* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Biological membranes, cell compartments
G protein–coupled receptor wikipedia , lookup
Gene expression wikipedia , lookup
Biosynthesis wikipedia , lookup
Biochemical cascade wikipedia , lookup
Magnesium transporter wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Mitochondrial replacement therapy wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Biochemistry wikipedia , lookup
Oxidative phosphorylation wikipedia , lookup
Lipid signaling wikipedia , lookup
Paracrine signalling wikipedia , lookup
Fatty acid metabolism wikipedia , lookup
Proteolysis wikipedia , lookup
Western blot wikipedia , lookup
Mitochondrion wikipedia , lookup
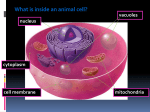
Biological membranes, cell compartments Bruno Sopko Content • Introduction • Plasma membrane – Lipid rafts • • • • • • • Nucleus Mitochondria Endoplasmic reticulum Golgi apparatus Lysosoms Peroxisoms Cytoplasm and cytoskeleton Introduction - Advantages of the compartmentalisation • Maintaining of the high local concentration of the chemical components in one compartment – higher reaction rates • Controlling the transport of intermediates among compartments is other effective regulation system of metabolic pathways, localized in more than one compartment • Protection from the environment • Protection from the “aggressive” compartment interior Introduction – subcellular compartments • • • • • • • • • • • Nucleus Nucleolus Golgi apparatus Mitochondrion Lysosoms Peroxisoms Cytoskeleton Cytosol Centriols Plasma membrane Endoplasmic reticulum Introduction – Transport across membrane • Diffusion • Assisted diffusion • Active transport Plasma membrane • Plasma membrane (plasmalemma) is biologic membrane separating inner parts of the cell from the environment. • Plasma membrane is the surface of all cells and is selectively-permeable. This is the regulation part for the intake and excretion of the chemical substances. • Composed of lipids (phospholipids, glycolipids and cholesterol esters) and proteins, participating in many cell processes as, for instance, cell adhesion, ion channels transport and cell signaling. • Inner part contains specific sites, binding components of intercellular cytoskeleton. • Organized in subcompartments – microdomains, rafts Plasma membrane - forming Lipid Rafts Lipid rafts are small (10-200nm), heterogeneous, highly dynamic, sterol- and sphingolipid-enriched domains that compartmentalize cellular processes. Lipid Rafts • Cholesterol and sphingolipid-enriched membrane microdomains or platforms – Cholesterol of double concentration – Sphingomyelin concentration elevated by 50% • Concentrate and segregate proteins within the membrane • More ordered and tightly packed than surrounding bilayer • Float freely in the surrounding membrane Types of lipid rafts • Caveolae: small, flask-shaped invaginations of the plasma membrane enriched in caveolin • Planar lipid rafts: found in neurons and enriched in flotillin • Caveolin and flotillin recruit signaling proteins • Signaling can be promoted or dampened Lipid Raft Proteins • “True resident proteins” – GPI-anchored proteins-prion protein (PrPc) – Caveolin – Flotillin • Signaling proteins – G-protein, non-receptor tyrosine kinases • Cytoskeletal/Adhesion proteins – actin, myosin, vinculin, cofilin, cadherin, ezrin GPI-anchored proteins Lipid Raft formation Lipid Rafts and Viruses • HIV virus – Budding may occur from lipid rafts • Influenza virus – Raft-associated glycoproteins in envelope Nucleus • DNA replication • Transcription, synthesis of mRNA, tRNA, rRNA, ribosomes • Surrounded by double membrane connected to ER • Transport of cytosolic compounds (active and passive) only via nuclear pores • Specialized subcompartments (nucleolus – ribosomes formation, DNA replication is localized etc.) Nucleus Nucleus – transport across nuclear envelope Nucleus –protein transfer Transfer into nucleus Transfer from nucleus Nucleus – RNA export Mitochondria • Two membranes – inner and outer are highly different in composition and enzymatic activity • Matrix of the mitochondria (mitosol) exhibits different biochemical functions • Majority of the mitochondrial proteins is coded in nucleolar DNA (and synthesized by free ribosomes in cytosol), but 13 mitochondrial proteins and some RNA are coded in circular mitochondrial DNA (mtDNA) Mitochondria • Synthesis of more then 90% ATP – oxidative phosphorylation (inner membrane) • Heat production • Apoptosis – inner mitochondrial pathway • Oxidative decarboxylation of pyruvate (pyruvate dehydrogenase complex) • ß-oxidation of fatty acids (shorter then 24-C) • Kreb’s cycle - mitosol • P450 - in inner membrane Mitochondria – protein import Mitochondria – elektron import from cytosolic NADH Michael W. King, Ph.D / IU School of Medicine / miking at iupui.edu Mitochondria – ATP export to cytosol Mitochondria – import of fatty acids for oxidation Mitochondria – acetyl-CoA export for synthesis of lipids and cholesterol Michael W. King, Ph.D / IU School of Medicine / miking at iupui.edu Mitochondria – the urea cycle • Only ornithin transcarbamoylase is located in mitosol • Carbamoylphosphate is synthetized in mitochondria from bicarbonate and ammonia, utilizing 2 ATP • Citrullin is exported via antiport with ornithine Endoplasmic reticulum • Proteosynthesis (rough ER = RER) • Primal steps of polysaccharides chains synthesis (N-bound glycoproteins (RER)) • Synthesis (smooth ER = SER) - phospholipids, triglycerides • Cholesterol and steroids (SER) synthesis (on the surface) • Hydroxylation of endogenous a exogenous compounds by cytochromes P450 (SER) • Calcium ions storage (SER , sarcoplasmic reticulum) • Connected with nucleolar envelope Golgi apparatus • Cooperates with endoplasmic reticulum • Enzymatic postranslation protein modifications (glycosylation, sulfatation) • Synthesis of new plasma membrane and participation in creation of primal lysosoms and peroxisoms Lysosoms • Intracellular digestion of intracellular, and extracellular compounds • Lysosomal enzymes are hydrolases with maximal activity at pH 5 (inside of lysosoms) • Hydrolysis of intracellular material – proteins nucleic acids, lipids together with organelles – autofagia • Extracellular material is hydrolyzed after the transport into the cell by endocytosis (pinocytosis and phagocytosis) heterofagia Lysosoms 1. Substances are in membrane enclosed vesicle 2. By fusion of the vesicle with primal lysosom secondary lysosom is formed 3. Lysosomal hydrolases digest the content of secondary lysosoms 4. Digested parts of the hydrolysate are transferred to cytosol for reuse 5. Non-digested material is accumulated in residual body and is excreted by exocytose 6. Remaining (nonexocytosed) residual bodies contain lipofuscin („age pigment“) Peroxisoms • Produce or use hydrogen peroxide • Differ in function and in number in different types of cell • More then 50 types of enzyme catalyzing oxidative and reductive biosynthetic reactions • oxidation of very long fatty acids chains (a- and boxidation) • synthesis glycerolipids, glycerol ether lipids (plasmalogens) and isoprenoids • enzymes for oxidation of D-aminoacids, 2-hydroxy acids and uric acid (uricase is absent at higher apes) • catalase • Known more then 25 perixosomes biogenesis disorders (Zellweger‘s syndrome = absence of peroxisomes and death occurs by age 6 months) • Some xenobiotics induce peroxisom proliferation Peroxisoms - -oxidation of long fatty acids Peroxisoms –plasmalogens synthesis Cytoplasm/cytoskeleton • Keeps phenotype (morphology) of the cell and participate in intracellular transport, cell mobility and cell division • Is formed by microtubules, intermedial filaments and actin filaments (microfilaments) Cytoplasm/cytoskeleton - microtubules • diameter 25 nm and length from 200 nm to 25 mm • Polymers of α- and βtubulin dimmers, polymerized end to end to protofilaments. Protofilaments form hollow microtubules Cytoplasm/cytoskeleton - intermedial filaments • diameter 10 nm • Domain structure of IF is conserved. Each protein contains non-a-helix domain at N and C-ends, which frame a-helix domain of the „stick“ • Basic unit of the intemedial filaments (IF) is dimmer • Known more then 70 gens for six basic types(I - VI) IF : – I a II - keratins (epithelial and higher) – III – e.g. desmin (sarcomers of muscle cells) and vimentin (e.g. fibroblasts - correct localization of organelles – V - nucleolar IF Cytoplasm/cytoskeleton - intermedial filaments Cytoplasm/cytoskeleton - microfilaments • The most dynamic part of cytoskeleton are microfilaments (actin filaments). • Diameter is 6nm • Formed by two linear polymers of actin subunits Cytoplasm/cytoskeleton - Centrosom and centriols • Centrosom is organell, main organisation centre of microtubules and regulates cell cycle • Centrosom is the cell region, forming microtubules • Centrioles are important parts of centrosomes. Cytoplasm/cytosol • • • • • • • • • • 54% cell volume Many reaction chains are located in parts of cytosol glycolysis (NAD+/ NADH) PPP - pentosophosphate pathway (NADPH) glycogenolysis glycogenesis (synthesa glykogenu) biosynthesis of fatty acids (fatty acid synthase) synthesis of active saccharides Proteosynthesis support Forming „subcompartments“ with different enzyme concentrations-> many reactions are carried out only in certain parts of cytosol Examples of pathways localized in a single compartment only • • • • Kreb’s cycle Glycolysis Proteosynthesis DNA replication Literature • • • • • • • • Gabriel Schlenstedt, Protein import into the nucleus, FEBS Letters 389 (1996) 7579 Walter Nickel and Catherine Rabouille, Mechanisms of regulated unconventional protein secretion, NATURE Reviews, Molecular cell Biology volume 10, february 2009 Marks´ Basic Medical Biochemistry, A Clinical Approach, third edition, 2009 (M. Lieberman, A.D. Marks) David A. Jans, Chong-Yun Xiao, and Mark H.C. Lam, Nuclear targeting signal recognition: a key control point in nuclear transport?, BioEssays 22:532-544, 2000 John Wiley & Sons, Inc The Cell: A Molecular Approach, Fourth Edition, GEOFFREY M. COOPER , ROBERT E. HAUSMAN, 2007, ASM Press, Washington, D.C. USA Yoshihiro Yoneda, How Proteins Are Transported from Cytoplasm to the Nucleus, J. Biochem. 121, 811-817 (1997) Alwin Köhler and Ed Hurt, Exporting RNA from the nucleus to the cytoplasm, NATURE Reviews, Molecular cell Biology volume 8, october 2007