* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Glioblastoma Multiforme (GBM) – Subtype Analysis
Public health genomics wikipedia , lookup
Metagenomics wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Gene desert wikipedia , lookup
Gene nomenclature wikipedia , lookup
Pharmacogenomics wikipedia , lookup
Minimal genome wikipedia , lookup
Gene therapy wikipedia , lookup
Genomic imprinting wikipedia , lookup
Genome evolution wikipedia , lookup
Ridge (biology) wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Oncogenomics wikipedia , lookup
Genome (book) wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
The Selfish Gene wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Mir-92 microRNA precursor family wikipedia , lookup
Microevolution wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Designer baby wikipedia , lookup
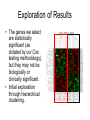
Glioblastoma Multiforme (GBM) – Subtype Analysis Lance Parsons Introduction • Clinicians (meat readers) determine histological categorization: Astrocytoma, Oligodendrocytoma, Mixed, or Glioblastoma multiforme (GBM) • GBM patients have poor prognosis, but some surive unexpectly long. • Molecularly and clinically distinct subtypes of GBMs Incorporating Biological Knowledge • “Tiers” of classification can assist with discovery of downstream groups • Glioma Classification – Histological Level – Clinical Level (Survival, Age, etc) – Transcriptiome (Gene Expression Level) • Gene Classification – GO Hierarchy – Pathway Databases – Expression Level (Microarray Data) Age Young Tumor Type Medium Old Astro Mixed Oligo GBM Gene Subset Expression X Y X Glioma Subtypes A B C Survival Short Long Age and Survival • • • Young patients show greater variability in survival Use this level of the “hierarchy” to assist in downstream analysis. Very simple method is to use only the Young samples and find the groups within that set of samples. Normalization • Making the numbers comparable – Log Transform – Equalize variance, lineraize data – Median Center Arrays – Correct for differences between arrays – Standardize to unit variance? Noise Filter • Removing noise from the dataset – Affymetrix software does some of this with Present/Absent calls – Fold-change filter? – Other methods? Feature (Gene) Selection • Find genes highly correlated with patient survival, within young sample group. • Cox Proportional Hazards model – Regression model that accounts for “censored” data • Permutation test can improve robustness • Simple Cox selects 39 genes (permutation pending) Exploration of Results • The genes we select are statistically significant (as dictated by our Cox testing methodology), but they may not be biologically or clinically significant. • Initial exploration through hierarchical clustering. Clinical Validation • Kaplan-Meier curves fit the two groups to a survival model Biological Validation • file:///C:/Documents%20and%20Settings/L ance/My%20Documents/Research/project s/HenryFord/HFAnalysis-GBMYoung_annotations.html