* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Base excision repair
Promoter (genetics) wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
DNA sequencing wikipedia , lookup
Real-time polymerase chain reaction wikipedia , lookup
Agarose gel electrophoresis wikipedia , lookup
Restriction enzyme wikipedia , lookup
DNA profiling wikipedia , lookup
Genomic library wikipedia , lookup
Community fingerprinting wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
SNP genotyping wikipedia , lookup
Biosynthesis wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
Gel electrophoresis of nucleic acids wikipedia , lookup
DNA repair protein XRCC4 wikipedia , lookup
Non-coding DNA wikipedia , lookup
Molecular cloning wikipedia , lookup
Transformation (genetics) wikipedia , lookup
Point mutation wikipedia , lookup
DNA supercoil wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
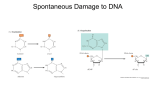
Mutations and DNA damage Mutations and DNA damage Mutations result from • • • • Changes in the nucleotide sequence of DNA Deletions Insertions Rearrangements of DNA sequences in the genome. These changes can be: • Spontaneous mutations that occurs as a result of natural processes in cells, for example DNA replication errors • Induced mutations Occur as a result of interaction of DNA with an outside agent that causes DNA damage (UV radiation, oxygen free radical, chemicals). Spontaneous mutations Point mutations The simplest type of mutation in which a nucleotide pair in a DNA duplex is replaced with a different nucleotide pair. Whether nucleotide substitutions have a phenotypic effect depends on where they are located in the DNA sequence. Transition mutations replace one pyrimidine base with another, or one purine base with another. Transversion mutations replace a pyrimidine with a purine or vice versa. Both mutations can be permanently incorporated by DNA replication. Point mutations: Silent and nonsense mutations Mutations that change the nucleotide sequence without changing the amino acid sequence are called synonymous mutations or silent mutations. Mutational changes in nucleotides that are outside of coding regions can also be silent. However, some noncoding sequences do have essential functions in gene regulation and, in this case, mutations in these sequences would have phenotypic effects. A nucleotide substitution that creates a new stop codon is called a nonsense mutation. It causes premature chain termination during protein synthesis. Nearly always a nonfunctional product. Point mutations: Missense mutations Nucleotide substitutions in protein-coding regions that result in changed amino acids are called nonsynonymous mutations or missense mutations. May alter the biological properties of the protein. The human hereditary disease sickle cell anemia is an A→T transversion which changes the codon for glutamic acid to valine in the -globin chain of the hemoglobin. It results in a decrease in the ability of red blood cells to carry oxygen throughout the body. Most people have only hemoglobin type A (Hb A) within RBC (normal genotype: Hb AA). In Sickle-cell anemia there is homozygosity for the mutation that causes HbSS. Point mutations: Insertions or deletions can cause frameshift mutations If the length of an insertion or deletion is not an exact multiple of three nucleotides, this results in a shift in the reading frame of the resulting mRNA. Usually leads to production of a nonfunctional protein. Example: human hereditary disease cystic fibrosis. The protein encoded by CFTR gene (cystic fibrosis transmembrane conductance regulator) is required to regulate the components of sweat, digestive fluids, and mucus. CFTR regulates the movement of chloride and sodium ions across epithelial membranes, such as the alveolar epithelia located in the lungs. The deletion of three base pairs in the nucleotide sequence of CFTR gene results in the loss of the codon for phenylalanine. CF develops when neither gene works normally (as a result of mutation) and therefore has autosomal recessive inheritance Induced mutations General classes of DNA damage Spontaneous damage to DNA can occur through the action of water in the aqueous environment of the cell. Degradation of DNA in water is due to DNA hydrolysis, that is, the breaking of DNA bonds through the addition of water. This type of damage occurs very rarely. A mutagen is any chemical agent that causes an increase in the rate of mutation above the spontaneous background. Three general classes of DNA damage • Single base changes • Structural distortion • DNA backbone damage Induced mutations Single base changes Single base changes: deamination A single base change or “conversion” affects the DNA sequence but has only a minor effect on overall structure. Deamination is the most frequent and important kind of hydrolytic damage, and can occur spontaneously from the action of water, or be induced by a chemical mutagen. The replacement of the amino group of cytosine with oxygen converts cytosine to uracil. When a UG base pair replaces a CG base pair this causes only a minor structural distortion in the DNA double helix. This type of damage is not likely to completely block the process of replication or transcription, but may lead to the production of a mutant RNA or protein product. Single base changes: deamination Vertebrate DNA frequently contains 5-methylcytosine in place of cytosine. Methylated cytosines are “hotspots” for spontaneous mutation in vertebrate DNA because deamination of 5-methylcytosine generates thymine. This results in the change of a GC base pair into an AT when damaged DNA is replicated Single base changes: alkylation Alkylation is the transfer of an alkyl group from one molecule to another. Alkyl groups range from single carbon compounds such as methyl groups to much longer chains of hydrocarbons. Alkylating agents such as nitrosamines lead to the formation of O6-methylguanosine. This modified base often mispairs with thymine. Can result in a GC→GT→AT point mutation after DNA replication. Single base changes: oxidation Oxidation is any chemical reaction in which a compound loses electrons. Potent oxidizing agents are generated by ionizing radiation and by chemical agents that generate free radicals. These reactive oxygen species (O2−, H2O2, and OH[·]) can generate 8oxoguanine (oxoG), a damaged guanine base containing an extra oxygen atom. OxoG can form a Hoogsteen base pair with adenine. Gives rise to a GC→TA transversion. One of the most common mutations found in human cancers. Induced mutations Structural distortion Structural distortion Thymine dimer UV radiation induces the formation of a cyclobutane ring between adjacent thymines, forming a T-T dimer. The T-T dimer distorts the double helix and can block transcription and replication. UV radiation can also induce dimers between cytosine and thymine. Structural distortion Structural distortion can be caused by intercalating agents and base analogs: Ethidium bromide has several flat polycyclic rings that insert between the DNA bases. 5-bromouracil, an analog of thymine, can mispair with guanine. Due to the resulting distortion of the double helix, intercalating agents can cause insertion or deletion of one or more base pairs during DNA replication. Induced mutations DNA backbone damage DNA backbone damage Formation of abasic sites Loss of the nitrogenous base from a nucleotide. Generated spontaneously by the formation of unstable base adducts. Double-stranded DNA breaks Induced by ionizing radiation (X-rays, radioactive materials) and a wide range of chemical compounds. Ionizing radiation can attack (ionize) the deoxyribose sugar in the DNA backbone directly or indirectly by generating reactive oxygen species. Double-strand breaks are the most severe type of DNA damage, since they disrupt both DNA strands. Cellular responses to DNA damage Damage bypass Damage reversal Damage removal Cellular responses to DNA damage Damage bypass Lesion bypass The normal replication machinery uses high-fidelity DNA polymerases. These high-fidelity polymerases accurately copy nondamaged template DNA, but are unable to bypass DNA lesions that cause structural distortion of the DNA helix Specialized low-fidelity, “error-prone” DNA polymerases transiently replace the replicative polymerases and copy past damaged DNA in a process called Translesion synthesis (TLS). Lesion bypass (1) The replication machinery is shown arrested (stop sign) at a thymine dimer on the template for leading strand synthesis. Multiple specialized errorprone DNA repair polymerases are stored in a subnuclear compartment near the arrested replication fork (DNA pol ι, λ, ζ, η, and κ). (2) When the replication fork stalls at the lesion, DNA pol δ is replaced by one of the specialized polymerases (DNA pol ι in this example). Lesion bypass (3) When translesion synthesis has bypassed the damage and extended to a suitable position downstream, another polymerase switch occurs. DNA pol ι is released and replaced by DNA pol δ. High-fidelity DNA replication continues Lesion bypass Error-prone DNA polymerases The error-prone DNA polymerases are able to copy damaged DNA templates permissively and efficiently. However, because they are error-prone they may generate mutations. Typical error rates range from 10-1 to 10-3 per base pair. Most lesions completely alter the pairing properties of the pre-existing bases. In this case, the polymerases May insert incorrect nucleotides opposite the lesion: nucleotide substitution May skip past and insert correct nucleotides opposite bases downstream: frameshift Lesion bypass An exception to the general tendency of repair polymerases to be error-prone is DNA polymerase eta (η) DNA polymerase eta () • Performs translesion synthesis past TT dimers by inserting AA. This results in the lesion being bypassed in an error-free manner • Has an extra wide active site that can accommodate two dNTPs instead of one. • Van der Waals forces and hydrogen-bonding interactions hold the TT dimer so that the two thymines can be paired with two adenines. • The T-T dimer is still present in the original parent strand of the DNA double helix after translesion synthesis Lesion bypass Translesion synthesis enables a cell to survive what might be a fatal block to replication, but with the risk of a higher mutation rate. Lesion bypass is not really a repair pathways; for example, the T-T dimer is still present in the original parent strand of the DNA double helix after translesion synthesis. What pathways are available to repair such damage???? Cellular responses to DNA damage Damage reversal Reversal of thymine-thymine dimers by DNA photolyase In most organisms, UV radiation damage to DNA can be directly repaired by a process called photoreactivation or “light repair”. DNA photolyase uses energy from near UV to blue light to break the covalent bonds holding two adjacent pyrimidines together. How does this take place????? Reversal of thymine-thymine dimers by DNA photolyase DNA photolyase has two cofactors: • A pigment that absorbs UV/blue light • Fully reduced flavin dinucleotide (FADH-) The TT dimer is flipped out of the DNA helix and brought very close to FADH-. An electron is transferred from FADH- and the dimer is split. Placental mammals, including humans, do not have a photoreactivation pathway!!!! Damage reversal by DNA methyltransferase Methyltransferase catalyzes the transfer of the methyl group on O6methylguanine to the sulfhydryl group of a cysteine residue on the enzyme. Methyltransferase are present in all organism DNA methyltransferase binds the minor groove of the DNA. The O6-methylguanine flips out from the double helix into the active site. A sulfhydryl group of a cysteine in the active site accepts the methyl group from guanine. Once the methyltransferase accepts the methyl group from guanine, the enzyme cannot be used again. Cellular responses to DNA damage Damage removal Repair systems that remove damaged DNA include: Repair of single base changes by two main pathways: • Base excision repair Involves the correction of single base changes that are due to conversion. • Mismatch repair Mismatched base pairs that result from DNA polymerase errors during replication are corrected by mismatch repair Repair of structural distortion • Nucleotide excision repair The nucleotide excision repair pathway is used for repair of structural distortion, for example bulges from thymine dimers induced by UV irradiation Repair of double-strand breaks • Repaired by homologous recombination or nonhomologous endjoining. Repair of single base changes Base excision repair Base excision repair Base excision repair is initiated by a group of enzymes called DNA glycosylases. These enzymes cleave the N-glycosidic bond connecting the deoxyribose sugar to the damaged base. There are DNA glycosylases that recognize oxidized/reduced bases, methylated bases, deaminated bases, or base mismatches. For example, uracil DNA glycosylase removes uracil from UA or UG base pairs, and the human oxoG repair enzyme, hOGG1, catalyzes excision of oxoG. The first step in base excision repair is recognition of the lesion How do DNA repair proteins find the rare sites of damage in a vast expanse of normal DNA is poorly understood overall. Base excision repair Model for DNA damage recognition by 8-oxoguanine DNA glycosylase 1 (hOGG1) DNA glycosylase 1 first binds nonspecifically to DNA. The damaged base goes through a series of “gates” or checkpoints within the enzyme. If the enzyme encounters a normal GC base pair, then: The G is transiently extruded into a G-specific pocket and returned to the double helix. If the enzyme encounters a oxoG-C base pair, then: The oxoG is extruded into the G-specific pocket and then inserted into a lesion recognition pocket where it is excised. Base excision repair The excision of the damaged base form an abasic site in the DNA Repair of deamination of cytosine to uracil The next step in repair is to remove the abasic nucleotide. In mammalian cells, the repair patch may be a single nucleotide (short patch) or 2–10 nucleotides in lenght (long patch). Base excision repair Short patch repair The enzymes glycosylase-associated β-lyase and APE1 (apurinic/apyrimidinic endonuclease) make nicks 3′ and 5′ to the abasic site in the DNA , respectively. DNA polymerase β replaces the missing nucleotide, and the DNA ligase 3–XRCC1 complex seals the gaps in the sugar– phosphate backbone. Base excision repair Long patch repair APE1 makes an incision 5′ to the abasic site. Then, DNA polymerase δ or ε, and PCNA, displace the strand 3′ to the nick to produce a flap of 2–10 nt. The flap is cut at the junction of the single to double strand transition by FEN1 (flap endonuclease-1). A patch of the same size is then synthesized by DNA polymerase δ or ε with the aid of PCNA and ligated by DNA ligase I. Repair of single base changes Mismatch repair Mismatch repair Mismatch repair corrects mistakes that occur during DNA replication and are not proofread by the DNA polymerase. Basic features of the mismatch repair pathway are conserved from E. coli to humans. Mismatch repair pathway in mammalian cells The first step in mismatch repair is the recognition of error by two proteins the MutS/MutL complex. Once the mismatch is recognized, a large region of DNA including the mismatch is excised. Mismatch repair One model proposes that MutS /MutL then diffuses for several thousand nucleotides either 5′ or 3′. • A 5′ or 3′ single-strand break is generated by EXO1 in association with PCNA (the replication clamp) and RFC (the clamp loader). • 5′→3′ or 3′→5′ progessive exonuclease activity of EXO1 removes the mismatch. Mismatch repair • 5′→3′ repair synthesis is mediated by DNA polymerase and associated factors. • Ligation of the remaining gap is catalyzed by DNA ligase I. Reconstitution of mismatch repair in an in vitro system showed that the recognition protein MutS, the exonuclease EXO1 and DNA polymerase are indispensible for repair. Repair of structural distortion Nucleotide excision repair Nucleotide excision repair Repair of structural distortion (e.g. bulges from thymine-thymine dimers induced by UV irradiation). Basic features of the nucleotide excision repair pathway are conserved from E. coli to humans. Xeroderma pigmentosum • Autosomal recessive disorder. • Photosensitivity. • Greatly increased risk of sunlightinduced skin cancer. • Neurological degeneration. • Defects in nucleotide excision repair or in T-T dimer translesion synthesis. Mammalian nucleotide excision repair pathway The repair pathway responsible for recognizing lesions in the whole genome is called global genome repair (GGR), while the transcription-coupled repair (TCR) pathway identifies lesions in the transcribed strand of active genes. In the first step, DNA damage is recognized by the cooperative binding of XPC, RPA, XPA, and TFIIH. • XPC binds first, followed by the other proteins (XPC, sliding along DNA molecule and jumping from one DNA molecule to another, is able to scan the genome rapidly and efficently) . • TFIIH is a multiprotein complex that also plays an important role in gene transcription. Mammalian nucleotide excision repair pathway Unwinding of the duplex DNA is promoted by the action of XPB and XPD helicases, which are subunits of the TFIIH complex. The endonuclease XPG makes a 3′ incision The endonuclease XPFERCC1 makes another 5′ incision approximately 20 nucleotides from the damage. Mammalian nucleotide excision repair pathway After the helicase activity of XPB and XPD unwinds the duplex DNA, the damage-containing DNA sequence (24–32 nt) is released. Repair synthesis is mediated by DNA polymerase or . Ligation of the remaining gap in the DNA backbone is mediated by DNA ligase I. Repair of double-strand breaks Homologous recombination Double-strand break repair by removal of DNA damage This type of DNA damage is the most harmful to cells and is often linked to cell death or cancer! For example, hereditary deficiencies in double-strand break repair are linked to an increased susceptibility to breast cancer. Double-strand breaks in DNA are induced by reactive oxygen species, ionizing radiation, and chemicals the generate reactive oxygen species (free radicals). Double-strand breaks in DNA are repaired by: Homologous recombination Repairs double-strand breaks by retrieving genetic information from an undamaged homologous chromosome. Nonhomologous end-joining (NHEJ) Rejoins double-strand breaks via direct ligation of the DNA ends without any requirement for sequence homology. Double-strand break repair by removal of DNA damage • Homologous recombination plays a major role in doublestrand break repair in prokaryotes and single-cell eukaryotes. • In mammalian cells, double-strand breaks are primarily repaired through NHEJ. • In mammalian cells, the main function of homologous recombination is to repair double-strand breaks at the replication fork Homologous recombination: repair of double-strand breaks in DNA Exposure of cells to ionizing radiation or other double strand break-inducing agents triggers an increase in ATM kinase activity (a serine–threonine kinase in the nucleus). A kinase is a type of enzyme that transfers phosphate groups from high-energy donor molecules, such as ATP, to specific substrates, a process referred to as phosphorylation. ATM is recruited to the break site and phosphorylates some of the proteins involved in DNA repair and cell cycle control. Humans that lack ATM suffer from a syndrome called ataxia telangiectasia, characterized by extreme sensitivity to radiation, increased susceptibility to developing cancer, immunodeficiency, premature aging, and neurodegenerative disorders. Model for mammalian DNA double-strand break repair by homologous recombination A double-strand break (DSB) is induced by ionizing radiation. The MRN (Mre11–Rad50–Nbs1) complex is rapidly recruited to the DSB site. Model for mammalian DNA double-strand break repair by homologous recombination MRN complexes form a bridge between free DNA ends via the coiled-coil arms of Rad50 dimers. Inactive ATM dimers are recruited to the DSBs through interaction with the carboxyl terminus of Nbs1, and by a less stable interaction with Rad50. Activating signals are delivered to ATM dimers, possibly through a conformational change in Nbs1. Model for mammalian DNA double-strand break repair by homologous recombination ATM undergoes phosphorylation accompanied by its conversion from a dimer to a monomer. Activated ATM monomers either remain near the DSB, where they phosphorylate proteins involved in DNA repair, or diffuse away from the DSB sites to phosphorylate nuclear substrates, such as p53 and Creb that are involved in cell cycle control. Model for mammalian DNA double-strand break repair by homologous recombination The 3′,5′-exonuclease activity of Mre11 generates 3′ single-strand DNA tails that are recognized by Rad52. Strand invasion of the 3′ tails with intact homologous sequences is initiated by Rad51. The proteins Rad54, Rad55, Rad57, BRCA1, and BRCA2 are also involved in homologous recombination, but their precise roles have yet to be determined. Model for mammalian DNA double-strand break repair by homologous recombination Strand exchange generates a hybrid molecule between damaged and undamaged duplex DNAs. Sequence information that is missing at the DSB site is restored by DNA synthesis (newly synthesized DNA is shown in red). The interlinked molecules are then processed by branch migration. Model for mammalian DNA double-strand break repair by homologous recombination Heteroduplex DNA: duplex DNA formed during recombination is composed of single DNA strands originally derived from different homologs. Holliday junction: an intermediate in which the two recombining duplexes are joined covalently by single crossovers. The Holliday junction is resolved into two duplexes by an enzyme complex called the resolvasome. The resolvasome has “resolvase” activity. Repair of double-strand breaks Nonhomologous end-joining Nonhomologous end-joining In mammals double-strand breaks in DNA are primarily repaired through nonhomologous end-joining. This is thought to be the major pathway for repair of double-strand breaks induced by ionizing radiation. Following a double-strand break, the broken ends of DNA are recognized by two heterodimers of the Ku70 and Ku80 proteins. The heterodimers form a scaffold that holds the broken ends in close proximity, allowing other enzymes to act. Nonhomologous end-joining The Ku heterodimer recruits the nuclease (Artemis/DNA-PKCS), the polymerases (DNA polymerases μ and λ), and the ligase complex (XRCC4– DNA ligase IV) to the damaged site. The endonuclease Artemis is activated after it is phosphorylated by the DNA-dependent protein kinase catalytic subunit (DNA-PKCS) Nonhomologous end-joining The activated Artemis/DNA-PKCS complex then trims excess or damaged DNA at the break site. DNA polymerases μ and λ, or the enzyme TdT are required for any nonhomologous end-joining events that need fill-in of gaps or extension of the 3′ or 5′ overhangs. The rejoining of the broken ends is carried out by DNA ligase IV in association with XRCC4. Nonhomologous end-joining • This double-strand break repair process can lead to mutation. • Two broken ends can be ligated together regardless of whether they came from the same chromosome. • Frequently results in insertions or deletions at the break site. • Trade-off between repair and otherwise lethal breaks in the genome.