* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Artemis as genome viewing and annotation tool
Gene expression programming wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Gene nomenclature wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Gene therapy wikipedia , lookup
DNA sequencing wikipedia , lookup
Genetic engineering wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Minimal genome wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Genome (book) wikipedia , lookup
Epigenomics wikipedia , lookup
Gene desert wikipedia , lookup
Gene expression profiling wikipedia , lookup
Copy-number variation wikipedia , lookup
Point mutation wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Public health genomics wikipedia , lookup
Transposable element wikipedia , lookup
Non-coding DNA wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
History of genetic engineering wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Microsatellite wikipedia , lookup
Human genome wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Microevolution wikipedia , lookup
Whole genome sequencing wikipedia , lookup
Designer baby wikipedia , lookup
Genomic library wikipedia , lookup
Genome evolution wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Human Genome Project wikipedia , lookup
Metagenomics wikipedia , lookup
Pathogenomics wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Genome editing wikipedia , lookup
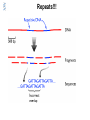
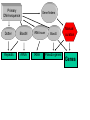
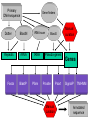
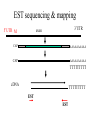
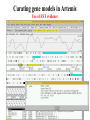
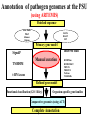
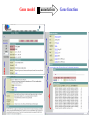
Sequence Analysis with Artemis & Artemis Comparison Tool (ACT) South East Asian Training Course on Bioinformatics Applied to Tropical Diseases - 2005 (Sponsored by UNDP/World Bank/WHO/TDR) International Centre For Genetic Engineering And Biotechnology , New Delhi, INDIA Workshop Overview Overview of the genome sequencing and sequence analysis. Demonstration of Artemis. Hands on guided exercise in Artemis. Demonstration of ACT . Hands on guided exercise in ACT Generating ACT comparison files The Wellcome Trust Sanger Institute •Funded by The Wellcome Trust, a registered charity. •Established in 1993 to begin the Human genome project. •First Draft (2000) complete (2003-4) Wellcome Trust Photo Library Data release policy: All sequence data is released immediately and is freely available via the internet in order to maximise its benefit for research. http://www.sanger.ac.uk ftp://ftp.sanger.ac.uk/ Wellcome Trust Photo Library Generating the complete genome sequence Infrastructure Levels of automation Colony picking robots Plasmid preps robots ABI3700 ABI3730 TOTAL:140 Automated sequencing Each ABI reads 96 DNA sequences at once. The machines are run 10 times a day, 7 days a week. Throughput of 1,200 to 1,300 96-well plates per day ± 120,000 DNA samples read each day. Each day, the Sanger Institute reads 60 million base pairs. That’s equal to one of the smaller human chromosomes and many times that of an average bacterial genome. Pathogen Sequencing Unit http://www.sanger.ac.uk/Projects/Microbes The Pathogen Group is funded by the Beowulf Genomics Initiative to sequence the genomes of a wide range of small Eukaryotes and microbes. Yeasts and Fungi: Saccharomyces cerevisiae Schizosaccharomyces pombe Aspergillus fumigatus Candida dubliniensis Candida parapsilosis Protozoa: Plasmodium falciparum X3 Plasmodium spp. X5 Leishmania spp. Trypanosoma spp. Eimeria Theileria Babesia Bacteria: M. tuberculosis M. leprae Y. pestis S. typhi C. Diphtheriae Bordetella spp. x3 B. pseudomallei S. aureus MRSA S. aureus MSSA E. carrotovora Sequencing strategy and assembly Shotgun sequencing – strategy DNA Contiguous sequence pUC clone end sequence physical gap sequence gap ‘Draft sequence’ Order of contigs? 95% coverage, 4-5x depth. ‘A genome in a day’ ‘15 in a month’ ‘High-quality draft sequence’ Shotgun sequencing – strategy DNA Contiguous sequence pUC clone end sequence physical gap sequence gap large clone end sequence Finished sequence: 100% coverage, 10x depth. Repeats!!! Shotgun assembly - Yersinia pestis Primary DNA sequence Gene finders Dotter BlastN tRNA scan Repeats rRNA tRNA BlastX Pseudo-genes Manual curation Genes Primary DNA sequence Gene finders Dotter BlastN tRNA scan Repeats rRNA tRNA Fasta BlastP Pfam BlastX Pseudo-genes Prosite Manual curation Psort Manual curation Genes SignalP TMHMM Annotated sequence PSU Projects Organism Database entry Finished genome Annotated genome Artemis Artemis • Sequence viewer and analysis tool – Visualization of sequence features • DNA • Six frame translation – Perform and view analysis • Basic analysis • Launch more complex analysis and searches • Import and view the results of other searches Outline of Artemis demonstration • • • • Artemis window features Open a genome sequence Changing the view Getting around – Goto Menu – Navigator – Feature Selector • Basic analysis – Edit a feature – Fasta search – Show feature plots Artemis Drop Down Menus Entry Button Line Main Sequence View Panel Sliders Magnified Sequence View Panel Feature Menu Sliders Artemis Curating gene models in Artemis Use of multiple lines of evidence Curating gene models in Artemis Use of FASTA evidence EST sequencing & mapping 5’UTR M intron exon stop 3’UTR CAP AAAAAAAAAA CAP AAAAAAAAAA mRNA TTTTTTTTT cDNA TTTTTTTTT EST EST Curating gene models in Artemis Use of EST evidence ESTs Curating gene models in Artemis Use of EST evidence Curation of gene models in Artemis Mapping proteome fragments to genome Curation and annotation in Artemis Mapping InterPro domain hits to genome Annotation of pathogen genomes at the PSU (using ARTEMIS) Finished sequence Gene Finder PHAT Glimmer Orpheus FASTA BLAST EST Primary gene model InterPro scan SignalP Manual curation TMHMM t-RNA scan HMMPfam HMMSMART PRINTS PROSITE ProDom TIGRFAMs Refined gene model Functional classification (GO / Riley) Organism-specific gene families Comparative genomics (using ACT) Complete Annotation Gene model annotation Gene function Top tips! Manual annotation. Use a several lines of evidence: - Run several available gene finding programs - Search programs: local (BLAST) and global (FASTA) alignments -Protein domains and motifs: Interpro (Pfam, prosite, SMART etc.) -Transmembrane / signal peptide prediction (TMHMM, SignalP) - Base your annotation on characterised proteins where possible (e.g. UNIPROT entry) - Read the literature (Pubmed entry) Sanger Front page