* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Ubiquitin-proteosome protein degradation ppt
Biochemistry wikipedia , lookup
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Mitogen-activated protein kinase wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Ultrasensitivity wikipedia , lookup
Gene regulatory network wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Metalloprotein wikipedia , lookup
Biochemical cascade wikipedia , lookup
Gene expression wikipedia , lookup
Magnesium transporter wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Signal transduction wikipedia , lookup
Expression vector wikipedia , lookup
Protein structure prediction wikipedia , lookup
Paracrine signalling wikipedia , lookup
Bimolecular fluorescence complementation wikipedia , lookup
Interactome wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Protein purification wikipedia , lookup
Western blot wikipedia , lookup
Protein–protein interaction wikipedia , lookup
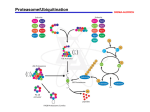
Intracellular Protein Degradation Chris Weihl MD/PhD [email protected] Department of Neurology How is trash handled? Protein Degradation in the Cell Ub Autophagy Nucleus Aggresome Ub UPS Ub Ub Endocytosis Consequence of impaired protein degradation • Protein aggregates • Ubiquitinated inclusions • Vacuolation • Damaged organelles • Secondary impairment in other cellular processes • Cell Death • Underlying pathogenesis of degenerative disorders (neurodegeneration, muscle degeneration, liver degeneration, lung disease, aging) Protein Degradation Turnover of protein is NOT constant Half lives of proteins vary from minutes to infinity “Normal” proteins – 100-200 hrs Short-lived proteins regulatory proteins enzymes that catalyze committed steps transcription factors Long-lived proteins Special cases (structural proteins, crystallins) Protein Degradation • May depend on tissue distribution Example: Lactic Acid Dehydrogenase Tissue Half-life Heart 1.6 days Muscle 31 days Liver 16 days • Protein degradation is a regulated process Example: Acetyl CoA carboxylase Nutritional state Half-life Fed 48 hours Fasted 18 hours Protein Degradation Ubiquitin/Proteasome Pathway 80-90% Most intracellular proteins • Lysosomal processes 10-20% Extracellular proteins Cell organelles Some intracellular proteins How are proteins selected for degradation? UBIQUITIN Small peptide that is a “TAG” 76 amino acids C-terminal glycine - isopeptide bond with the e-amino group of lysine residues on the substrate Attached as monoubiquitin or polyubiquitin chains G K Ubiquitination of proteins is a FOUR-step process First, Ubiquitin is activated by forming a link to “enzyme 1” (E1). Then, ubiquitin is transferred to one of several types of “enzyme 2” (E2). Then, “enzyme 3” (E3) catalizes the transfer of ubiquitin from E2 to a Lys e-amino group of the “condemned” protein. Lastly, molecules of Ubiquitin are commonly conjugated to the protein to be degraded by E3s & E4s AMP The UPS is enormous! The UPS is enormous! The genes of the UPS constitutes ~5% of the The genes of the UPS constitutes ~5% of the genome genome E1’s- 1-2 activating enzymes • E1’s1-210-20 activating enzymesenzymes E2’sconjugating • E2’s10-20 conjugating enzymes E3’s500-800 ubiquitin ligase- drives specificity • E3’s500-800 specificity DUBs100ubiquitin ubiquitin ligasespecificdrives proteasesregulators of pathway • DUBs- 100 ubiquitin specific proteases- regulators of pathway PROTEASOME COMPONENTS 20S Proteasome 19S Particle ATP 26S Proteasome Hydrolysis peptide bonds after: hydrophobic a.a. = CHYMOTRYPSINLIKE - 5 acidic a.a. = (-) CASPASE-LIKE -1 basic a.a. = (+) TRYPSIN-LIKE -2 DEUBIQUITINATION De-ubiquitinating Pathways controlled by regulated proteolysis Mechanism of muscle atrophy MURF/Atrogin Knockout of Atrogin Rescues atrophy 00 0 00 00 00 1: 1 1: 1 la ct la ct la ct tre at m en t 1: 1 no Relative Light Units 900000 800000 700000 600000 500000 400000 300000 200000 100000 0 Proteasome inhibition increases Usp14 ubiquitin-hydrolase activity Usp14 Uch37 Borodovsky, A et al EMBO J. 20:5187-96 2001 The proteasomal DUB Usp14 impairs protein degradation Lee, BH et al Nature 467:179-84 2010 Decrease steady-state levels of aggregate prone proteins in the absence of Usp14 Lee, BH et al Nature 467:179-84 2010 Lyosomal degradation • Autophagy Autophagy • Lysosomal degradation of proteins and organelles • Occurs via three routes • Macroautophagy • Microautophagy (direct uptake of cellular debris via the lysosome) • Chaperone mediated autophagy (selective import of substrates via Hsc70 and Lamp2a) Yeast Genetics meets Human Genetics • Identification of >50 autophagy essential proteins with mammalian homologs Macroautophagy Lysosome FOXO3 Beclin ATG7 mTOR ATG5-ATG12-ATG16L1 Induction Nucleation Phagophore Autophagosome Sequestration Trafficking & Cargo loading “Autophagic Flux” Autolysosome Fusion Degradation Genetic knockout of autophagy initiating proteins Complete loss of ATG5 leads to lethality Tissue specific knockout of autophagy • Degeneration of CNS tissue; Hara et al 2006 • Hepatomegaly in Liver; Komatsu et al 2005 • Atrophy and weakness of skeletal muscle; Masiero et al 2009 • Pathologic similarities • Ubiquitinated inclusions • Aberrant mitochondria • Oxidatively damaged protein Basal Autophagy • Autophagy has a “housekeeping” role in the maintenance of cellular homeostasis • Autophagy is responsible for the clearance of ubiquitinated proteins Selective Autophagy • Aggregaphagy– p62/SQSTM1, Nbr1 • Mitophagy – Parkin, Nix • Reticulophagy – endoplasmic reticulum • Ribophagy – translating ribosomes • Xenophagy – e.g. Salmonella via optineurin • Lipophagy – autophagy mediated lipolysis • Performed by an expanding group of ubiquitin adaptors p62 as an autophagic tool • p62 associates with ubiquitinated proteins and LC3 • p62 is an autophagic substrate LC3 as an autophagic tool LC3-I (18kD) LC3-II (16kD) GFP-LC3 starved IBMPFD myopathy LC3II protein levels (A.U) p62 protein levels (A.U) 2 1 0 2 1 0 Con WT RH9 RH12 Con WT RH9 RH12 Ju et al, JCB 2009 Ju et al, JCB 2009 Upregulation of functional autophagosomes Decrease in autophagosome degradation or “autophagic flux” Phagophore closure Autophagosome-lysosome fusion Absence of functional lysosomes VCP Ju et al, JCB 2009 Ub Nucleus Immunosuppressant used to treat transplant rejection Inhibits the mTOR pathway mTOR integrates extrinsic growth signals and cellular nutrient status and energy state Active mTOR Protein synthesis and cell growth Inactive mTOR (or rapamycin treatment) Inhibition of protein synthesis and increased autophagic degradation of protein Ub Ub Increase autophagic stimulus Nucleus Depending upon the disease, stimulating or inhibiting autophagy may be appropriate. Identifying drugs that “facilitate” autophagy.