* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Instructions for FUEL-mLoc Web-server
Survey
Document related concepts
SNARE (protein) wikipedia , lookup
Gene expression wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Magnesium transporter wikipedia , lookup
Gene regulatory network wikipedia , lookup
Paracrine signalling wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Protein adsorption wikipedia , lookup
Protein moonlighting wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Cell membrane wikipedia , lookup
Cell-penetrating peptide wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Intrinsically disordered proteins wikipedia , lookup
Western blot wikipedia , lookup
Transcript
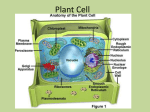
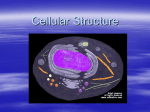
Instructions for FUEL-mLoc Server Instructions for FUEL-mLoc Web-server Shibiao Wan∗,‡ and Man-Wai Mak‡ email: [email protected], [email protected] ∗ ‡ Princeton University, NJ, USA The Hong Kong Polytechnic University, Hong Kong SAR, China August 2016 Back to FUEL-mLoc Server Contents 1 Significance of Subcellular Localization Prediction 2 2 Major organelles in a typical eukaryotic cell 2 3 Specific information about FUEL-mLoc 4 4 Some Notes 7 1 Instructions for FUEL-mLoc Server 1 Significance of Subcellular Localization Prediction Proteins must be transported to the correct organelles of a cell and folded into correct 3-D structures to properly perform their functions. Therefore, knowing the subcellular localization is one step towards understanding its functions. Proteins can exist in different locations within a cell, and some proteins can even simultaneously reside at, or move between, two or more different subcellular locations. As an essential and indispensable topic in proteomics research and molecular cell biology, protein subcellular localization is critically important for protein function annotation, drug target discovery, and drug design. Efficient and reliable computational methods are developed to assist the biological experiments such as fluorescent microscopy imaging. Proteins with multiple locations play important roles in some metabolic processes taking place in more than one compartment. 2 Major organelles in a typical eukaryotic cell Most of the biological activities performed by proteins occur in organelles. An organelle is a cellular component or subcellular location within a cell that has specific functions. Fig. 1 illustrates some organelles in a typical eukaryotic cell. (Note that human cells and plant cells are two subsets of eukaryotic cells.) In eukaryotic cells, major organelles include cytoplasm, mitochondria, chloroplast, nucleus, extracellular space, endoplasmic reticulum (ER), Golgi apparatus and plasma membrane. Cytoplasm takes up most of the cell volume where most of the cellular activities, such as cell division and metabolic pathways, occur. Mitochondrion is a membrane-bound organelle found in most eukaryotic cells. It is mainly responsible for supplying energy for cellular activities. Chloroplast is 2 Instructions for FUEL-mLoc Server Figure 1: Organelles or subcellular locations in a typical eukaryotic cell. Major eukaryotic organelles include cytoplasm, mitochondria, chloroplast, nucleus, extracellular space, endoplasmic reticulum, Golgi apparatus and plasma membrane. an organelle existing in plant or algal cells. Its role is to conduct photosynthesis to store energy from sunlight. Nucleus is a membrane-enclosed organelle containing most of the genetic materials for a cell. Its main function is to control the activities of the cell by regulating gene expression. Extracellular space refers to the space outside the plasma membrane, which is occupied by fluid. ER is a type of organelle that forms an interconnected membranous network of cistemae which serves the functions of folding protein molecules in cistemae and transporting synthesized proteins to Golgi apparatus. 3 Instructions for FUEL-mLoc Server Golgi apparatus is an organelle which is particularly important in cell secretion. Plasma membrane or cell membrane is a biological membrane that separates the intracellular environment from extracellular space. Its basic function is to protect the cell from its surroundings. Some proteins locate in peroxisome, vacuole, cytoskeleton, nucleoplasm, lysosome, acrosome, cell wall, centrosome, cyanelle, endosome, hydrogenosome, melanosome, microsome, spindle pole body, synapse, etc. For Gram-positive bacteria species, their proteins are usually located in cell membrane, cell wall, cytoplasm and extracellular space. For Gram-negative bacteria species, their proteins are located in eight subcellular locations, including cell inner membrane, cell outer membrane, cytoplasm, extracellular space, fimbrium, flagellum, nucleus and periplasm. For the virus species, viral proteins are usually located within host cells, which are distributed in subcellular locations such as host cytoplasm, host nucleus, host cell membrane, host ER, host nucleus as well as viral capsid. 3 Specific information about FUEL-mLoc FUEL-mLoc is an interpretable multi-label predictor which uses unified features to yield sparse and interpretable solutions for large-scale prediction of both single-label and multilabel proteins of different species, including eukaryota, human, plant, Gram-positive bacteria. Gram-negative bacteria and virus. Given a query protein sequence in a particular species, a set of GO terms are retrieved from a newly created compact databases, namely ProSeq-GO. The frequencies of GO occurrences are used to formulate frequency 4 Instructions for FUEL-mLoc Server vectors with dimensionality of more than eight thousand. By using the one-vs-rest ENbased (elastic net-based) classifiers, much fewer GO terms are selected. Subsequently, the dimension-reduced feature vectors are classified by a multi-label EN classifier. Based on the selected essential GO terms, the user of FUEL-mLoc can not only determine where a protein resides, but also explains why it is located there. Similar to R3P-Loc [1, 2, 3, 4], instead of using the Swiss-Prot and GOA databases as previous predictors [5, 6, 7, 8, 9, 10, 11], FUEL-mLoc uses two newly-created compact databases, namely ProSeq and ProSeq-GO, for GO information transfer. The ProSeq database is a sequence database in which each amino acid sequence has at least one GO term annotated to it. The ProSeq-GO comprises GO terms annotated to the protein sequences in the ProSeq database. An important property of the ProSeq and ProSeqGO databases is that they are much smaller than the Swiss-Prot and GOA databases, respectively. FUEL-mLoc is designed to be able to predict subcellular localization of proteins from six species, namely eukaryota, human, plant, Gram-positive bacteria, Gram-negative bacteria and virus. • For eukaryotic proteins, 22 subcellular locations can be predicted, including: (1) acrosome; (2) cell membrane; (3) cell-wall; (4) centrosome; (5) chloroplast; (6) cyanelle; (7) cytoplasm; (8) cytoskeleton; (9) endoplasmic reticulum; (10) endosome; (11) extracellular; (12) golgi apparatus; (13) hydrogenosome; (14) lysosome; (15) melanosome; (16) microsome; (17) mitochondrion; (18) nucleus; (19) peroxisome; (20) spindle pole body; (21) synapse; and (22) vacuole. The predictor is not designed for predicting the subcellular localization of non-eukaryotic proteins 5 Instructions for FUEL-mLoc Server when selecting predicting the eukaryotic proteins. Therefore, the prediction results of non-eukaryotic proteins in this case are arbitrary and meaningless. • For human proteins, 14 subcellular locations can be predicted, including: (1) centrosome; (2) cytoplasm; (3) cytoskeleton; (4) endoplasmic reticulum; (5) endosome; (6) extracellular; (7) Golgi apparatus; (8) lysosome; (9) microsome; (10) mitochondrion; (11) nucleus; (12) peroxisome; (13) plasma membrane; and (14) synapse. The predictor is not designed for predicting the subcellular localization of non-human proteins. Therefore, the prediction results of non-human proteins are arbitrary and meaningless. • For plant proteins, 12 subcellular locations can be predicted, including: (1) cell membrane; (2) cell wall; (3) chloroplast; (4) cytoplasm; (5) endoplasmic reticulum; (6) extracellular; (7) golgi apparatus; (8) mitochondrion; (9) nucleus; (10) peroxisome; (11) plastid; and (12) vacuole. Note (11) plastid here includes those plastid groups except for (3) chloroplast. The predictor is not designed for predicting the subcellular localization of non-plant proteins when selecting predicting the plant proteins. Therefore, the prediction results of non-plant proteins in this case are arbitrary and meaningless. • For Gram-positive bacterial proteins, 4 subcellular locations can be predicted, including: (1) cell membrane; (2) cell wall; (3) cytoplasm; and (4) extracellular space. The predictor is not designed for predicting the subcellular localization of non-plant proteins when selecting predicting the plant proteins. Therefore, the prediction results of non-plant proteins in this case are arbitrary and meaningless. 6 Instructions for FUEL-mLoc Server • For Gram-negative bacterial proteins, 8 subcellular locations can be predicted, including: (1) cell inner membrane; (2) cell outer membrane; (3) cytoplasm; (4) extracellular space; (5) fimbrium; (6) flagellum; (7) nucleus; and (8) periplasm. The predictor is not designed for predicting the subcellular localization of non-Gramnegative-bacterial proteins when selecting predicting the plant proteins. Therefore, the prediction results of non-plant proteins in this case are arbitrary and meaningless. • For viral proteins, 6 subcellular locations can be predicted, including: (1) viral capsid; (2) host cell membrane; (3) host endoplasmic reticulum; (4) host cytoplasm; (5) host nucleus; (6) secreted. The predictor is not designed for predicting the subcellular localization of non-viral proteins. Therefore, the prediction results of non-viral proteins are arbitrary and meaningless. 4 Some Notes The following notes for FUEL-mLoc web-server should be paid attention to: 1. FUEL-mLoc can make prediction for the input of amino acid sequences of proteins in FASTA format. 2. Users can optionally provide email addresses to receive the prediction results. If no (or wrongly-formatted) email address is provided, the results will be shown on the webpage. On the other hand, if an email address is provided, the prediction results will be sent via email. For users’ convenience, the prediction results are still downloadable from the webpage even if the results are sent to their emails. 7 Instructions for FUEL-mLoc Server 3. FUEL-mLoc is an interpretable predictor and the interpretations for a particular query protein (or a list of proteins) are presented in a formatted table in an HTML file which can also be downloadable. 4. Because FUEL-mLoc uses two new compact databases, namely ProSeq and ProSeqGO, instead of Swiss-Prot and GOA, it consumes much less memory and predicts faster. References [1] S. Wan, M. W. Mak, and S. Y. Kung, “Sparse regressions for predicting and interpreting subcellular localization of multi-label proteins,” BMC Bioinformatics, vol. 17, no. 97, 2016. [2] S. Wan, M. W. Mak, and S. Y. Kung, “Mem-ADSVM: a two-layer multi-label predictor for identifying multi-functional types of membrane proteins,” Journal of Theoretical Biology, vol. 398, pp. 32–42, 2016. [3] S. Wan, M. W. Mak, and S. Y. Kung, “Mem-mEN: predicting multi-functional types of membrane proteins by interpretable elastic nets,” IEEE/ACM Transactions on Computational Biology and Bioinformatics, vol. 13, pp. 706–718, 2016. [4] S. Wan, M. W. Mak, and S. Y. Kung, “R3P-Loc: A compact multi-label predictor using ridge regression and random projection for protein subcellular localization,” Journal of Theoretical Biology, vol. 360, pp. 34–45, 2014. 8 Instructions for FUEL-mLoc Server [5] K. C. Chou, Z. C. Wu, and X. Xiao, “iLoc-Hum: using the accumulation-label scale to predict subcellular locations of human proteins with both single and multiple sites,” Molecular BioSystems, vol. 8, pp. 629–641, 2012. [6] S. Wan, M. W. Mak, and S. Y. Kung, “mGOASVM: Multi-label protein subcellular localization based on gene ontology and support vector machines,” BMC Bioinformatics, vol. 13, pp. 290, 2012. [7] S. Wan, M. W. Mak, and S. Y. Kung, “GOASVM: A subcellular location predictor by incorporating term-frequency gene ontology into the general form of Chou’s pseudoamino acid composition,” Journal of Theoretical Biology, vol. 323, pp. 40–48, 2013. [8] S. Wan, M. W. Mak, and S. Y. Kung, “HybridGO-Loc: Mining hybrid features on gene ontology for predicting subcellular localization of multi-location proteins,” PLoS ONE, vol. 9, no. 3, pp. e89545, 2014. [9] S. Wan, M. W. Mak, and S. Y. Kung, “mPLR-Loc: An adaptive decision multi-label classifier based on penalized logistic regression for protein subcellular localization prediction,” Analytical Biochemistry, vol. 473, pp. 14–27, 2015. [10] S. Wan and M. W. Mak, Machine learning for protein subcellular localization prediction, De Gruyter, 2015. [11] S. Wan and M. W. Mak, “Predicting subcellular localization of multi-location proteins by improving support vector machines with an adaptive-decision scheme,” International Journal of Machine Learning and Cybernetics, pp. 1–13. 9