* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Lecture 9: Biological Pathway Simulation
Deoxyribozyme wikipedia , lookup
Ligand binding assay wikipedia , lookup
Lipid signaling wikipedia , lookup
Isotopic labeling wikipedia , lookup
Metalloprotein wikipedia , lookup
Enzyme inhibitor wikipedia , lookup
Secreted frizzled-related protein 1 wikipedia , lookup
Biochemistry wikipedia , lookup
Clinical neurochemistry wikipedia , lookup
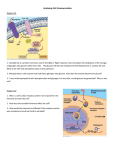
Western blot wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Proteolysis wikipedia , lookup
Photosynthetic reaction centre wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Basal metabolic rate wikipedia , lookup
Oxidative phosphorylation wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Multi-state modeling of biomolecules wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Gene regulatory network wikipedia , lookup
Biosynthesis wikipedia , lookup
Ultrasensitivity wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Mitogen-activated protein kinase wikipedia , lookup
Metabolic network modelling wikipedia , lookup
Signal transduction wikipedia , lookup
LSM3241: Bioinformatics and Biocomputing Lecture 9: Biological Pathway Simulation Prof. Chen Yu Zong Tel: 6874-6877 Email: [email protected] http://xin.cz3.nus.edu.sg Room 07-24, level 7, SOC1, NUS Biomolecular Interaction: Enzyme + Substrate E + S ==> E + P • This is a generalization of how a biochemist might represent the function of enzymes. 2 Biomolecular Interaction: Enzyme + Substrate E + S ==> E + P kinase-ATP complex + inactive-enzyme ==> Kinase + ADP + active enzyme K P ATP ADP • Here is an example of the generalization represented by two different ways. 3 Biomolecular Interaction: Enzyme + Substrate inactive enzyme Active enzyme Kinase-ATP complex ADP • This is another representation. 4 Spoke and Matrix Models of Protein-Protein Interactions Vrp1 (bait), Las17, Rad51, Sla1, Tfp1, Ypt7 Spoke Possible Actual Topology Simple model Intuitive, more accurate, but can misrepresent Matrix Theoretical max. no. of interactions, but many FPs Bader&Hogue Nature Biotech. 2002 Oct 20(10):991-7 5 Cell Polarity Cell Wall Maintenance Cell Structure Mitosis Chromosome Structure DNA Synthesis DNA Repair Unknown Others Synthetic Genetic Interactions in Yeast 6 Tong, Boone Metabolic Pathway: ATP Production • Glycolysis – Phosphorylation – Pyruvate • Anaerobic respiration • Lactate production • 2 ATPs produced 7 Cyclic Metabolic Pathway 8 Methods of Metabolic Engineering 9 Generic Signaling Pathway Signal Receptor (sensor) Transduction Cascade Targets Metabolic Enzyme Response Altered Metabolism Gene Regulator Altered Gene Expression Cytoskeletal Protein Altered Cell Shape or Motility 10 Components of Signaling What can be the Signal? External message to the cell • • • • • • • • Peptides / Proteins- Growth Factors Amino acid derivatives - epinephrine, histamine Other small biomolecules - ATP Steroids, prostaglandins Gases - Nitric Oxide (NO) Photons Damaged DNA Odorants, tastants Signal = LIGAND Ligand- A molecule that binds to a specific site on another molecule, usually a protein, ie receptor 11 Components of Signaling What are Receptors? Sensors, what the signal/ligand binds to initiate ST Cell surface Cell-Surface Receptor Hydrophillic Ligand Plasma membrane Intracellular Hydrophobic Ligand Carrier Protein Intracellular Receptor Nucleus 12 Generic Signal Transduction 13 RTK Signal Transduction 14 Signal Transduction Downstream effectors Protein Signaling Modules (Domains) SH2 and PTB bind to tyrosine phosphorylated sites SH3 and WW bind to proline-rich sequences PDZ domains bind to hydrophobic residues at the C-termini of target proteins PH domains bind to different phosphoinositides FYVE domains specifically bind to Pdtlns(3)P (phosphatidylinositol 3-phosphate) 15 Mechanisms for Activation of Signaling Proteins by RTKs Activation by membrane translocation Activation by a conformational change Activation by tyrosine phosphorylation 16 Mechanisms for Attenuation & Termination of RTK Activation 1) Ligand antagonists 2) Receptor antagonists 3) Phosphorylation and dephosphorylation 4) Receptor endocytosis 5) Receptor degradation by the ubiquitin-proteosome pathway 17 Activation of MAPK Pathways by Multiple Signals 18 Growth, differentiation, inflammation, apoptosis -> tumorigenesis Overview of MAPK Signaling Pathways 19 The MAPK Pathway Activated by RTK 20 P RTK ST- PI3K pathway 21 Apoptosis Pathways 22 TGF Pathway 23 Constructing a pathway model: things to consider 1. Dynamic nature of biological networks. Biological pathway is more than a topological linkage of molecular networks. Pathway models can be based on network characteristics including those of invariant features. 24 Constructing a pathway model: things to consider 2. Abstraction Resolution: • How much do we get into details? • What building blocks do we use to describe the network? High resolution (A) Substrates and proteins (B) Pathways (C) “special pathways” Low resolution 25 Constructing a pathway model Step I - Definitions We begin with a very simple imaginary metabolic network represented as a directed graph: Vertex – protein/substrate concentration. Edge - flux (conversion mediated by proteins of one substrate into the other) How do we define a biologically significant system boundary? Internal flux edge External flux edge 26 Constructing a pathway model Step II: Interaction Kinetics E + S ==> E + P kinase-ATP complex + inactive-enzyme ==> Kinase + ADP + active enzyme K P ATP ADP Reversibility of Chemical Reactions: Equilibrium H2 2H • Chemical reactions are reversible • Under certain conditions (concentration, temperature) both reactants and products exist together in equilibrium state 28 Reaction Rates Net reaction rate = forward rate – reverse rate • In equilibrium: Net reaction rate = 0 • When reactants “just” brought together: Far from equilibrium, focus only on forward rate • But, same arguments apply to the reverse rate 29 The Differential Rate Law • How does the rate of the reaction depend on concentration? E.g. 3A + 2B C + D rate = k [A]m[B]n (Specific reaction) rate constant Order of reaction with respect to A m+n: Overall order of the reaction Order of reaction with respect to B 30 Rate Constants and Reaction Orders • Each reaction is characterized by its own rate constant, depending on the nature of the reactants and the temperature • In general, the order with respect to each reagent must be found experimentally (not necessarily equal to stoichiometric coefficient) 31 Elementary Processes and Rate Laws • Reaction mechanism: The collection of elementary processes by which an overall reaction occurs • The order of an elementary process is predictable Unimolecular A* B K+ [A] First order Bimolecular A +BC+D K+ [A] [B] Second order Trimolecular A + B + C D + E K+ [A] [B] [C] Third order 32 Elementary Processes and Rate Laws • Reaction mechanism: The collection of elementary processes by which an overall reaction occurs • The order of an elementary process is predictable Unimolecular A* B K+ [A] – K- [B] First order Bimolecular A + B C + D K+ [A] [B] – K- [C] [D] Second order Trimolecular Third A + B + C D + E K+ [A] [B] [C] – K- [D] [E] order 33 Constructing a pathway model Step III - Dynamic mass balance Concentration vector Stoichiometry Flux vector Matrix dx S v dt 34 A ‘simple’ ODE model of yeast glycolysis 35 A model pathway system and its time-dependent behavior Positive Feedback Loop 36 A model pathway system and its time-dependent behavior 37 A model pathway system and its time-dependent behavior 38