* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Microbial Genetics - University of Montana
Quantitative trait locus wikipedia , lookup
Ridge (biology) wikipedia , lookup
Public health genomics wikipedia , lookup
Human genome wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Point mutation wikipedia , lookup
Gene expression profiling wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Non-coding DNA wikipedia , lookup
Primary transcript wikipedia , lookup
Genetic engineering wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Designer baby wikipedia , lookup
Helitron (biology) wikipedia , lookup
Genome (book) wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Genome evolution wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Microevolution wikipedia , lookup
Minimal genome wikipedia , lookup
Genome editing wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
History of genetic engineering wikipedia , lookup
Genomic library wikipedia , lookup
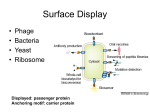
Microbial Genetics MICB404, Spring 2006 Lecture #17 Bacteriophage I Lytic phage • Announcements • Today we finish conjugation • Bacteriophage I: Lytic phage Hfr gene mapping E. coli chromosome genetic map Selected Marker hisG Selected Markers argH rif trpA 7% 6% 1% Plot frequency of unselected markers Selected = argH Selected = hisG 89% 100 7 12% 6 1 100 90 argH 10 20 30 trpA rif Prime factor selection Prime Factors • Selection – Early transfer of distal markers • Markers that were far from oriT in chromosome of Hfr will be close in F’, and transferred early • Recipient is merodiploid and transconjugant, not recombinant • Capable of conjugation Prime Factors • Selection – Replicons • Capable of replication independent of chromosome • If recipient is defective in recombination, prime factor transconjugants can acquire and transmit selected marker but no recombinants from Hfr individuals form Bacteriophage • Viruses that infect bacteria Bacteriophage • Important in the history of genetic research – – – – – – replication transcription recombination gene regulation DNA is the genetic material mRNA is an intermediate in translating genetic information from DNA to protein – http://www.asm.org/division/m/blurbs/Secrets.html • Important tools in molecular biology – Enzymes (T4 DNA Ligase, T7 RNA polymerase, λ exonuclease, cre recombinase) – λ DNA – phage cloning vectors – phage display Confirmation that genes consist of DNA Bacteriophage • Bacterial parasites – Exploits host metabolism for replication and production of new phage particles • amino acids, cellular energy, translation – Phage genes encode factors required to re-direct cellular activity to phage manufacture • Minimum functions for survival – – – – Protection of genome in environment Delivery of genome into bacterium Conversion of infected bacterium to phage factory Release of progeny phage Bacteriophage • 10-100+ genes • Structure – Icosahedral tailless – Icosahedral tailed – Filamentous Tail Fibers Bacteriophage • Head: Coat or Capsid – Contains nucleic acid • Tail & fibers – Interaction with bacterial surface – Injection of DNA into bacterium Bacteriophage • Nucleic acid – single-stranded DNA – double-stranded DNA • linear or circular molecules – RNA • generally, linear & single-stranded – Genome size • • • • M13: 7200 bases ssDNA λ: 49,000 bp T4: 160,000 bp packaging Lytic Phage Life Cycle • • • • • • • • Adsorption Injection Transcription Take over Production Packaging Assembly Lysis Phage Life Cycle • Adsorption – binding to receptors in bacterial cell surface • receptors play various roles for cells, e.g. solute uptake • reversible phase • irreversible phase Phage Life Cycle • Injection – DNA passes through bacterial cell wall and membrane • Medium exposure – None for tailed phage – Occurs with tailless phage • Transcription – use host-like promoters for expression of early genes Phage Life Cycle • Take over – phage shut down host replication, transcription, and translation – re-program cell to produce phage coat proteins and genome • alter activity of host enzymes – phage-specific sigma factors • express phage-specific enzymes – DNA & RNA polymerases, etc. • evade host defense mechanisms • Production Phage Life Cycle – phage nucleic acids and proteins synthesized in host cell • structural proteins • catalytic proteins (maturation) • Packaging – Nucleic acid inserted into heads • Assembly – coat proteins assembled into capsids – tail proteins assembled • Lysis Phage Life Cycle – Lysozyme or endolysin one of the late-expressed enzymes – Disrupts cell wall – Phage released to environment – Note: filamentous phage are extruded through cell wall without lysis or cell death • Yield: 10-1000 phage per cell Lytic Development Cycle • Early genes • Middle genes • Late genes Lytic Development Cycle • Early genes • Middle genes • Late genes Studying phage • Plaques: clear zones in lawns of plated bacteria caused by cell lysis – Plaque morphology affected by phage genotype • small, large • sharp, haloed Studying phage Bacteria Phage Plate 108 cells and 1 phage Infection and lysis 100 phage 2nd round of infection & lysis 10,000 phage Lawn Plaque Result: all cells are killed and lysed surrounding site of original phage particle 1 phage results in 1 plaque (106 phage) Studying phage • Multiplicity of Infection MOI – Average number of phage adsorbed per bacterium • mixing 107 E. coli with 3x107 T4 phage: MOI = 3 – High, >1 – Low, <1 Studying phage • Calculate the proportion of cells infected by a specific number of phage • Poisson distribution • P(n) = (mn * e-m)/n! where: m = MOI n = number of phage infecting a cell P(n) = probability cell will be infected with n phage (e.g. MOI =1: at most 63% cells infected) Studying phage • Burst size – Number of phage produced by infected cell = (number of phage produced after lysis) (number of infected cells) – Typical burst size is 100 for wildtype phage • mutants with burst size <10 can produce plaques Host defense • Restriction-Modification • Expression of surface receptors • Activity Phage genetics • Bacteriophage T4 – Life cycle & replication – Genetic analysis Phage genetics • Bacteriophage T4 • > 200 genes T series of bacteriophage Morphology Name Plaque size Head (nm) Tail (nm) Latent period (min) Burst size T1 medium 50 150 x 15 13 180 T2 small 65 x 80 120 x 20 21 120 T3 large 45 invisible 13 300 T4 small 65 x 80 120 x 20 23.5 300 T5 small 100 tiny 40 300 T6 small 65 x 80 120 x 20 25.5 T7 large 45 invisible 13 T-even and T-odd phage The T-even phages, T2, T4 and T6, are all related serologically and all have large genomes; T4 has a genome 168,895 bp in length The T-odd phages fall into three serological groups: T3 and T7 are related to each other but not to T1 or to T5, which are unrelated. The T7 genome was sequenced in 1983; it is 39,937 bp in length 200-300 300 T4 • Seymour Benzer, 1950’s – Fine-structure mapping of rII (rapid lysis mutants type II) locus • rII will complete life cycle in E. coli B but not on K12λ – but can infect K12λ • Complementation indicated rII has 2 genes or “complementation groups” T4 • r mutants distinguished based on plaque morphology Complementation Co-infect K12 with 2 rII mutant phage: A a1 a b B Defects in different genes: infection has functional copy of each gene: complementation. Massive lysis with progeny mostly of parental Ab or aB genotypes a2 B B Overlapping defects in same gene: no complementation. No lysis, no progeny phage Recombination Recombination frequency • The closer two sequence regions are to one another, the less room for, and less likelihood of, crossover. • Frequency of recombinants is therefore a measure of how far apart mutations are in the sequences. • Recombination frequency. Recombination Co-infect E. coli B with 2 rII mutant phage: a1 a2 B B a1 a2 B A B Non-overlapping defects in same gene: recombination. Low degree of K12 lysis, progeny are almost entirely AB, wild-type Collect phage lysate Recombination Co-infect E. coli B with 2 rII mutant phage: a1 a2 B B a1 a2 B Mapping Infect both B and K12 B: plaques represent total number of progeny phage K12: wildtype are ½ of total recombinants Map distance: 2 x f(wild-type progeny) x 100 A B • Monday’s lecture: – Bacteriophage II – Reading • Continue with Snyder and Champness, Chapter 7