* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Molecular Basis for Relationship between Genotype and Phenotype
Endogenous retrovirus wikipedia , lookup
Alternative splicing wikipedia , lookup
RNA interference wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Gene regulatory network wikipedia , lookup
Proteolysis wikipedia , lookup
Real-time polymerase chain reaction wikipedia , lookup
Community fingerprinting wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Metalloprotein wikipedia , lookup
Amino acid synthesis wikipedia , lookup
RNA silencing wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Non-coding DNA wikipedia , lookup
Biochemistry wikipedia , lookup
Messenger RNA wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Polyadenylation wikipedia , lookup
Eukaryotic transcription wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Point mutation wikipedia , lookup
Transcriptional regulation wikipedia , lookup
Genetic code wikipedia , lookup
RNA polymerase II holoenzyme wikipedia , lookup
Gene expression wikipedia , lookup
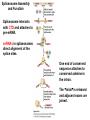
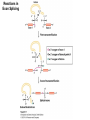
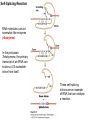
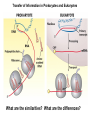
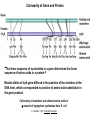
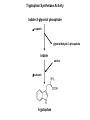
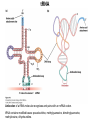
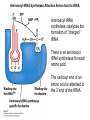
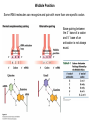
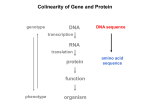
Molecular Basis for Relationship between Genotype and Phenotype genotype DNA DNA sequence transcription RNA translation protein function phenotype organism amino acid sequence Transcription Initiation in Eukaryotes TATA binding protein (TBP), part of TFIID complex, must bind to promoter before other GTFs and RNA polymerase II can form preinitiation complex (PIC). Phosphorylation of carboxyl tail domain (CTD), the protein tail of b subunit of RNA polymerase II, allows separation of RNA polymerase II from GTFs to start transcription. Cotranscriptional Processing of RNA State of phosphorylation of CTD determines the type of proteins that can associate with the CTD (thus defining cotranscriptional process). 5’ end of pre-mRNA is capped with 7-methylguanosine. This protects the transcript from degradation; capping is also necessary for translation of mature mRNA. Cotranscriptional Processing 3’ end of the transcript typically contains AAUAAA or AUUAAA. This sequence is recognized by an enzyme that cleaves the newly synthesized transcript ~20 nucleotides downstream. At the 3’ end, a poly(A) tail consisting of 150 200 adenine nucleotides is added. Polyadenylation is another characteristic of transcription in eukaryotes. Complex Patterns of Eukaryotic RNA Splicing Different mRNA can be produced; different a-tropomyosin can be produced. Alternative splicing is a mechanism for gene regulation. Gene product can be different in different cell types and at different stages of development. Intron Splicing: Conserved Sequences exons - coding sequences introns - noncoding sequences Small nuclear ribonucleoprotein particles (snRNPs) recognize consensus splice junction sequence of GU/AG. snRNPs are complexes of protein and small nuclear RNA (snRNA). Several snRNPs comprise a spliceosome. Spliceosome directs the removal of introns and joining of exons. Spliceosome Assembly and Function Spliceosome interacts with CTD and attaches to pre-mRNA. snRNAs in spliceosomes direct alignment of the splice sites. One end of conserved sequence attaches to conserved adenine in the intron. The “lariat” is released and adjacent exons are joined. Reactions in Exon Splicing Self-Splicing Reaction RNA molecules can act somewhat like enzymes (ribozymes). In the protozoan Tetrahymena, the primary transcript of an rRNA can excise a 413-nucleotide intron from itself. These self-splicing introns are an example of RNA that can catalyze a reaction. Transfer of Information in Prokaryotes and Eukaryotes What are the similarities? What are the differences? Colinearity of Gene and Protein genotype DNA DNA sequence transcription RNA translation protein function phenotype organism amino acid sequence Colinearity of Gene and Protein “The linear sequence of nucleotides in a gene determines the linear sequence of amino acids in a protein.” Mutant alleles of trpA gene differed in the position of the mutation at the DNA level, which corresponded to position of amino acid substitution in the gene product. Colinearity of mutations and altered amino acids in a subunit of tryptophan synthetase from E. coli C. Yanofsky, 1967. Scientific American Tryptophan Synthetase Activity indole-3-glycerol phosphate a subunit glyceraldehyde 3-phosphate indole serine b subunit tryptophan Molecular Basis for Relationship between Genotype and Phenotype genotype DNA DNA sequence transcription RNA translation protein function phenotype organism amino acid sequence tRNA Anticodon of a tRNA molecule recognizes and pairs with an mRNA codon. tRNA contains modified bases: pseudouridine, methylguanosine, dimethylguanosine, methylinosine, dihydrouridine. Genetic Code Aminoacyl-tRNA Synthetase Attaches Amino Acid to tRNA Aminoacyl-tRNA synthetase catalyzes the formation of “charged” tRNA. There is an aminoacyltRNA synthetase for each amino acid. The carboxyl end of an amino acid is attached to the 3’ end of the tRNA. Wobble Position Some tRNA molecules can recognize and pair with more than one specific codon. Base-pairing between the 3’ base of a codon and 5’ base of an anticodon is not always exact.