* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Biomolecules and Nanotechnology
Survey
Document related concepts
Cell nucleus wikipedia , lookup
Magnesium transporter wikipedia , lookup
Endomembrane system wikipedia , lookup
Multi-state modeling of biomolecules wikipedia , lookup
Protein (nutrient) wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Protein phosphorylation wikipedia , lookup
Circular dichroism wikipedia , lookup
Protein moonlighting wikipedia , lookup
Signal transduction wikipedia , lookup
Protein structure prediction wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Intrinsically disordered proteins wikipedia , lookup
Chemical biology wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Transcript
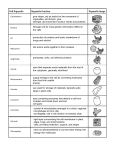
A reprint from American Scientist the magazine of Sigma Xi, The Scientific Research Society This reprint is provided for personal and noncommercial use. For any other use, please send a request to Permissions, American Scientist, P.O. Box 13975, Research Triangle Park, NC, 27709, U.S.A., or by electronic mail to [email protected]. ©Sigma Xi, The Scientific Research Society and other rightsholders Biomolecules and Nanotechnology Evolution has forced innovative solutions to biomolecular problems. Some may inform the growing field of nanotechnology David S. Goodsell T he term “nanotechnology” commonly refers to a speculative field that proposes to build machinery so small its components are measured on a scale of billionths of a meter (nanometers) using many of the principles of macroscopic engineering. In his books, K. Eric Drexler has popularized the design and computer modeling of many of these machines, including nano-scale manipulators to build objects atom by atom, bearings and axles built of diamond-like lattices of carbon, waterwheel-like pumps to extract and purify molecules and tiny computers with moving parts whose size is within atomic scale. The goals of these compelling machines are precision, with every structure and action controlled at the level of individual atoms, and parsimony, performing tasks at the minimum size necessary. You might be surprised to learn that nanotechnology was perfected more than three billion years ago. Indeed, working examples of each of these machines exist today within living cells. Nanoscale manipulators for building molecule-sized objects were discovered by the earliest cells and are now used to build proteins and other molecules atom by atom according to defined instructions. Rotating bearings are found in many forms: Clamps that encircle DNA David S. Goodsell is an assistant professor in the Department of Molecular Biology at the Scripps Research Institute. Trained in x-ray crystallography, he now splits his time between computer-aided drug design and research into the basic principles of macromolecular structure and function. His illustrated books The Machinery of Life and Our Molecular Nature (SpringerVerlag, New York) explore biological molecules and their diverse roles within living cells. Address: The Scripps Research Institute, Department of Molecular Biology, 10550 North Torrey Pines Road, La Jolla, CA 92037. Internet: http://www.scripps.edu/pub/goodsell 230 American Scientist, Volume 88 and slide along its length may be found in the simplest bacteria. Our own cells contain a rotary motor used not to power motion but instead to generate energy. Cells use a large collection of moleculeselective pumps to import ions, amino acids, sugars, vitamins and all of the other nutrients needed for living. Cells also use molecular computers, which, by altering their shapes, “read” the concentration of surrounding molecules and compute the proper functional outcome. By evolutionary search and modification over trillions of generations, living organisms have perfected a plethora of molecular machines, structures and processes. Figure 2 presents a few examples of the rich bio-nanotechnology that may be found in every modern cell. Biological molecules are proven examples of the feasibility, and the utility, of nanotechnology. Our lives depend on them. They are foreign, however, to our everyday experience, with unusual organic shapes and unfamiliar properties. Bio-nanomachines are often the same size and complexity as the speculative nanomachines being designed today, but they bear little resemblance to the machinery of our macroscopic world. Eric Drexler’s nanomanipulators and gears seem more familiar, because they are built by engineers along the familiar rigid, rectilinear designs of our macroscopic world. To understand the organic, flexible forms of bio-nanomachines, we must forget the processes of design and engineering in our familiar world and look instead at the forces that shaped the evolution of life. Evolutionary Legacy The process of evolution by natural selection places strong constraints on the form that biological molecules may adopt. Because genetic information is passed directly from generation to generation, cells must maintain a living line back to the earliest primordial cells. If a cell fails to generate a living descendent, all of its biological discoveries will be lost. This is far more limiting than the technology of our familiar world. If we create machines that don’t function, we scrap them and go back to the drawing board. But if a cell takes a gamble and changes a critical machine, it had better get it right the first time or the result will be disastrous. The picture is not entirely grim, however, as cells have several levels of redundancy within which to develop new Figure 1. Biomolecular machines are comparable in size and complexity to the engineeringinspired nanomachines currently being proposed, but the forms and characteristics of the two are entirely different. Opposite are two solutions to atom-level synthesis. Model nanomanipulators (top right) build their products one atom at a time, like robots on an automobile assembly line. The biological approach to protein synthesis, at bottom, is less rigid. Many soluble machines read information from a strand of RNA, working together to build a new protein one piece at a time. Both of these illustrations are drawn at the same scale, so the sizes and shapes of the machines may be directly compared. Individual atoms are about the size of a grain of salt. Above are two approaches to a rotary bearing, drawn at higher magnification to show individual atoms. The model bearing at left is a perfectly symmetric arrangement of carbon in a diamondoid lattice. The bacterial sliding clamp, by comparison, is more organic in shape, composed of two C-shaped globular protein chains. Atomic coordinates were taken from file 2pol in the Protein Data Bank. (All images were prepared by the author.) © 2000 Sigma Xi, The Scientific Research Society. Reproduction with permission only. Contact [email protected]. © 2000 Sigma Xi, The Scientific Research Society. Reproduction with permission only. Contact [email protected]. 2000 May–June 231 information storage and retrieval (DNA, DNA polymerase, RNA) powered motion (flagellar motor) warfare (cholera toxin) chemical catalysis (enzymes) novel materials (elastin) signal transduction (hormone receptor) information-driven synthesis (ribosome) electrical insulation (myelin) containment (lipid bilayer) infrastructure (actin filament) molecular recognition (antibody) light emission (luciferase) electrical switching (voltage-sensitive channel) light sensing (opsin) photosynthesis (reaction center) packaging and delivery (rhinovirus) Figure 2. Using a few basic structural plans, cells have developed molecular nanomachines to fill every need. machines. First, the plans for a given machine may be duplicated, which allows the duplicate to be modified and ultimately perfected to perform a function different from the original. This is very common in the evolution of life. Hemoglobin, the protein that carries oxygen in our blood, is an example. Our cells contain information for building several different types of hemoglobin. One is optimized for carrying oxygen in the blood of adults, whereas another is found in the blood of a fetus. The fetal hemoglobin has a higher affinity for oxygen, allowing it to capture oxygen from the mother’s blood. About 200 million years ago, a gene duplication allowed the fetal hemoglobin to be perfected separately. Second, biology seldom involves a single cell. A population of cells—billions, trillions—is the biologically relevant entity. Within this population there exists ample room for experimentation. Millions of modifications may be tried, even if most are ultimately lethal. The population will still survive and individuals with rare improvements may grow to dominate in later generations. Human immunodeficiency virus (HIV) shows the benefits of evolutionary 232 American Scientist, Volume 88 small charged polar hydrophobic bulky Figure 3. Most of the cell’s molecular machinery is built of protein. Proteins are synthesized as a linear chain of amino acids. The 20 biological amino acids have diverse properties, some of which are indicated here, which allow the creation of proteins with widely differing properties. If properly designed, with the appropriate combination of charged and hydrophobic amino acids , the protein chain will fold into a compact globule when placed in water. change, accelerated so that we can see the effects in months instead of millennia. HIV reverse transcriptase, the enzyme that copies the virus’s genetic information, is particularly error-prone. Because of this, the population of viruses within an infected individual contains viruses with all possible single-site mutations—thousands of variants on the wild-type virus. The best of these will dominate, but even the weakest are continually created and recreated in subsequent generations by the low-fidelity copying mechanism. Thus, when an infected individual is treated with anti-HIV drugs, the population has a wide range of different mutants to choose from, some of which may be resistant to the drug: The virus is made more efficient by its very inefficiency. The hallmark of biological evolution is the plasticity provided by mutation and genetic recombination. Within a population, or through genetic duplication within a single cell, a great many variants may be tested and the occasional improvement saved. Evolution carries with it one important drawback, however: the problem of legacy. Once a key piece of machin- © 2000 Sigma Xi, The Scientific Research Society. Reproduction with permission only. Contact [email protected]. ery is perfected, it is difficult to replace it or make major modifications without killing the cell. This is particularly true for major molecular processes, such as protein synthesis, energy production and reproduction, which require the concerted action of many different molecular machines. This leads to the remarkable uniformity of all earthly living things when observed at the molecular level. All are built of the same basic components. Modern Molecular Machinery As a consequence of the evolution of life from a single primordial cell, all known living things on earth share a common molecular plan. All living things are made of four basic molecular building blocks: protein, nucleic acid, polysaccharide and lipid. Other small molecules are specially synthesized for specific functions, but the everyday work of the cell is performed by the four basics. The earliest cells chose these materials to the exclusion of others, and subsequent generations of cells, right up to our own, have been forced to work with them. Two different approaches are taken to synthesize these molecules, resulting in characteristic forms and functions. Proteins and nucleic acids are built in modular form by stringing subunits together based on genetic information. Proteins and nucleic acids may be built in any size and with subunits in any order. This gives remarkable flexibility to the form and function of these molecules. In contrast, lipids and polysaccharides are built by dedicated machines. Each new type of lipid molecule requires the creation of an entirely new suite of synthetic machines. Likewise, a new suite of machines must be created to build each new type of polysaccharide linkage. The result is that lipids and polysaccharides appear in fewer forms than proteins and are used in much more limited, albeit essential, roles. Our distant relatives developed a standard for biological information, choosing a particular 20 amino acids to be used in proteins, encoded by five types of nucleotides found in the nucleic acids DNA and RNA. Today, every protein is made of these 20 amino acids (at least initially). In their defense, these primordial cells chose an excellent set of building blocks, including flexible and rigid components, charged, uncharged, acidic, basic and neutral amino acids, large and small amino acids, and several with attractive chemically reactive properties (Figure 3). The amino acids may be used to create proteins with a wide range of properties. These include very flexible proteins with changeable shapes and very rigid crosslinked proteins designed to retain their shape under harsh conditions. Other proteins are highly basic or highly acidic, designed to perform their jobs under extreme acidic or alkaline conditions. Some are covered with carbonrich groups that repel water and seek out membranes for binding; others have polar surfaces and perform their duties in the watery cytoplasm. Modular synthesis allows proteins to be built in many shapes and sizes. As a consequence, most of the processes of modern cells are performed by proteins. Evolutionary legacy, however, places several limits on the design of proteins. As noted above, proteins are limited to the 20 components encoded in the DNA genome. Evolution also limits the size of proteins, limits them to aqueous environments and requires that they automatically assemble themselves within the crowded confines of the cell. In spite of these limitations, the breadth of protein form and function in modern cells is remarkable. The size of a protein is limited by the error rate of the protein-synthesis machinery, which in theory could produce a protein of any length. Missense errors, which misread the genetic information and substitute an incorrect amino acid at one position, occur at an average frequency of about 1 in 2,000. For a protein composed of 500 amino acids, one out of four proteins will typically have an error, but nearly every protein of 2,000 amino acids will have one. More important, however, are processivity errors, which cause protein synthesis to abort prematurely. These errors have been estimated to occur at a rate of about 1 in 3,000, so long proteins of several thousand amino acids are only rarely constructed in full. The average size of a typical protein chain, 300 to 500 amino acids, is the compromise adopted by most cells. Error rates keep the chain length low, so larger proteins must be built as complexes of multiple protein chains. Figure 4. Molecules must perform their tasks under the cell’s very crowded conditions. They must seek out their proper substrates and interact only with the proper partners. This cross section through the cytoplasm of a typical human cell depicts such macromolecules. The large purple molecules are ribosomes that are reading genetic information from the snaky white messenger RNA molecules. At the same time, orange, L-shaped transfer RNA molecules are aligned on the ribosome in order to build a new protein. Actin filaments and intermediate filaments criss-cross the space, providing support to the cell and a scaffold for hundreds of different enzymes, which are working on their metabolic tasks. The spaces between the macromolecules are also filled with small molecules and water, not depicted here for clarity, forming a very busy environment. Note that this picture is a static snapshot of the cell. In reality, all of these components are in rapid motion. © 2000 Sigma Xi, The Scientific Research Society. Reproduction with permission only. Contact [email protected]. 2000 May–June 233 Figure 5. Contact sites between proteins are highly specific, ensuring that a protein interacts only with its proper partners. Enolase, an enzyme involved in the breakdown of sugar, is shown here. Active enolase is formed from two identical subunits held together by an extensive protein-protein interface. As with most proteins composed of several subunits, enolase relies on complementary shape and a collection of many weak interactions to ensure specificity. In the upper illustration, the two subunits, colored green and blue, wrap arms around one another, with each arm fitting into a complementary groove. In the lower illustration, the two subunits have been separated slightly, and lines have been drawn to connect atoms that form hydrogen bonds across the interface. Atomic coordinates were taken from the file 4enl at the Protein Data Bank. Proteins were invented in “warm, salty pools,” so life on earth now requires a warm, aqueous environment (either externally or carried around inside). Water is essential for protein structure and function because of an emergent property of water solutions, termed the hydrophobic effect. Water has peculiar properties, which are used to great advantage by biological molecules. Portions of a protein that are rich in carbon interact weakly with water and are termed hydrophobic. When placed in solution, these hydrophobic regions crowd together in a globule, minimizing contact with water and allowing the water to escape and interact with more favorable environments. The hydrophobic effect is a major stabilizing force for protein folding, where carbon-rich portions of the chain are folded within the protein globule (as well as for formation of the lipid membranes that surround every cell, where the carbonrich portions of the lipids are packed inside the membrane). Because our molecules rely on hydrophobicity for their structural integrity, we could never live in vacuum or in organic solvents. Our proteins simply would not fold. Perhaps the most difficult limitation to overcome is the need for self-assembly. Biological molecules are designed to assemble themselves within cells: Proteins are created as unstructured, linear chains of amino acids that must fold into a stable, functional conformation (sometimes with a little chaperoning in the proper direction). Often, the folded chain spontaneously associates with others to form larger stable complexes. This is a major limitation to the design of proteins: Not only must the protein be functional in its active conformation, but the protein chain must also be designed to fold into this active conformation using only the folding tools available in the cell. Biomolecular Self-Assembly The forces involved in biomolecular structure and interaction are different from those at play in the macroscopic world, and thus our intuition may play us false when attempting to understand protein self-assembly. In our macroscopic world, much of engineering is based on the effect of gravity on solid objects. The strength of concrete and steel and the different frictional properties of Teflon and rubber are familiar quantities. The molecular world, on the other hand, is dominated by the effect of thermal motion on the atomic interactions within and between molecules. Molecules are endowed with kinetic energy proportional to the temperature, which manifests itself as translational, rotational and vibrational motion. The forces holding molecules together are continually fighting against these motions and are often overcome by them. The cellular environment is unusual in another respect, as shown in Figure 4. Figure 6. Closed point-group symmetry combined with a subunit of a given size allow cells to create objects of a defined size. These symmetrical complexes are used for measuring nanoscale distances, surrounding molecular targets or enclosing defined volumes. DNA-binding proteins such as CAP (left, with DNA in gray) use twofold symmetry to create allosteric “calipers” that measure the repeat length of DNA. Higher cyclic symmetries are used to make channels and cavities, such as the GroEL chaperonin (center), which uses sevenfold symmetry to create an enclosed environment within which new proteins are folded. Cubic symmetries are used to create larger containers. Ferritin (right) uses octahedral symmetry to create a container for iron ions, and viruses (see Figure 7) build large icosahedral capsids to enclose and deliver their genetic material to new host cells. Atomic coordinates were taken from files 1cgp, 1der and 1hrs at the Protein Data Bank. 234 American Scientist, Volume 88 © 2000 Sigma Xi, The Scientific Research Society. Reproduction with permission only. Contact [email protected]. Figure 7. Quasisymmetry allows viruses to build structures that are larger than is possible with perfect symmetry. At top is satellite tobacco necrosis virus, which shows perfect icosahedral symmetry. One subunit is highlighted. Exactly 60 subunits, all with identical structures, make up the capsid. At bottom is the tomato bushy stunt virus, which comprises 180 subunits and consequently is larger. In this virus, the symmetry is not perfect, and the subunits fall into three classes (red, orange and yellow), each with slightly different structures. Atomic coordinates were taken from the files 2tbv and 2stv at the Protein Data Bank. Proteins are synthesized in cells and left to float freely, diffusing to their ultimate site of action amid a crowded collection of competitors. Thus, a typical protein will come into contact with thousands of other types of proteins and must be able to discriminate its unique target from all others. This is quite different from the macroscopic world, where an engineer can selectively fit two parts together. For instance, the concept of a #6 screw would never work inside the cell. When building a chair, we are able to use the same screw to fasten many different pieces together, because we actively choose where each goes. In the cell, however, each molecule must be designed with a unique fastener, ensuring that it binds only to its proper target and no other. Before atomic structures of proteins were known, physicist H. R. Crane provided two design concepts that are required for biological self-assembly. First, “for a high degree of specificity the contact or combining spots on the two particles must be multiple and weak.” An array of many weak interactions, such that all are needed to provide the necessary stability, will form a specific site for interaction. If only a few very strong interactions are used, there is an increased chance that a protein will find a similar interaction with improper proteins. Second, “one particle must have a geometrical arrangement which is complementary to the arrangement on the other.” In other words, the shape of the interacting surfaces must form a good fit, and this fit must be different from that with other proteins. Specificity is provided by the complementary shape of the interacting surfaces, fitting knobs into holes, and by the complementary arrangement of hydrogen-bonding groups and charge-charge pairs. These two principles—that protein-protein interfaces are extended, with many weak interactions, and that protein-protein interfaces are complementary—have been proved in numerous protein structures, such as the one shown in Figure 5. Symmetry of Proteins The process of evolutionary selection has yielded an unusual result: Evolution of proteins favors perfect symmetry. The majority of soluble and membrane-bound proteins found in cells are symmetrical complexes formed by several subunits. Most proteins are oligomeric, composed of multiple copies of one or more types of subunits. Nearly all of these oligomeric proteins are also beautifully symmetrical, with identical subunits packed in identical environments. A complex interplay of conflicting functional needs has driven evolution to this surprisingly aesthetic conclusion. The major evolutionary force is the need for large proteins. Large proteins are preferred over smaller proteins and peptides for several reasons. Some functional roles simply require a molecule that is physically big. Large protein complexes form structural elements that span entire cells; they form rings that encircle DNA and rulers that measure lengths of DNA; they create pores of many sizes through cell membranes; they form large spherical containers for storage and delivery and small cylindrical containers that create exactly the proper environment for protein folding. Large proteins are also well suited for cooperative functions, such as allostery Figure 8. Allosteric motion of entire subunits within a complex is often used to regulate an enzyme’s activity. The active form of fructose-1,6-bisphosphatase, an enzyme involved in sugar metabolism, is a flat complex of four subunits, shown in profile on top. The binding of AMP (green, lower image) causes a scissor-like opening of the complex, inactivating the enzyme. Atomic coordinates were taken from files 4fbp and 5fbp at the Protein Data Bank. (discussed below) and multivalent binding, which require a molecule with several identical active sites. Multivalent binding increases the binding strength of a molecule to a target by reduction of entropy. Once one site on the protein has bound, the other sites are held in close proximity to the target, increasing their probability of binding. Many of the molecules of the immune system have a distinctive shape, composed of many flexible arms, in order to take advantage of this cooperativity. Large proteins also have attractive physicochemical characteristics. They are more stable against denaturation, having a more stabilized internal structure than small proteins. Large © 2000 Sigma Xi, The Scientific Research Society. Reproduction with permission only. Contact [email protected]. 2000 May–June 235 Figure 9. Immune system is designed for recognition of molecules on surfaces. Soluble proteins of the immune system seek out foreign molecules on the surfaces of viruses, bacteria and cancer cells, and cells of the immune system pass messages by contact of proteins on their surfaces. To perform these functions, many molecules of the immune system share a similar molecular plan, with globular binding domains, which seek out foreign molecules or pass messages, connected by flexible linkers, which allow more latitude when searching for the target. Antibodies and protein C1 from the complement system (top) are soluble proteins with several flexible arms, allowing them to explore surfaces of invading organisms. Cell-bound molecules, such as multiple histocompatibility complex and CD molecules (bottom), have only a single binding site and are connected to cells with a flexible linker. proteins also have a lower ratio of surface area to volume, making them less prone to damage and degradation by other enzymes. Unfortunately, the accuracy of the protein-synthetic machinery limits the size of proteins that may be constructed. As noted above, protein chains of 300 to 500 amino acids may be consistently synthesized, but longer chains will become increasingly riddled with errors. The answer is to build a complex from subunits when a large protein is needed, which allows any faulty subunits to be discarded. This also allows new possibilities for regulation: Large structures may be built and disassembled at will, or subunits may be transported to a distant site (or even outside the cell) and assembled there. Nearly all of these oligomeric proteins in cells form closed, symmetrical complexes based on ideal point-symmetry groups. In general, if a complex contains several identical subunits, 236 American Scientist, Volume 88 they will adopt identical symmetrical positions in the complex. Asymmetric complexes and random aggregates are almost completely unknown. Symmetrical association is favored over asymmetric association because it provides stability and control. The stability of closed, symmetrical complexes is a consequence of two factors. First, interfaces between proteins are highly specific and highly directional, so in most cases evolution selects and improves only a single type of association between subunits. Second, given these specific, directional interfaces, the maximum number of intersubunit contacts is formed by closed complexes. Closed, symmetrical complexes also ensure that the level of oligomerization is tightly controlled. Unwanted aggregation is very dangerous for cells— pathological aggregation of mutant proteins leads to diseases such as sickle-cell anemia, Alzheimer’s disease and prionrelated diseases. Selection of a closed, symmetrical complex defines the size and shape of the resultant complex. Under special circumstances, symmetry may be broken for a given functional need. For instance, viruses often need to build shells that are too large to construct with typically sized proteins in perfect symmetry—the highest point-group symmetry is icosahedral, so the largest perfectly symmetrical capsid is limited to 60 subunits. If larger shells are needed, more subunits must be used. Viruses often turn to quasisymmetrical complexes, where hundreds to thousands of identical subunits combine in similar, but not perfectly symmetrical, positions. Quasisymmetry was first conceived as a method of tiling an icosahedron with a triangular network, much like the geodesic domes designed by Buckminster Fuller. Protein subunits are arranged in this triangular lattice. Small elastic deformations allow the subunits to adopt similar contacts in each of the different positions. A series of different networks can be defined containing 60T subunits, where T is a “triangulation number.” Only certain triangulation numbers yielded smooth networks, according to the relation T = h2 + hk + k2, where h and k are integers. When structures were obtained for viral capsids, this model for quasiequivalence was surprisingly successful. The arrangement of subunits of most capsids corresponded closely to one of the triangulation numbers: Examples with T = 1 (perfect icosahedral) symmetry and T = 3 symmetry are shown in Figure 7. However, elastic deformations were not observed. Instead, subunits typically accommodated different positions through the use of structural “switches,” where the subunit adopts two or more significantly different conformations. Often, the subunits are composed of two domains connected by a flexible linker, and flexure of the subunit is used to adopt different conformations. Biomolecular Flexibility and Dynamics Engineers in our macroscopic world typically build rigid structures that stoically resist the forces of nature. Nature, however, has taken a different approach, developing machines that flex over the course of their action. Is a totally rigid nanostructure needed or even desired? Apparently not. In fact, biological molecules take advantage of flexibility for many aspects of their function. Many of these functions © 2000 Sigma Xi, The Scientific Research Society. Reproduction with permission only. Contact [email protected]. would be severely compromised, or not even possible, given a rigid molecule. Subtle motions can have surprisingly large effects on reaction rates or assembly. Biological molecules are perfectly placed to take advantage of these subtle motions. The step-by-step optimization provided by evolution allows a moderately active protein to be improved, through small changes modifying structure and flexibility, to yield a machine ideally tailored to fulfill its function. This process is easy for evolution but far more difficult for biotechnological design. We design our machines in one step, instead of through many small random optimization steps, and we expect to get it right with a minimum of tweaking and redesign. Thus, to anticipate all of the subtle effects of motion, our design techniques must be accurate enough to predict conformation and flexibility of molecules at scales far smaller than the radius of an atom. All biological molecules are flexible to some extent and are battered into different conformations by the constant pressure of surrounding water and the kinetic energy of their own atoms. At physiological temperatures, biological molecules constantly flex. Most of the interactions holding a protein together are conserved—covalent bonds remain connected, hydrogen bonds and salt bridges link portions of the chain—but entire elements of secondary structure flex, bending slightly or separating momentarily from the globule. These motions are often termed “breathing.” Breathing is essential in the function of myoglobin, a deep red protein that stores oxygen in muscle cells. Oxygen is bound to myoglobin in a pocket that is completely buried within the protein. Looking at the static structures provided by x-ray crystallography, there are no channels leading into or out of the pocket. For the oxygen to enter and exit, the molecule must breathe, transiently forming channels that allow passage. Many proteins use a carefully designed change of shape to regulate their action. These allosteric (“other shape”) proteins are composed of several subunits, each of which performs identical functions. In the simplest model of their action, each subunit may adopt two conformations, one functionally active, the other less active. Regulation is performed by propagation of the shape change from one subunit to its neighbors. For instance, phosphofructoki- nase, a key enzyme in sugar metabolism, uses allosteric regulation to modify its action. Phosphofructokinase is composed of four identical subunits (a tetramer), each containing a reactive site for the sugar molecules. The tetramer also contains binding sites for the energy molecule adenosine triphosphate (ATP) in the cleft between subunits. When ATP binds to this second site, it forces the entire enzyme complex into a different shape, which is less active than the original form. In the cell, this regulation is used as a negative-feedback loop. ATP is one of the final products of the sugar-breaking process that the enzyme performs. When ATP is plentiful, it binds to the regulatory site in phosphofructokinase, shutting down its own synthesis. The enzyme that performs the opposite reaction, shown in Figure 8, is also allosterically regulated. Many protein chains rely on “induced fit” to mediate their function. The chain may remain in a partially unfolded conformation that only completely folds when it binds to its target. Induced fit may be used to create doorways that allow ligands to enter protein cavities that are shielded from the surrounding environment. HIV-1 protease is an example. The active site is a cylindrical tunnel, with the cleavage machinery at its center. Somehow, a polypeptide must be threaded through this tunnel in order for the cleavage reaction to occur. This problem is solved through the use of two flexible flaps that cover the top of the tunnel. When free in solution, these flaps are disordered, opening a path to the active site. When the protease wraps around its target, the flaps close, forming a stable structure that positions the polypeptide accurately for cleavage. Flexible linkages are common in the molecular world. Protein chains may be made more flexible through addition of many molecules of the amino acid glycine, which are less hindered in bond rotation because of the lack of a side chain, or through addition of many charged residues, which favor exposure to solvent over forming a compact globule. The rigid kink formed by proline, surprisingly, is also commonly found in flexible regions, because it does not fit comfortably within compactly folded structures. The immune system contains many examples of flexible linkages that enhance multivalent binding, as shown in Figure 9. Prospects Biological molecules are examples of solved problems in nanotechnology— lessons from nature that may be used to inform our own design of nanoscale machines. The entire discipline of biotechnology has emerged to harvest this rich field of biological wealth. We routinely edit and rewrite the information in DNA to build custom proteins tailored for a given need. Today, for instance, bacteria are engineered to produce hormones, genes for disease resistance are added to agricultural plants, and cells are cultured into artificial tissues. Principles of protein structure and function also yield insights for nanotechnological design and fabrication. The diversity of protein structure and function shows the power of modular, information-driven synthesis, as well as the limitations imposed by modular design once a dedicated modular plan is chosen. Proteins demonstrate that extended, complementary interfaces are essential prerequisites for molecular self-assembly. The prevalence of protein complexes proves that error-prone synthesis may be accommodated through the use of subunits and symmetry to build large objects accurately and economically. And contrary to our macroscopic experience, motion and flexibility may be assets, not liabilities. The principles observed in the mobile, organic shapes of biological molecules may be applied to the controlled rectilinear forms of diamondoid lattices, fullerines or whatever nanoscale primitives are ultimately successful. We must not be too impatient, however. Nature has had some three or four billion years to perfect her machinery; so far, we have had only a few decades. Bibliography Crane, H. R. 1950. Principles and problems of biological growth. Scientific Monthly 70:376–389. Drexler, K. E. 1992. Nanosystems, Molecular Machinery, Manufacturing and Computation. New York: John Wiley & Sons. Goodsell, D. S. 1996. Our Molecular Nature: The Body’s Motors, Machines and Messages. New York: Springer-Verlag. Goodsell, D. S., and A. J. Olson. 1993. Soluble proteins: Size, shape and function. Trends in Biochemical Sciences 18:65–68. Goodsell, D. S., and A. J. Olson. 2000. Structural symmetry and protein function. Annual Reviews in Biophysics and Biomolecular Structure 29: 105–153. Larsen, T. A., A. J. Olson and D. S. Goodsell. 1998. Morphology of protein–protein interfaces. Structure 6:421–427. Protein Data Bank is available on-line at http://www.rcsb.org/pdb © 2000 Sigma Xi, The Scientific Research Society. Reproduction with permission only. Contact [email protected]. 2000 May–June 237