variations of the T4-lysozyme strategy. The two "2AR structures:T4L vs FAB5 2.4Å resolution 3.4/3.7Å resolution Rosenbaum et al, Science, 2007. The T4L-"2AR structure Cherezov et al, Science, 2007. The T4L-"2AR structure Carazolol "2AR Bound lipids (Cholesterol and palmitic acid) T4 Lysozyme Cherezov et al, Science, 2007. The T4L-"2AR structure:Contacts with T4L There are only a limited number of intramolecular interactions between the T4 Lysozyme (T4L-green) and the "2AR (gold). Cherezov et al, Science, 2007. The T4L-"2AR structure:Ligand binding site Binding site cleft Negative charge Positive charge Cherezov et al, Science, 2007. Ligand Binding:T4L-"2AR versus rhodopsin "2AR Rhodopsin "2AR ECL2 (A) contains a short helix and two disulfide bonds (yellow). One disulfide bond connects to ECL1 the second connects to helix III. Phe193 of ECL2 interacts with the ligand carazolol (purple). In contrast, ECL2 in rhodopsin (B) does not have direct access to retinal (peach). Cherezov et al, Science, 2007. More Recent "2AR Crystal Structure •! There has been three new crystal structures of the "2AR using similar techniques and in two cases with a new twist. •! There are two crystal structures of the "2AR with an agonist bound in the active site (active confirmation). One was done with the T4L in the third EC loop as before (Rosenbaum et al, Nature V469, pg 236, 2011) and one was done with the T4L and a nanobody (Rasmussen et al, Nature V469, pg 175, 2011). •! There is also a crystal structure of the "2AR with an agonist bound in the active site bound to a Gs protein. In this complex, the T4L was placed at the amino-terminal end of the protein and a nanobody was also used (Rasmussen et al, Nature V477, pg 549, 2011). G Protein Cycle for the "2AR –Gs Complex Rasmussen et al, Nature(2011), 477: 549-555. "2AR –Gs Complex Rasmussen et al, Nature(2011), 477: 549-555. Other Crystal Structures of GPCRs A2A Adenosine receptor- Jaakola et al, Science (2008), 322: 1211-1217. CXCR4 Chemokine Receptor- Wu et al, Science (2010), 330: 1066-1071. Sphingosine 1-phosphate Receptor- Hanson et al, Science (2012), 335: 851-855. M2 Muscarinic receptor- Haga et al, Nature(2012), 482: 547-552. M3 Muscarinic receptor- Kruse et al, Nature(2012), 482: 552-559. Kappa-opioid receptor- Wu et al, Nature(2012), 485: 327-334. FQ-opioid receptor- Thompson et al, Nature(2012), 485: 395-400. Delta-opioid receptor- Granier et al, Nature(2012), 485: 400-405. µ$opioid receptor- Manglik et al, Nature(2012), 485: 321-327. The A2A$denosine Receptor Inserted T4L into the 3rd intracellular loop and truncated c-terminal end of the protein. Jaakola et al, Science (2008), 322: 1211-1217. The A2A$denosine Receptor ZM241385 is an antagonist of the A2A receptor Jaakola et al, Science (2008), 322: 1211-1217. The CXCR4 Receptor Inserted T4L into the 3rd intracellular loop and a thermostable point mutant. Wu et al, Science (2010), 330: 1066-1071. The CXCR4 Receptor IT1t is a small molecule and CVX15 is a cyclic peptide that bind to CXCR4. Wu et al, Science (2010), 330: 1066-1071. The M2 Muscarnic Receptor Inserted T4L into the 3rd intracellular loop. M2 Muscarinic receptor- Haga et al, Nature(2012), 482: 547-552. The M2 Muscarinic Receptor QNB is a small molecule that binds selectively to the M2 receptor. M2 Muscarinic receptor- Haga et al, Nature(2012), 482: 547-552. The M3 Muscarinic Receptor Inserted T4L into the 3rd intracellular loop. M3 Muscarinic receptor- Kruse et al, Nature(2012), 482: 552-559. The M3 Muscarinic Receptor Tiotropium is a small molecule that binds selectively to the M3 receptor. M3 Muscarinic receptor- Kruse et al, Nature(2012), 482: 552-559. The Delta Opioid Receptor Inserted T4L into the 3rd intracellular loop. Delta-opioid receptor- Granier et al, Nature(2012), 485: 400-405. . The Delta Opioid Receptor Naltrindole is a small molecule that binds selectively to the Delta opiod receptor. Delta-opioid receptor- Granier et al, Nature(2012), 485: 400-405. . The Kappa Opioid Receptor Inserted T4L into the 3rd intracellular loop. Kappa-opioid receptor- Wu et al, Nature(2012), 485: 327-334. The Kappa Opioid Receptor JDTic is a small molecule that binds selectively to the Kappa opioid receptor. Kappa-opioid receptor- Wu et al, Nature(2012), 485: 327-334. The FQ Opioid Receptor Inserted the thermostabilize apocytrochrome b562RIL (BRIL) in place of 43 amino acids at the n-terminus and deleted 31 residues as the c-terminus. FQ-opioid receptor- Thompson et al, Nature(2012), 485: 395-400. The FQ Opioid Receptor C24 is a small molecule that binds selectively to the FQ receptor. FQ-opioid receptor- Thompson et al, Nature(2012), 485: 395-400. The µ Opioid Receptor Inserted T4L into the 3rd intracellular loop. µ$opioid receptor- Manglik et al, Nature(2012), 485: 321-327. The µ Opioid Receptor "-funaltrexamine ("-FNA)is a small molecule that binds selectively to the µ opioid receptor. µ$opioid receptor- Manglik et al, Nature(2012), 485: 321-327. Section #4: DNA as an allosteric ligand of proteins. Reference: S.H Meijsing et al., Science 324, 407-410 (2009). Glucocorticoid Receptor •! The glucocorticoid receptor (GR) is a potent transcriptional activator that contains a ligand-binding domain for dexamethasone. •! Ligand binding is required prior to binding DNA and inducing gene expression. •! It induces expression of many target genes whose by associating with specific DNA-binding sites, the sequences of which differ dramatically between genes. •! GR binds to palindromic and non-palindromic sequences as a dimer and binding to DNA induces dimerization. Schematic of the functional domains of GR •! GR is composed of three functional domains: Activation Function 1 (AF1: residues 1-406), DBD (residues 440-525) and LBD/AF2 (residues 550-770). •! The DNA-binding domain (DBD) contains two zinc fingers. Binding to DNA occurs subsequent to hormone binding. •!The DBD binding site consists of two hexameric half sites separated by 3- or 4-base pairs (imperfect palindromic sequences). GR target DNA sites (GBS) differ in sequence •! All sequences are derived from endogenous target genes. •! All sequences have comparable basal activity but differ considerably when induced with dexamethasone (dex). •! There is not a good correlation between in vitro binding and in vivo transcriptional activity. Mutations in GR domains effect GBS activity •! Mutations were introduced into the dimerization motif (red), the AF1 (yellow) and in the AF2 (green). •! The mutations had different effects on different GBS sites. •! This suggested that binding to different targets were causing different interactions at regions distant from the DNA-binding residues. The role of the lever arm in GR function •! In 13 crystal structures of GR-DBD:GBS complex, the structures were virtually identical except the lever arm. •! In one chain H472 was flipped in and in the other chain H472 was flipped out. •! In the flipped out chain, the conformation of the lever arm was heterogeneous and this suggests that the DNA sequence directs distinct changes in the conformation of the lever arm. The role of the lever arm in GR function Chain A versus chain B Chain B: 3 bp versus 4 bp spacer The role of the lever arm in GR function •! The GR% splice variant differs from GR! in the lever arm. •! The two isoforms display similar abilities to repress the osteocalcin gene . •! The two isoforms display varying abilities to activate genes depending on the target GBS sequence. The role of the lever arm in GR function •! The lever arm and interactions with residues in the core help to define the specificity of the steroid hormone receptors. DNA as an allosteric ligand of proteins Summary: •! The binding interface of the 13 GR:GBS complexes are very similar and suggest the different roles is not due to the specific interactions with DNA. •! The structures demonstrate the importance of the lever arm to the function of the GR and potentially to other steroid hormone receptor. •! The results suggest that binding DNA can alter the properties of the GR. Thus, DNA is acting as an allosteric ligand for the proteins functions.