* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download VirusEvoution2005
Human genome wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Metagenomics wikipedia , lookup
Epigenetics of human development wikipedia , lookup
History of genetic engineering wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Non-coding DNA wikipedia , lookup
Polyadenylation wikipedia , lookup
Genome evolution wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
RNA interference wikipedia , lookup
Viral phylodynamics wikipedia , lookup
Epitranscriptome wikipedia , lookup
Primary transcript wikipedia , lookup
Nucleic acid tertiary structure wikipedia , lookup
Microevolution wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
History of RNA biology wikipedia , lookup
RNA silencing wikipedia , lookup
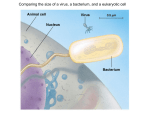
Ecology and Evolutionary Biology of Viruses SOME CONSEQUENCES AND EFFECTS OF VIRUS INFECTION • Like other life forms, viruses promote the propagation of their own kind • Like other life forms, viruses evolve in response to selection pressure • Viruses are major factors in promoting the evolution of higher organisms • Viruses help control populations of their hosts, including humans Host properties influence the virus types found in that host group • • • • Vertebrates have broad range of viruses Plants have mostly small RNA viruses Fungi have mostly dsRNA viruses Single-celled organisms have mostly large dsDNA viruses • True Fungi Overview of Virus Properties – RNA – 2.5-28 kb – DNA – none – Enveloped ones have no capsid – Little genome complexity – Little morphological complexity – Some divided genomes • Prokaryote – RNA – 5-8 kb – DNA – 10-200 kb – Few enveloped – Range of complexity – Range of morphologies – Few divided genomes • Animal – RNA – 5-30 kb – DNA: 5-350 kb – Many enveloped – Range of complexity – Range of morphologies – Some divided genomes • Plant – RNA – 0.3-28 kb – DNA – 3-10 kb – Few enveloped – Little genome complexity – Little morphological complexity – Many divided genomes • Lower eukaryote – RNA – 5-10 kb – DNA – 180-1200 kb – Internal envelope – Range of complexity – Range of morphologies – No divided genomes Virus transmission • Animal viruses are transmitted through air, through wounds, through orifices, or by vectors • Most plant viruses are transmitted by vectors, especially homopterous insects • Most fungal viruses are transmitted horizontally only by hyphal fusion • Bacterial viruses are transmitted by attachment of free virus to bacterial cell walls or pili; injection of nucleic acid • How do these transmission modes affect their ecology and evolutionary biology? Virus Evolution • Viruses origins are unknown • Theories of virus origin: – Regressive evolution: viruses degenerated from previously independent life forms, lost many functions required by cellular organisms – Cellular origins: viruses assembled from cellular components into independent entities capable of moving cell-to-cell and, later, gaining ability capacity for transmission – Independent entities: viruses evolved independently and in parallel with complex organisms from self-replicating molecules in the primordial RNA world • These theories are not mutually exclusive – different virus lineages may have different origins Virus evolution • Virus evolution is contemporary and observable • Large numbers of progeny contribute to potential high rate of evolution of viruses • Mutation rate is higher for RNA than for DNA • Evolution rate does not necessarily reflect mutation rate • Mutation rate for a particular virus may be different in different tissues • Different parts of viral genomes evolve at different rates • Ability to generate large amounts of sequence data has greatly enhanced ability to study evolution Sequence variation during virus replication • Intrinsic error rates of polymerases are difficult to quantify • DNA polymerase has proof-reading capability; intrinsic error rate is low, usually ~ 10-6 to 10-5 • RNA polymerase has no proof-reading capability; intrinsic error rate is high, usually ~ 10-4 to 10-3 • Error rates may be different in different genome regions, e.g., “hotspots” • Homologous or non-homologous recombination may occur in RNA or DNA viruses • Change of templates is referred to as copy choice Means of RNA virus evolution • Minor replication error (substitution, single base change) – no error correction by RdRp • Major replication error (deletion) • Intragenomic gene duplication • Intergenomic recombination – gene duplication – acquisition of additional genes – coding sequence or noncoding sequence substitution • Virus evolution may be accelerated by coinfection Positive-strand RNA virus evolution • Positive-strand RNA viruses evolve rapidly; only important functional domains are conserved • RNA viruses are made up of a limited number of building blocks; only RdRp is required and is the ultimate basis of rational phylogenies • Evolution of RNA viruses involves conservation of required genes and recombination/shuffling of gene blocks • Widespread recombination of a relatively small number of genes makes it impossible to generate single phylogenetic trees (reticulate evolution) • Basic RNA virus replication machinery likely evolved more than once Positive-strand RNA virus evolution A. Organization of conserved replication-associated genes of major groups of positive-sense RNA viruses; B. Phylogenetic reconstruction of selected alphaviruses based on RdRp gene. From your text: Flint et al., 2004 Viruses closely related to RNA bacteriophages (Leviviridae) Virus Location Host Capsid? OMV4 (2.6 kb) * CMV1/NB631 (2.7 kb) * SNV/23S (2.9 kb) * Qβ (3.6 kb) TBSV (4.8 kb) PEMV (4.2 kb) DRV (4.1 kb) * * * * Mito Fungus No Mito Fungus No Cyto Yeast No (Cyto) Bacterium Yes Cyto Plant Yes Cyto Plant No Cyto Fungus No = stop codon position * = core RNA-dependent RNA polymerase domain = capsid protein gene Genome organizations of negative-sense RNA viruses, and homologies among genes From text: Flint et al., 2004 Mechanisms of recombination of viral RNAs Left: Generalized viral RNA recombination. Bottom: Three classes of intergenomic RNA recombination: 1) Requiring substantial base pairing but no identifiable RNA secondary structures or regulatory elements; 2) Occurring in association with identifiable RNA ,structures or regulatory elements, but not requiring substantial base pairing; 3) A combination of 1 and 2. From your text: Flint et al., 2004 Variation among viral sequences • The term “quasi species” is used predominately for RNA viruses • Because of absence of proofreading, many variants are found in an RNA virus population; the “quasispecies cloud” is the mutant spectrum derived from the dominant master copy • A genetic bottleneck occurs when a virus population is constrained, resulting in loss of diversity – can be because of: – vector constraints – host defense constraints • A small founder population coming through a genetic bottleneck may give rise to a skewed population Viral quasispecies, population size, bottlenecks, and fitness From your text: Flint et al., 2004 Intrinsic mutation rates among RNA viruses vary In the absence of selection, spontaneous mutation rates of different viral RNA polymerases are high, and vary by ~10 fold. From Drake & Holland, 1999, PNAS 96:13909