* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Slide 1
Group selection wikipedia , lookup
Frameshift mutation wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Genetic engineering wikipedia , lookup
Ridge (biology) wikipedia , lookup
Gene expression programming wikipedia , lookup
Polymorphism (biology) wikipedia , lookup
Minimal genome wikipedia , lookup
Genomic imprinting wikipedia , lookup
Genome evolution wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Gene expression profiling wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Genome (book) wikipedia , lookup
Designer baby wikipedia , lookup
Population genetics wikipedia , lookup
Genetically modified crops wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Genetically modified organism containment and escape wikipedia , lookup
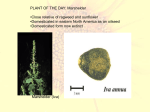
PLANT OF THE DAY: Marshelder •Close relative of ragweed and sunflower •Domesticated in eastern North America as an oilseed •Domesticated form now extinct Marshelder (Iva) Crop domestication Big Questions •Why were plants domesticated? •Where were plants domesticated? •When were plants domesticated? •How quickly were they domesticated? •What were the selection pressures that caused domestication? •What kinds of genetic changes are under selection during domestication? •Do analyses of evolution under domestication inform us about evolution under natural selection? •Why haven’t any major crops been domesticated recently? Centres of Plant Domestication • Concept first devised by Vavilov in 1919 • Archaeological evidence suggests that hunter-gatherers independently began cultivating food plants in at least 11 regions of the world (Doebley et al. 2006) Domestication ‘Domestication is the process by which humans actively interfere with and direct crop evolution.’ • It involves a genetic bottleneck: • Often only few genes are actively selected and account for large shifts in phenotype. • Crops exhibit various levels of domestication. What is a domestication syndrome? A domestication syndrome describes the properties that distinguish a certain crop from it’s wild progenitor. Typically such characteristics are: • larger fruits or grains • more robust plants • more determinate growth / increased apical dominance • loss of natural seed disperal • fewer fruits or grains • decrease in bitter substances in edible structures • changes in photoperiod sensitivity • synchronized flowering Tomato - Fewer and Larger Fruits Sunflowers - reduced branching, larger seeds, increased seed set per head Wheat - reduced seed shattering, increased seed size Squash – larger, fleshier fruits Corn – reduced fruitcase, softer glume, more kernels per cob, no dispersal, reduced branching, apical dominance Lettuce – leaf size/shape, fewer secondary compounds Rice – no shattering, larger grains Domestication is a process • The distinction ‘domesticated’ or ‘not domesticated’ is an oversimplification • Some crops have moved further along this process further than others. • We can recognize different levels of domestication • How can we decide which level? • Different domestication traits were selected for progressively •Distinction between selection under domestication vs. crop diversification more targeted, ‘conscious’ selection during diversification • ‘Slow’ rate of evolution of different domestication traits despite faster rates suggested by models • Artificial selection can be “similar across different taxa, geographical origins and time periods” • Parallel evolution for “sticky glutinous varieties” in rice and foxtail millets, all through selection at the waxy locus • Most QTL studies suggest that many domestication traits are controlled by a few genes of large effect – not though in sunflower • Population genomic studies in maize suggest 2 – 4% of genes show evidence of artificial selection Domestication of Maize How often has maize been domesticated? – Sampling (Matsuoka et al, 2002) How often has maize been domesticated? – Once. (Matsuoka et al, 2002) Tracking footprints of maize domestication and evidence for a massive selective sweep on chromosome 10 (Tian et al., 2009 PNAS) Teosinte branched 1 (tb1) • was identified as a major QTL controlling the difference in apical dominance between maize and its progenitor, teosinte (Doebley et al., 1997; Doebley, 2004) • is a member of the TCP family of transcriptional regulators, a class of genes involved in the transcriptional regulation of cell-cycle genes. •Differences in tb1 expression patterns between maize and teosinte indicate that human selection was targeted at regulatory differences that produced a higher level of tb1 message in maize. • Lack of any fixed amino acid differences between maize and teosinte in the TB1 protein supports this hypothesis. •First domestication gene cloned. “For maize tb1 … [the selection coefficient] is in the range 0.05 to 0.2, comparable to cases of natural selection.” (Purugannan and Fuller, 2009) Teosinte glume architecture1 (tga1) • was identified as a QTL controlling the formation of the casing that surrounds the kernels of the maize ancestor, teosinte (Wang et al., 2005) • is a member of the squamosa-promoter binding protein (SBP) family of transcriptional regulators. • tga1 has phenotypic effects on diverse traits including cell lignification, silica deposition in cells, three-dimensional organ growth, and organ size •The difference in function between the maize and teosinte alleles of tga1 appears to be the result of a single amino acid change. The fact that there are no discernable differences in gene expression supports this interpretation. Flowering Locus T (FT) Flowering Locus T (FT) protein is main component of florigen Four flowering time gene homologs – all members of the FT gene family – experienced selective sweeps during a stage of sunflower domestication Elite Landrace Wild Reduced nucleotide diversity (π) in HaFT paralogs with early domestication or improvement Genetics 2011; 187:271-287 Genetic and functional analyses identify causative mutations in HaFT paralogs Domesticated Allele of HaFT1 has frameshift mutation Current Biology 2010; 20:629-635 Loss of function mutation at FT also contributes to heterosis in other species “already kicking ass in commercial tomato production.” D. Zamir, Dec. 11, 2012, Asilomar Krieger et al. 2010 Homozygote (sFT) Heterozygote (sft Homozygote (sft) X sFT) Nature Genetics 42, 459-465, (2010) b Floret number 80 a 40 0.0 a Homozygote Heterozygote Homozygote Domesticated Wild (frameshift) (in-frame) Seed weight (g) Heterozygotes with frameshift allele exhibit heterosis 2.0 b a 1.0 0.0 a Homozygote Heterozygote Homozygote Domesticated Wild (frameshift) (in-frame) Current Biology 2010; 20:629-635 Domestication genes in plants FT Sunflower Transcriptional regulator; flowering time Yes Yes frameshift mutation Crop Diversification genes in plants The genetic basis of the evolution of non-shattering Non-shattering is often regarded as the hallmark of domestication in most seed crops because it renders a plant species primarily dependent on humans for survival and propagation: • rice gene sh4 (similar to the genes encoding MYBlike transcription factors in maize) • rice quantitative trait locus (QTL) qSH1, which encodes a homeobox-containing protein • the wheat gene Q, which is similar to genes of the AP2 family in other plants • In sunflower likely controlled by multiple genes Domestication genes in plants • Maize and rice domestication seem to suggest few loci of large effect are important • Sunflower domestication seems to suggest many loci of small to intermediate effect are important • 9 domestication genes in plants so far, as well as 26 other loci known to underlie crop diversity • Of the 9 domestication loci, 8 encode transcriptional activators. • More than half of crop diversification genes encode enzymes. Domestication seems to be associated with changes in transcriptional regulatory networks, whereas crop diversification involves a larger proportion of enzymeencoding loci (lots of them loss-of-function alleles). The role of polyploidy in domestication Where does the cultivated gene pool come from? Sclerotinia resistance locus Wild Introgressions H. petiolaris H. argophyllus H. annuus landraces Unanswered questions Why do