* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download (Rfg, Rbg), (Gfg, Gbg)
Public health genomics wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Oncogenomics wikipedia , lookup
Minimal genome wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Metagenomics wikipedia , lookup
Genome evolution wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
History of genetic engineering wikipedia , lookup
Genome (book) wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Genomic imprinting wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Microevolution wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Designer baby wikipedia , lookup
Gene expression programming wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Ridge (biology) wikipedia , lookup
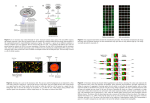
BIOINFORMATICS Lecture 8 Analyzing Microarray Data Dr. Aladdin Hamwieh Khalid Al-shamaa Abdulqader Jighly Aleppo University Faculty of technical engineering Department of Biotechnology 2010-2011 Microarray can monitor many genes at once, a DNA microarray is an inert, solid, flat and transparent surface (e.g.: a microscopic slide) onto which 20,000 to 60,000 short DNA probes of specified sequences are orderly tethered. Each probe corresponds to a particular short section of a gene. So a single gene is covered by several probes which span different parts of the gene sequence. REPOSITORIES OF MICROARRAY STUDIES Due to the large use of microarrays, data repositories have flourished world-wide. Three of the largest databases of gene expression are: 1. The Gene Expression Omnibus (GEO) 2. National Center for Biotechnology Information (NCBI) 3. Stanford Microarray Data Base (SMD) And for PLANTS Plant Expression database PLEXdb DNA microarrays measure the RNA abundance with either 1 channel (one color) or 2 channels (two colors). Affymetrix GeneChip has 1 channel and use either fluorescent red dye Cy5 or green fluorescent dye, Cy3 Stanford microarrays measure by competitive hybridization the relative expression under a given condition (fluorescent red dye Cy5) compared to its control (labeled with a green fluorescent dye, Cy3) (Two channels) Biological question Differentially expressed genes Sample class prediction etc. Experimental design Microarray experiment 16-bit TIFF files Image analysis (Rfg, Rbg), (Gfg, Gbg) Normalization R, G Estimation Testing Clustering Biological verification and interpretation Discrimination Biological question Differentially expressed genes Sample class prediction etc. Microarray experiment 16-bit TIFF files Image analysis (Rfg, Rbg), (Gfg, Gbg) Normalization R, G Estimation Testing Clustering Biological verification and interpretation Discrimination VIDEO Biological question Differentially expressed genes Sample class prediction etc. Experimental design 16-bit TIFF files Image analysis (Rfg, Rbg), (Gfg, Gbg) Normalization R, G Estimation Testing Clustering Biological verification and interpretation Discrimination MICROARRAY EXPERIMENT 1. 2. 3. 4. 5. 6. 7. Isolate mRNA Make labelled cDNA library Apply your DNA on the slide Scan the slide Purify the picture Extract the data Analyse your data RESULTS The colors denote the degree of expression in the experimental versus the control cells. Gene not expressed in control or in experimental cells Only in control cells Mostly in control cells Same in both cells Mostly in experimental cells Only in experimental cells Let us talk about the analysis and the mathematical problems: Now we have a lot of pictures which contain a huge information so: 1- we have to purify the picture 2- we have to extract our data. Biological question Differentially expressed genes Sample class prediction etc. Experimental design Microarray experiment 16-bit TIFF files (Rfg, Rbg), (Gfg, Gbg) Normalization R, G Estimation Testing Clustering Biological verification and interpretation Discrimination IMAGE ANALYSIS The raw data from a cDNA microarray experiment consist of pairs of image files, 16-bit TIFFs, one for each of the dyes. Image analysis is required to extract measures of the red and green fluorescence intensities for each spot on the array. STEPS IN IMAGE ANALYSIS 1. Addressing. Estimate location of spot centers. 2. Segmentation. Classify pixels as foreground (signal) or background. 3. Information extraction. For each spot on the array and each dye • foreground intensities; • background intensities; • quality measures. WHY DO WE CALCULATE THE BACKGROUND INTENSITIES? Motivation behind background adjustment: A spot’s measured fluorescence intensity includes a contribution that is not specifically due to the hybridization of the target to the probe, but to something else, e.g. the chemical treatment of the slide, autofluorescence etc. Want to estimate and remove this unwanted contribution. QUANTIFICATION OF EXPRESSION For each spot on the slide we calculate Red intensity = Rfg - Rbg fg = foreground, bg = background, and Green intensity = Gfg – Gbg CDNA GENE EXPRESSION DATA Data on p genes for n samples down-regulated gene Genes 1 2 3 4 5 Up-regulated gene unchanged sample1 sample2 sample3 sample4 sample5 … expression 0.46 0.30 0.80 1.51 0.00 ... -0.10 0.49 0.24 0.06 0.46 ... 0.15 0.74 0.04 0.10 0.20 ... -0.45 -1.03 -0.79 -0.56 -0.32 ... -0.06 1.06 1.35 1.09 -1.09 ... mRNA samples Gene expression level of gene 5 in mRNA sample 4 = log2( Red intensity / Green intensity) Biological question Differentially expressed genes Sample class prediction etc. Experimental design Microarray experiment 16-bit TIFF files Image analysis (Rfg, Rbg), (Gfg, Gbg) R, G Estimation Testing Clustering Biological verification and interpretation Discrimination NORMALIZATION Why? To correct for systematic differences between samples on the same slide, or between slides, which do not represent true biological variation between samples for example: 1. Dyes activity 2. Dyes quantity 3. scanning parameters 4. location on the array 5. Air bubbles SELF-SELF HYBRIDIZATIONS How do we know it is necessary? By examining self-self hybridizations, we label one sample from the same tissue with two dyes Cy3 , Cy5 so We find dye biases. Biological question Differentially expressed genes Sample class prediction etc. Experimental design Microarray experiment 16-bit TIFF files Image analysis (Rfg, Rbg), (Gfg, Gbg) Normalization R, G Biological verification and interpretation HOMOGENEITY AND SEPARATION PRINCIPLES Homogeneity: Elements within a cluster are close to each other Separation: Elements in different clusters are further apart from each other …clustering is not an easy task! Given these points a clustering algorithm might make two distinct clusters as follows BAD CLUSTERING This clustering violates both Homogeneity and Separation principles Close distances from points in separate clusters Far distances from points in the same cluster GOOD CLUSTERING This clustering satisfies both Homogeneity and Separation principles CLUSTERING TECHNIQUES Agglomerative: Start with every element in its own cluster, and iteratively join clusters together Divisive: Start with one cluster and iteratively divide it into smaller clusters Hierarchical: Organize elements into a tree, leaves represent genes and the length of the pathes between leaves represents the distances between genes. Similar genes lie within the same subtrees HIERARCHICAL CLUSTERING 4 5 1 2 3 6 7 9 1 2 3 4 5 6 7 9 8 8 HIERARCHICAL CLUSTERING ALGORITHM 1. 2. 3. 4. 5. 6. 7. 8. 9. 10. 11. Hierarchical Clustering (d , n) Form n clusters each with one element Construct a graph T by assigning one vertex to each cluster while there is more than one cluster Find the two closest clusters C1 and C2 Merge C1 and C2 into new cluster C with |C1| +|C2| elements Compute distance from C to all other clusters Add a new vertex C to T and connect to vertices C1 and C2 Remove rows and columns of d corresponding to C1 and C2 Add a row and column to d corrsponding to the new cluster C return T The algorithm takes a nxn distance matrix d of pairwise distances between points as an input. K-MEANS CLUSTERING PROBLEM: FORMULATION Input: A set, V, consisting of n points and a parameter k Output: A set X consisting of k points (cluster centers) that minimizes the squared error distortion d(V,X) over all possible choices of X 1-MEANS CLUSTERING PROBLEM: AN EASY CASE Input: A set, V, consisting of n points Output: A single points x (cluster center) that minimizes the squared error distortion d(V,x) over all possible choices of x 1-MEANS CLUSTERING PROBLEM: AN EASY CASE Input: A set, V, consisting of n points Output: A single points x (cluster center) that minimizes the squared error distortion d(V,x) over all possible choices of x 1-Means Clustering problem is easy. However, it becomes very difficult (NP-complete) for more than one center. An efficient heuristic method for K-Means clustering is the Lloyd algorithm K-MEANS CLUSTERING: LLOYD ALGORITHM expression in condition 2 5 4 x1 3 x2 2 1 x3 0 0 1 2 3 4 expression in condition 1 5 K-MEANS CLUSTERING: LLOYD ALGORITHM 1. 2. 3. 4. 5. Lloyd Algorithm Arbitrarily assign the k cluster centers while the cluster centers keep changing Assign each data point to the cluster Ci corresponding to the closest cluster representative (center) (1 ≤ i ≤ k) After the assignment of all data points, compute new cluster representatives according to the center of gravity of each cluster, that is, the new cluster representative is ∑v \ |C| for all v in C for every cluster C *This may lead to merely a locally optimal clustering. THANK YOU