* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Minireview Shifty Ciliates: Frequent Programmed
Neuronal ceroid lipofuscinosis wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Minimal genome wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Gene expression programming wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Gene nomenclature wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Genome evolution wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Microevolution wikipedia , lookup
Genome (book) wikipedia , lookup
Protein moonlighting wikipedia , lookup
Designer baby wikipedia , lookup
Messenger RNA wikipedia , lookup
Helitron (biology) wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Transfer RNA wikipedia , lookup
Point mutation wikipedia , lookup
Gene expression profiling wikipedia , lookup
Epitranscriptome wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Expanded genetic code wikipedia , lookup
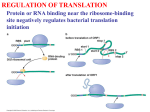
Cell, Vol. 111, 1–20, December 13, 2002, Copyright 2002 by Cell Press Shifty Ciliates: Frequent Programmed Translational Frameshifting in Euplotids Lawrence A. Klobutcher1,3 and Philip J. Farabaugh2 Department of Biochemistry University of Connecticut Health Center Farmington, Connecticut 06032 2 Department of Biological Sciences University of Maryland Baltimore County Baltimore, Maryland, 21250 1 Recent work suggests that there is a high frequency of programmed ⫹1 translational frameshifting in ciliates of the Euplotes genus. Frequent frameshifting may have been potentiated by stop codon reassignment, which is also a feature of this group. The way in which genetic information is translated into proteins had been considered universal: all organisms were thought to use a standard genetic code. Amazingly, the hard wiring of translation implicit in the genetic code now appears itself to be subject to change. The ciliated protozoa appear to have taken the greatest liberties with the so-called universal genetic code. Members of this group have the heaviest concentration of nuclear non-standard genetic codes recognized to date, typically using canonical stop codons to code instead for amino acids (see Lozupone et al., 2001; Tourancheau et al., 1995). For example, members of the genus Euplotes decode the conventional UGA stop codon as cysteine. In contrast, Tetrahymena and Paramecium retain a UGA stop codon, but decode the other two stop codons as glutamine, and it appears that this code has evolved independently up to five times. Changes to decoding can be subtler than the evolution of a non-standard genetic code. Often, mRNA sequences evolve to force a local change in the rules of decoding, for example decoding a specific termination codon as sense, or shifting the ribosome’s reading frame. A number of reports over the last few years indicate that euplotids have also taken these types of liberties with the code. Though the absolute numbers remain small, a significant fraction of available euplotid genes appear to require a ⫹1 translational frameshift to produce a functional protein. Sequence identities among the putative frameshift sites suggest that they occur as the ribosome encounters a termination codon, suggesting that the phenomenon of non-standard codes and non-canonical decoding may be mechanistically related. Programmed translational frameshifts occur when special sequences in mRNAs manipulate the ribosome to cause it to change its reading frame (reviewed in Baranov et al., 2002; Stahl et al., 2002). The outcome is that genetic information encoded discontinuously in the mRNA is expressed into a continuous protein product. Many metazoan viruses encode open reading frames (ORFs) that overlap, for example the retroviral gag and pol genes. Ribosomes that reach the end of the gag 3 Correspondence: [email protected] Minireview gene occasionally shift reading frames and continue synthesis into pol, producing a Gag-Pol fusion protein. The frequency of this frameshift is as much as 10,000fold greater than the estimated rate of spontaneous translational frameshifting. The sequence of the region in and around the site of frameshifting stimulates this impressive increase in “error.” Among the simplest types of programmed frameshifts are “⫹1 shifty stops” (Weiss et al., 1987). These sites consist of a poorly recognized termination codon immediately preceded by a sequence that can allow a tRNA to slip ⫹1 on the mRNA while still maintaining at least two base pairs. For example, in the prfB gene of Escherichia coli, a ⫹1 frameshift occurs at the sequence CUUUGA-C, shown in codons of the upstream, unshifted ORF (reviewed in Baranov et al., 2002). The signal UGA-C is recognized particularly poorly by peptide release factor II (RF2), itself the product of the prfB gene. RF2 recognizes UGA stop codons and triggers termination of translation. When the concentration of RF2 falls below the optimal concentration, recognition of the “internal” UGA codon in prfB mRNA is slowed, causing a translational pause with the peptidyl-tRNA in the ribosomal P site bound to the CUU codon. Frameshifting is thought to occur by the peptidyl-tRNA slipping from CUU to the UUU codon overlapping in the ⫹1 reading frame. This mechanism provides an autogenous control loop, optimizing expression of RF2, since a decrease in RF2 concentration would tend to increase its expression, but an increase would tend to decrease it. Frequency of Frameshifting in Euplotes The first report of a putative frameshifting event in a euplotid gene was for an open reading frame (ORF2) encoding a tyrosine-type recombinase in the Tec2 transposons of Euplotes crassus (Doak et al., 2003; Jahn et al., 1993). Because a number of mobile elements were known to require frameshifting for gene expression (reviewed in Farabaugh, 2000), this observation provided little reason to believe that frameshifting would be common in Euplotes cellular genes. However, more recent studies indicate that frameshifting may indeed be frequent. Evidence for frameshifting has been obtained for genes encoding the regulatory subunit of cAMPdependent protein kinase (PKAR) and a nuclear protein kinase (EoNdr2) in E. octocarinatus (Tan et al., 2001a, 2001b), a La motif protein (p43) in E. aediculatus (Aigner et al., 2000), and the telomerase reverse transcriptase (TERT) of E. crassus (Wang et al., 2002). The GenBank database includes sequences for the complete coding regions of only 67 genes from various Euplotes species. By contrast, among the ⵑ6000 genes in the yeast Saccharomyces cerevisiae, frameshifting is required for the expression of only two genes, encoding a subunit of telomerase, EST3 (Morris and Lundblad, 1997), and an actin binding protein, ABP140 (Asakura et al., 1998). Thus, only about 0.03% of yeast genes require frameshifting, while the limited euplotid data suggest that ⬎5% of genes may require a frameshift for expression. Cell 2 Figure 1. Frameshifting in Euplotes (A) The E. octocarinatus Eondr2 gene (black rectangle) is shown. The frame 0 ORF begins with an ATG initiation codon and encodes all of conserved protein kinase domain I, as well as a part of domain II. The second ORF (frame ⫹1) encodes a part of domain II, and the remaining 10 conserved protein kinase domains. Start and stop codons for the 0 and ⫹1 frame ORFs are indicated above and below the gene map, respectively. Figure based on data in Tan et al. (2001b). (B) The sequence of the E. aediculatus La motif protein mRNA in the vicinity of the frameshift site is shown, along with the predicted translation products from the 0 and ⫹1 frame ORFs. The AAA-UAAA motif is highlighted in gray. A frameshift at positions “1” or “2” would produce amino acid sequences that conform to the consensus of the “La motif,” including a highly conserved phenylalanine (underlined F) residue (for details, see Aigner et al., 2000). The Evidence for Translational Frameshifts Two types of evidence are typically necessary to support translational frameshifting: (1) solid sequencing data for the gene and mRNA, and (2) identification of a single protein produced from the separate reading frames. In all Euplotes cases, DNA sequencing revealed two separate ORFs that, if joined by translational frameshifting, would produce a single protein product similar to homologous proteins in other species. For example, serine/ threonine protein kinases contain twelve well-conserved domains. In the E. octocarinatus Eondr2 protein kinase gene (Figure 1A; Tan et al., 2001b), the first domain and part of the second are encoded by one ORF (frame O), while the remainder of second domain, and the other 10 domains, are encoded by a second out-of-frame ORF (frame ⫹1). Producing a protein with all the characteristic domains would require shifting the reading frame forward by one base in the region of overlap between the 0 and ⫹1 ORFs. In the case of the E. crassus TERT gene, three separate ORFs exist, so two ⫹1 frameshifts would be required to produce the protein (Wang et al., 2002). With the exception of the Tec2 transposon ORF2 gene and the E. crassus TERT gene, cDNA copies of the mRNAs for all of the genes have been sequenced (Aigner et al., 2000; Tan et al., 2001a, 2001b). All show the same arrangement of ORFs, indicating that mRNA editing does not result in the joining of the separate ORFs in the mRNA prior to translation. An additional concern is whether DNA sequencing errors might erroneously indicate frameshifting, particularly since some of the genes and mRNAs have been isolated by the polymerase chain reaction (PCR), which can introduce errors. This seems unlikely, as multiple independent clones of PCR products have often been analyzed, and direct sequencing of PCR products has also been performed (Aigner et al., 2000; Tan et al., 2001a, 2001b). While the nucleic-acid-based evidence for frameshifting is strong, there is less information available on the proteins. Indeed, only the La motif protein associated with telomerase in E. aediculatus has been isolated and analyzed (Aigner et al., 2000). A number of peptides derived from the purified La motif protein were sequenced. One of the peptides was encoded within the 0 frame ORF, while the remainder were encoded by the ⫹1 frame ORF, providing a clear indication that a single protein was produced by frameshifting. Conserved Sequences near ⫹1 Frameshift Sites During programmed frameshifting, the mRNA manipulates the translational machinery to cause a shift in reading frame. Frameshift stimulatory mRNA signals can include quite distant sequences or structural features. However, in each case the sequence of from 4 to 7 nt at the site of frameshifting is critical, as in the case of the prfB frameshift described above. The Euplotes frameshift genes share a common sequence motif strongly resembling known ⫹1 frameshift signals. In each gene, an AAA codon, coding for lysine, immediately precedes the stop codon of the 0 frame ORF. The stop codon is UAA, except for the Tec2 ORF 2 gene, which has a UAG stop codon (Jahn et al., 1993). There is also a strong tendency for the first base following the stop codon to be an A, so that the 0 frame ORF typically ends in the sequence 5⬘-AAA-UAA-A-3⬘. In the vicinity of this motif, there is currently no compelling evidence for other conserved sequence elements, or structural motifs such as a secondary structure-forming region, a common element of some types of frameshift sites. Tan et al. (2001b) have noted that the sequence 5⬘-CAAGAA3⬘ is often present within the 41 bases preceding the AAA-UAA-A motif, but exact matches to this sequence are not seen in all of the genes, nor is it clear how it could influence frameshifting. The precise position of the frameshift is unknown, as the amino acid sequence for this region has not been determined for any of the Euplotes frameshift proteins. However, the frameshift likely occurs near the AAAUAA-A motif. This is best illustrated using the La motif protein of E. aediculatus (Figure 1B). In this case, there is an extremely small overlap between the two ORFs, as the ⫹1 frame ORF has a termination codon located 11 bases upstream of the termination codon of the 0 frame ORF. As a result, the frameshift must occur somewhere in this short region. Moreover, this region represents part of the conserved La motif, and a frameshift at either the first or second codon upstream of the 0 frame stop codon would optimize amino acid sequence conservation (Figure 1B and see Aigner et al., 2000). Similar arguments can be made for the other genes. In Minireview 3 Table 1. Frequency of Stop Codon Usage in Euplotesa Stop Codon-Trinucucleotide UAA (87.9%) UAG (12.1%) Stop Codon-Tetranucleotide UAA-A (37.9%) UAG-A (1.7%) UAA-G (15.5%) UAG-G (0.0%) UAA-C (3.4%) UAG-C (5.2%) UAA-U (31.0%) UAG-U (5.2%) a Based on 58 Euplotes non-transposon, protein-coding genes available in the Transterm database (http://uther.otago.ac.nz/ Transterm.html). Figure 2. Model of the Euplotes ⫹1 Translational Frameshift The figure shows a ribosome encountering the AAA-UAA-A frameshift site. See text for details. the case of the EoNdr2 protein kinase, a frameshift at the lysine codon preceding the 0 frame stop codon would produce a protein with maximum sequence similarity to protein kinase domain II (Figure 1A; see Tan et al., 2001b). Possible Mechanism of the ⫹1 Frameshift The putative frameshift signal in euplotid genes can be considered a shifty stop (Weiss et al., 1987). A ribosome translating up to the site would stop with the AAA codon in the ribosomal P site and the UAA (or UAG) stop codon in the A site (Figure 2). If a peptidyl-tRNALys were to slip ⫹1 from AAA to AAU then another lysyl-tRNALys could enter the A site, accept the transfer of the peptide, and translocate to the P site. Translation would continue in the ⫹1 frame. Two questions arise from the above model. First, why is AAA always the last codon decoded in the 0 frame rather than other potential “slippery” codons such as UUU, CCC, or GGG? The finding implies that AAA, or its cognate tRNA, has some special feature. In the yeast Saccharomyces cerevisiae, ⫹1 frameshifting results from an abnormal codon·anticodon interaction at the equivalent codon (reviewed in Stahl et al., 2002). Whether this is true in euplotids is unclear since no information is available about their tRNAs. Second, why is termination at the UAA codon slow enough to allow frameshifting? Termination codons recognized poorly by release factor (RF) can stimulate frameshifting (reviewed in Bertram et al., 2001). In both prokaryotes and eukaryotes, RF appears to recognize a tetranucleotide sequence consisting of the termination codon and its 3⬘ nearest neighbor nucleotide. Certain tetranucleotides are recognized poorly by RF in vitro, and these same signals can stimulate frameshifting in vivo, presumably because their recognition is slow enough to allow time for the rare stochastic tRNA slippage thought to cause the shift in frame. The proposed model requires that UAA, and more specifically UAA-A, is a poorly recognized termination signal in euplotids. At first glance, this does not appear to be the case. Based on the analysis of 58 non-transposon, protein-coding genes, 87.9% of the Euplotes open reading frames (mainly predicted) have UAA as a terminator, and 37.9% end in UAA-A (Table 1). This raises the following conundrum: if UAA-A is frequently used as the “normal” termination signal, how is it capable of stimulating translational error at frameshift sites? We suggest that the reassignment of the UGA stop codon to a cysteine codon in euplotids has also resulted in poor recognition, and inefficient termination, for UAA codons. Stop codon reassignment is thought to require two steps (Osawa et al., 1990): (1) the development of a tRNA capable of decoding one of the stop codons, and (2) the loss of the ability of the RF to recognize the stop codon. In regard to the second step, a single release factor (eRF1) recognizes all three conventional stop codons in eukaryotes. Changes in the amino acid sequence of eRF1 at positions involved in interacting with one or more of the three nucleotides comprising a stop codon are thought to be necessary for the loss of recognition for particular stop codons. Indeed, a number of workers have characterized the sequences of eRF1 proteins from ciliates that have undergone stop codon reassignment in an attempt to determine which residues are involved in recognizing stop codons (e.g., Inagaki and Doolittle, 2001; Lozupone et al., 2001; Muramatsu et al., 2001). Since the three stop codons share nucleotide determinants, the same amino acid changes that block recognition of one stop codon may reduce the efficiency of recognition of one or both of the remaining stop codons. In the case of Euplotes, recognition of the UAA stop codon might be particularly impaired, as it shares two nucleotide positions in common with the reassigned UGA stop codon, while the UAG stop shares only one. Impaired or slow recognition of stop codons at natural termination sites may cause few problems in protein synthesis, but in the context of a shifty codon (such as AAA in Euplotes), it could serve to enhance the frequency of frameshifting. There is some evidence in support of this hypothesis. Seit-Nebi et al. (2002) introduced amino acid changes into the human eRF1 protein, some Cell 4 of which corresponded to residues present in the Paramecium eRF1, which recognizes only UGA stop codons. A number of these altered human eRF1 proteins displayed greatly impaired recognition of UAA and UAG stops in vitro, but, in addition, showed modest reductions in the recognition of UGA stop codons. Outstanding Issues Information on the mechanism of the ⫹1 frameshifting in euplotids is clearly needed. If translational frameshifting proves to be as common as the current limited data set indicates, the euplotids may present particular advantages for generally understanding the molecular mechanism underlying ⫹1 programmed translational frameshifts. Further analyses of eRF1, as well as analysis of tRNALys, may provide clues as to whether changes in stop codon usage can lead to proliferation of programmed frameshifts. It will also be important to determine if the AAA-UAA-A sequence is sufficient for frameshifting. An additional issue is whether frameshifting is involved in regulating gene expression in euplotids. While there is no obvious relationship between all of the Euplotes frameshift genes, two of them encode proteins involved in telomerase function (p43 and TERT), raising the possibility that frameshifting is involved in some form of coordinate regulation. Frameshifting can regulate protein expression in other systems (reviewed in Baranov et al., 2002; Farabaugh, 2000), as with the autogenous regulation of RF2 in E. coli. In other instances, the frequency of frameshifting is thought to set an optimal ratio of protein products. This is believed to be the case for retroviruses, where the ratio of the Gag and Gag-Pol fusion proteins depends on the frequency at which ribosomes frameshift. Whether Euplotes employs either of these regulatory mechanisms is unclear. Alternatively, it is possible that frameshifting has no regulatory role in euplotids. That is, these organisms may have evolved an efficient enough frameshifting system that generation of a frameshift site in a gene results in a negligible, perhaps selectively neutral, decrease in protein expression. A final area concerns the evolution of frameshifting. The process has been observed in E. crassus, E. aediculatus, and E. octocarinatus, and it will be of interest to determine if other euplotids and closely related groups also display frameshifting. There is already some indication that new frameshift sites have arisen during the evolution of euplotids, as the E. aediculatus TERT gene (Lingner et al., 1997) lacks the two frameshift sites present in the E. crassus TERT gene (Wang et al., 2002). A more general evolutionary question arises from the proposal that stop codon reassignment and frameshifting are related. That is, is the tendency to frameshifting a necessary outcome of reassigning termination codons to be decoded as sense? Most codon reassignments are in fact of termination codons, perhaps because that type of reassignment is less deleterious, but also because the reassignment may require only a small adjustment of the competition between RF and nonsense suppressor tRNA in the ribosomal A site (Lozupone et al., 2001). Adjusting competition certainly would eventually require restricting recognition of the reassigned termination codon, and, as we have discussed, this may reduce recognition of one of the remaining terminators. Pro- grammed frameshifting frequently results when the rate of a canonical process, such as termination, is reduced sufficiently to allow a normally extremely unlikely noncanonical process, like frameshifting, to proceed. It may be, therefore, that reassigning terminators will inevitably enhance the ability to evolve programmed frameshifts. It will be of great interest to see if shifty stop ⫹1 frameshifting is frequent in other species that have undergone termination codon reassignment. Selected Reading Aigner, S., Lingner, J., Goodrich, K.J., Grosshans, C.A., Shevchenko, A., Mann, M., and Cech, T.R. (2000). EMBO J. 19, 6230–6239. Asakura, T., Sasaki, T., Nagano, F., Satoh, A., Obaishi, H., Nishioka, H., Imamura, H., Hotta, K., Tanaka, K., Nakanishi, H., and Takai, Y. (1998). Oncogene 16, 121–130. Baranov, P.V., Gesteland, R.F., and Atkins, J.F. (2002). Gene 286, 187–201. Bertram, G., Innes, S., Minella, O., Richardson, J., and Stansfield, I. (2001). Microbiology 147, 255–269. Doak, T.G., Witherspoon, D.J., Jahn, C.L., and Herrick, G. (2003). Eukaryotic Cell, in press. Farabaugh, P.J. (2000). Prog. Nucleic Acid Res. Mol. Biol. 64, 131–170. Inagaki, Y., and Doolittle, W.F. (2001). Nucleic Acids Res. 29, 921–927. Jahn, C.L., Doktor, S.Z., Frels, J.S., Jaraczewski, J.W., and Krikau, M.F. (1993). Gene 133, 71–78. Lingner, J., Hughes, T.R., Shevchenko, A., Mann, M., Lundblad, V., and Cech, T.R. (1997). Science 276, 561–567. Lozupone, C.A., Knight, R.D., and Landweber, L.F. (2001). Curr. Biol. 11, 65–74. Morris, D.K., and Lundblad, V. (1997). Curr. Biol. 7, 969–976. Muramatsu, T., Heckmann, K., Kitanaka, C., and Kuchino, Y. (2001). FEBS Lett. 488, 105–109. Osawa, S., Muto, A., Jukes, T.H., and Ohama, T. (1990). Proc. R. Soc. Lond. B Biol. Sci. 241, 19–28. Seit-Nebi, A., Frolova, L., and Kisselev, L. (2002). EMBO Rep. 3, 881–886. Stahl, G., McCarty, G.P., and Farabaugh, P.J. (2002). Trends Biochem. Sci. 27, 178–183. Tan, M., Heckmann, K., and Brunen-Nieweler, C. (2001a). J. Eukaryot. Microbiol. 48, 80–87. Tan, M., Liang, A., Brunen-Nieweler, C., and Heckmann, K. (2001b). J. Eukaryot. Microbiol. 48, 575–582. Tourancheau, A.B., Tsao, N., Klobutcher, L.A., Pearlman, R.E., and Adoutte, A. (1995). EMBO J. 14, 3262–3267. Wang, L., Dean, S.R., and Shippen, D.E. (2002). Nucleic Acids Res. 30, 4032–4039. Weiss, R.B., Dunn, D.M., Atkins, J.F., and Gesteland, R.F. (1987). Cold Spring Harb. Symp. Quant. Biol. 52, 687–693.