* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Results from the GAIT project: Genetic analysis of
Y chromosome wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Ridge (biology) wikipedia , lookup
Pharmacogenomics wikipedia , lookup
Genomic imprinting wikipedia , lookup
Minimal genome wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Nutriepigenomics wikipedia , lookup
X-inactivation wikipedia , lookup
Genome evolution wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Population genetics wikipedia , lookup
Human genetic variation wikipedia , lookup
Gene expression profiling wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Gene expression programming wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Genetic testing wikipedia , lookup
Genetic engineering wikipedia , lookup
History of genetic engineering wikipedia , lookup
Public health genomics wikipedia , lookup
Behavioural genetics wikipedia , lookup
Designer baby wikipedia , lookup
Microevolution wikipedia , lookup
Heritability of IQ wikipedia , lookup
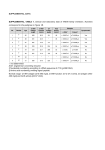
Which of these phenotypes share the same genetic influences? Pleiotropy: Textbook Example Cystic Fibrosis, simple autosomal recessive One gene influences multiple organs: lungs, liver, pancreas, small intestine, reproductive tract, skin (sweat glands) Trait 1 VA Trait 2 VA No Pleiotropy Trait 1 VA Trait 2 VA Partial Pleiotropy Trait 2 VA Trait 1 VA Complete Pleiotropy Phenotypic correlations can be broken down into genetic and environmental components P 2 1 h and h h 2 2 2 1 h G 1 h 2 2 2 1 1 h E 2 2 = heritabilities for traits 1 and 2 P = phenotypic correlation G = additive genetic correlation E = environmental correlation Extension to multivariate analysis G 2 + E I = Kronecker product operator E = residual environment covariance matrix G = additive genetic covariance matrix Nested models for bivariate analyses Parameters estimated Model e1 & e2 a1 & a2 e g ---------------------------------------------------------------------------General additive + + + + No pleiotropy + + + 0 Complete pleiotropy + + + 1 Examples from the GAIT Project. Phenotypic correlations can be broken down into genetic and environmental components P 2 1 h and h h 2 2 2 1 h G 1 h 2 2 2 1 1 h E 2 2 = heritabilities for traits 1 and 2 P = phenotypic correlation G = additive genetic correlation E = environmental correlation Phenotypic correlations among vitamin K dependent proteins. FVII FIX FX 0.39 0.46 0.54 0.35 0.48 0.43 FII FVII FIX FX Protein C Total Protein S PC 0.45 0.46 0.42 0.41 tPS 0.29 0.20 0.30 0.45 0.31 fPS 0.41 0.29 0.29 0.45 0.35 0.63 Genetic and environmental correlations among vitamin K dependent proteins. FII FII FVII FIX FX Protein C Total PS Free PS 0.55 0.41 0.82 0.59 0.30 0.19 FVII FIX FX 0.23 0.53 0.35 0.08 0.14 0.55 0.23 0.71 0.53 0.51 0.48 0.61 0.24 0.23 0.37 0.21 0.30 0.30 PC 0.31 0.42 0.33 0.15 tPS 0.46 0.22 0.35 0.65 0.33 0.33 0.25 0.55 fPS 0.86 0.57 0.51 0.86 0.67 0.80 Discrete Continuous (disease) (liability) 0 1 t Correlations with liability to thrombosis Trait p APCR FVII FVIII FIX FXI FXII Homocysteine t-PA vWF -0.23* 0.03 0.29* 0.15 0.21* 0.17* 0.23* 0.18* 0.26* g -0.65* -0.35 0.69* 0.60* 0.56* 0.35* 0.65* 0.75* 0.73* e 0.67* 0.57* -0.13 -0.20 0.07 -0.15 -0.28 -0.10 -0.18 Genetic correlations Liability to Thrombosis -0.65 ± 0.14 p = 0.000001 APCR -0.54 ±0.12 p = 0.009 0.69 ± 0.15 p = 0.0005 Factor VIII levels Inference A single gene or set of genes influences variation in risk for thrombosis, factor VIII levels, von Willebrand factor levels, and activated protein C resistance. However, each of these traits is also affected by additional genes not shared with the others. Genetic correlations 0.65 ± 0.20 p = 0.002 Homocysteine -0.20 ± 0.19 -0.16 ± 0.23 Liability to Thrombosis APCR Factor VIII Inference Evidence exits for a single gene or set of genes which influences both homocysteine levels and risk for thrombosis. This is a different gene or set of genes than the one with common influences on factor VIII, vWF, APCR, and thrombosis. Another way of looking at G FXII - liability G = 0.35, h2 = 0.68 for FXII, 0.61 for liability 12.25% of genetic variance shared in common (i.e. G squared) FXII: 8.3% of variance genes shared w/ liability, 59.7% of variance genes not shared w/liability Liability: 7.5% of variance genes shared w/FXII, 53.5% of variance genes not shared w/FXII Extension to multivariate analysis ˆ G 2 + E I Q P = Kronecker product operator Q = additive genetic covariance matrix for QTL G = residual additive genetic covariance matrix E = environmental covariance matrix Nested models for bivariate analyses Parameters estimated Model e1 & e2 a1 & a2 q1 & q2 e g q --------------------------------------------------------------------------------Sporadic + 0 0 + Additive + + 0 + + Linkage + + + + + + Pleiotropy + + + + + 1 Coincident + + + + + 0 Pleiotropy: a single gene influencing two or more phenotypes, q = 1 or -1 Coincident Linkage: two closely placed genes, each influencing different phenotypes, q = 0 Why do multivariate linkage analysis? • Exploits pleiotropy to improve power to detect linkage • Allows a formal test of pleiotropy versus coincident linkage • Improves estimate of QTL location and effect size Correlations with liability to thrombosis Trait p APCR FVII FVIII FIX FXI FXII Homocysteine t-PA vWF -0.23* 0.03 0.29* 0.15 0.21* 0.17* 0.23* 0.18* 0.26* g -0.65* -0.35 0.69* 0.60* 0.56* 0.35* 0.65* 0.75* 0.73* e 0.67* 0.57* -0.13 -0.20 0.07 -0.15 -0.28 -0.10 -0.18 Results of FXII genome screen (LODs > 1) Location LOD Ch 5, 193 cM Ch 10, 38 cM Ch 2, 9 cM Ch 11, 10 cM Ch 14, 63 cM Ch 15, 79 cM 4.73 3.53 2.26 1.30 1.16 1.03 FXII – Chromosome 10 QTL FXII – Chromosome 5 QTL FXII levels are genetically correlated with risk of thrombosis. Do either of the QTLs detected through FXII levels influence liability to thrombosis? Does the QTL identified through trait1 also influence trait2? Parameters estimated Model e1 & e2 a1 & a2 q1 & q2 e g q -------------------------------------------------------------------------------------No pleiotropy + + +/0 + + Pleiotropy + +/+ + + + 1 Bivariate analysis with liability to thrombosis Chromosome 10 – no improvement in likelihood Chromosome 5 – h2q for liability > 0, p = 0.004 Inference: The FXII QTL on chromosome 5 also influences susceptibility to thrombosis. Summary Two QTLs on chromosomes 5 and 10 influence FXII levels. The QTL on chromosome 5 also influences liability to thrombosis and is likely to be the FXII structural gene. FXII 46C/T appears to functionally influence FXII levels, but our results suggest additional functional variants exist in or near FXII.