* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download video slide
Signal transduction wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Peptide synthesis wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Magnesium transporter wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Gene regulatory network wikipedia , lookup
Expression vector wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Transcriptional regulation wikipedia , lookup
Epitranscriptome wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Western blot wikipedia , lookup
Interactome wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Metalloprotein wikipedia , lookup
Gene expression wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Genetic code wikipedia , lookup
Point mutation wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Biosynthesis wikipedia , lookup
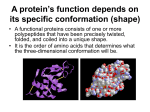
The Structure and Function of Macromolecules Chapter 5 3 -- Proteins Macromolecules: The Molecules of Life Carbohydrates Nucleic Acids Proteins Lipids 2 Proteins Polypeptides -- polymers of amino acids Protein -- one or more polypeptides Proteins -- many structures with wide range of functions Proteins -- more than 50% of the dry mass of most cells Proteins -- include structural support, storage, transport, cellular communications, movement, and defense against foreign substances 3 4 Amino Acid Monomers Organic molecules with carboxyl and amino groups Different properties due to differing side chains, called R groups 20 amino acids to make thousands of proteins 5 LE 5-UN78 a carbon Amino group Carboxyl group LE 5-17a Glycine (Gly) Alanine (Ala) Valine (Val) Leucine (Leu) Isoleucine (Ile) Nonpolar Methionine (Met) Phenylalanine (Phe) Tryptophan (Trp) Proline (Pro) LE 5-17b Polar Serine (Ser) Threonine (Thr) Cysteine (Cys) Tyrosine (Tyr) Asparagine (Asn) Glutamine (Gln) LE 5-17c Acidic Basic Electrically charged Aspartic acid (Asp) Glutamic acid (Glu) Lysine (Lys) Arginine (Arg) Histidine (His) Amino Acid Polymers Amino acids -- linked by peptide bonds A polypeptide -- polymer of amino acids Polypeptide length -- few monomers to more than a thousand Each polypeptide has a unique linear sequence of amino acids 10 Determining the Amino Acid Sequence of a Polypeptide First determined by chemical methods Now? Mostly automated – DNA sequencer 11 LE 5-20a Amino end Primary structure Amino acid subunits unique sequence of amino acids Carboxyl end LE 5-20b Secondary structure Interactions between backbone components Hydrogen bonds Typical secondary structures are coils (alpha helix) and a folded structure (beta pleated sheet) b pleated sheet Amino acid subunits a helix 13 Tertiary Structure LE 5-20d Interactions between R groups Include hydrogen bonds, ionic bonds, hydrophobic interactions, and van der Waals interactions Disulfide bridges may reinforce the protein’s conformation Hydrophobic interactions and van der Waals interactions Polypeptide backbone Hydrogen bond Disulfide bridge Ionic bond 14 Four Levels of Protein Structure Primary structure -- unique sequence of amino acids Secondary structure – interactions between backbone components Tertiary structure -- interactions between various side chains (R groups) Quaternary structure – proteins consisting of multiple polypeptide chains 15 Proteins with Quaternary Structure Collagen is a fibrous protein consisting of three polypeptides coiled like a rope Hemoglobin is a globular protein consisting of four polypeptides: two alpha and two beta chains 16 17 Sickle-Cell Disease: A Simple Change in Primary Structure A slight change in primary structure can affect a protein’s conformation and ability to function Sickle-cell disease, an inherited blood disorder, results from a single amino acid substitution in the protein hemoglobin 10 µm Red blood Normal cells are cell shape full of individual hemoglobin molecules, each carrying oxygen. 10 µm Red blood cell shape Fibers of abnormal hemoglobin deform cell into sickle shape. 18 LE 5-21b Sickle-cell hemoglobin Normal hemoglobin Primary structure Val His 1 2 Leu Thr 3 4 Pro Glu 5 6 Secondary and tertiary structures 7 b subunit Quaternary Normal hemoglobin structure (top view) Primary structure Secondary and tertiary structures Molecules do not associate with one another; each carries oxygen. His Leu Thr Pro Val Glu 1 2 3 4 5 6 7 Exposed hydrophobic region b subunit a Quaternary structure b Val b a Function Glu Sickle-cell hemoglobin b a Function Molecules interact with one another to crystallize into a fiber; capacity to carry oxygen is greatly reduced. b a LE 5-19 Groove A ribbon model Groove A space-filling model Conformation and Function Conformation – 3-D shape Functional protein -- one or more polypeptides twisted, folded, and coiled into a unique shape Sequence -- determines a protein’s threedimensional conformation Conformation -- determines its function Ribbon models and space-filling models can depict a protein’s conformation 21 What Determines Protein Conformation? In addition to primary structure, physical and chemical conditions can affect conformation Alternations in pH, salt concentration, temperature, or other environmental factors can cause a protein to unravel This loss of a protein’s native conformation is called denaturation A denatured protein is biologically inactive 22 LE 5-22 Denaturation Normal protein Denatured protein Renaturation The Protein-Folding Problem Prediction of conformation is non-trivial Thousands of possible conformations! Most proteins probably go through several states on their way to a stable conformation Chaperonins assist the proper folding of other proteins 24 LE 5-23a Cap Hollow cylinder Chaperonin (fully assembled) LE 5-23b Polypeptide Steps of Chaperonin Action: An unfolded polypeptide enters the cylinder from one end. Correctly folded protein The cap attaches, causing the cylinder to change shape in such a way that it creates a hydrophilic environment for the folding of the polypeptide. The cap comes off, and the properly folded protein is released. Scientists use X-ray crystallography to determine a protein’s conformation Another method is nuclear magnetic resonance (NMR) spectroscopy, which does not require protein crystallization 27 X-ray diffraction pattern 3D computer model The Flow of Genetic Information The information content -- DNA sequence DNA – directs synthesis of proteins Gene manufacture Transcription Translation Ribosome -- where translation happens 29 LE 17-4 Gene 2 DNA molecule Gene 1 Gene 3 DNA strand (template) 5 3 TRANSCRIPTION mRNA 5 3 Codon TRANSLATION Protein Amino acid 30 Nutritional Mutants in Neurospora Beadle and Tatum – irradiated mold resulting in inability to synthesize certain molecules Three classes of arginine-deficient mutants 3 different enzymes necessary for synthesizing arginine “One gene–one enzyme” hypothesis 31 The Products of Gene Expression: A Developing Story Not all proteins are enzymes! One gene–one protein Quaternary structure -- each component needs its own gene ATP synthase has 16 subunits! Now -- Beadle and Tatum’s hypothesis “one gene–one polypeptide” 32 Basic Principles of Transcription and Translation Transcription -- synthesis of RNA under the direction of DNA produces messenger RNA (mRNA) “language” of DNA to “language” of RNA Translation -- synthesis of a polypeptide under the direction of mRNA Ribosomes are the sites of translation “language” of Nucleic Acids to “language” of Amino Acids 33 LE 17-4 Gene 2 DNA molecule Gene 1 Gene 3 DNA strand (template) 5 3 TRANSCRIPTION mRNA 5 3 Codon TRANSLATION Protein Amino acid 34 DNA to RNA DNA in eukaryotes is in the nucleus Protein synthesis occurs at ribosomes in the cytoplasm DNA information -- from nucleus to cytoplasm intermediary (RNA) 35 RNA Intermediaries 36 A ribosome has three binding sites for tRNA: P site -- holds the tRNA The A site -- holds the tRNA with next amino acid The E site -- exit site, where discharged tRNAs leave the ribosome 37