* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Direct Interaction between Survivin and Smac/DIABLO Is Essential
Tissue engineering wikipedia , lookup
Cell growth wikipedia , lookup
Endomembrane system wikipedia , lookup
Extracellular matrix wikipedia , lookup
Cell culture wikipedia , lookup
Cellular differentiation wikipedia , lookup
Cell encapsulation wikipedia , lookup
Green fluorescent protein wikipedia , lookup
Organ-on-a-chip wikipedia , lookup
Signal transduction wikipedia , lookup
Cytokinesis wikipedia , lookup
Programmed cell death wikipedia , lookup
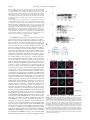
THE JOURNAL OF BIOLOGICAL CHEMISTRY © 2003 by The American Society for Biochemistry and Molecular Biology, Inc. Vol. 278, No. 25, Issue of June 20, pp. 23130 –23140, 2003 Printed in U.S.A. Direct Interaction between Survivin and Smac/DIABLO Is Essential for the Anti-apoptotic Activity of Survivin during Taxol-induced Apoptosis* Received for publication, January 29, 2003, and in revised form, March 19, 2003 Published, JBC Papers in Press, March 26, 2003, DOI 10.1074/jbc.M300957200 Zhiyin Song, Xuebiao Yao, and Mian Wu‡ From the Department of Molecular and Cell Biology, Key Laboratory of Structural Biology, School of Life Sciences, University of Science and Technology of China, Hefei, Anhui 230027, People’s Republic of China Survivin is a member of the inhibitor of apoptosis protein (IAP) family that has been implicated in both apoptosis inhibition and cell cycle control. However, its inhibitory mechanism and subcellular localization remain controversial. In this report, we provided evidence for the first time that Survivin physically interacts with Smac/DIABLO both in vitro and in vivo. A point mutation (D71R) in the baculovirus IAP repeat motif and a C-terminal deletion mutant (Surv-BIR) of Survivin fail to bind to Smac/DIABLO and abrogate its ability to inhibit apoptosis. The N-terminal of mature Smac/DIABLO is absolutely required for Survivin䡠Smac complex formation. Subcellular distributions of Survivin and Smac/ DIABLO showed that they co-localized within the cytosol during interphase. In addition, Survivin was found to be incapable of binding to caspase. We also identified that the co-presence of Smac/DIABLO and XIAP was required for Survivin to inhibit caspase cleavage in a cell-free system. In conclusion, our results provide the first evidence that the interaction between Smac/DIABLO and Survivin is an essential step underling the inhibition of apoptosis induced by Taxol. Among the regulators of apoptosis, considerable interest has been focusing on the inhibitors of apoptosis protein (IAP)1 family, which were first identified as negative regulators in programmed cell death characterized by the presence of one to three copies of the baculovirus IAP repeat (BIR) domain (1). However, anti-apoptotic function in physiological cell death has not been established for all IAPs, because some members in this family play essential roles in cell division, rather than merely acting as the regulators of apoptosis (2). Survivin (16.5 KDa) is a special member of the inhibitor-of-apoptosis proteins (IAPs) family containing a single BIR and lacking a RING finger motif. It is mostly expressed in the vast majority of tumors or in the embryonic development, but not in normal * This research was supported by the Key Project Fund (KSCX2-201-004), a special grant (to M. W.) from the Chinese Academy of Sciences, the National Natural Science Foundation of China (Grants 30121001 and 90208027), and a 973 grant (Grant 2002CB713700) from the Ministry of Science and Technology of China. The costs of publication of this article were defrayed in part by the payment of page charges. This article must therefore be hereby marked “advertisement” in accordance with 18 U.S.C. Section 1734 solely to indicate this fact. ‡ To whom correspondence should be addressed. Tel.: 86-551-3606264; Fax: 86-551-360-6264; E-mail: [email protected]. 1 The abbreviations used are: IAP, inhibitor of apoptosis protein; BIR, baculovirus IAP repeat; GST, glutathione S-transferase; Ab, antibody; GFP, green fluorescence protein; EGFP, enhanced GFP; BSA, bovine serum albumin; PBS, phosphate-buffered saline; FITC, fluorescein isothiocyanate; cyto c, cytochrome c; TRAIL, tumor necrosis factor-related apoptosis-inducing ligand. adult tissues (3, 4). Survivin is cell cycle-regulated and predominantly expressed in the G2/M phase (5). The degradation of Survivin is believed to be through the ubiquitin-proteasome pathway (6). Accumulating evidence supports the idea that Survivin is a bifunctional protein that acts as a cell division regulator (7–14) and an apoptosis suppressor (3, 5, 15, 16). The mechanism of how Survivin inhibits apoptosis remains controversial. Tamm et al. (16) demonstrated that Survivin was able to bind to the effector caspase-3 and caspase-7 in vitro and proposed that Survivin may inhibit caspase activity in physiological cell death by the similar mechanism. However, comparison between the x-ray crystallographic structures of Survivin and that of the XIAP (BIR2)䡠caspase-3 complex fails to reveal any clues of how Survivin could directly suppress caspase-3 (17). Verdecia et al. (18) and Banks et al. (19) have demonstrated that Survivin is unable to directly bind to caspase-3 in vitro and does not inhibit caspase-3 activity. Wang and colleagues identified a novel mitochondrial-associated protein Smac/DIABLO that was able to interact with IAPs (20, 21) and may help explain the inhibitory function of Survivin. Smac/DIABLO is a mitochondria protein but is released into the cytosol in response to some of apoptotic stimuli, including UVB-irradiation, etoposide, or glucocorticoids (20 –23). After mitochondria import, the N terminus of precursor Smac/DIABLO is removed by limited proteolysis to generate a mature form of the molecule (20, 27). Mature Smac/DIABLO was found to promote caspase activation by binding and neutralizing the IAPs, including XIAP, CIAP-1, and CIAP-2 (20, 21, 23). Numbers of reports confirmed that mature Smac/DIABLO interacts with BIR2 and BIR3 of XIAP, and the N terminus of mature Smac/DIABLO is absolutely required for this interaction (24, 26 –29). However, whether Smac/DIABLO physically interacts with Survivin has yet to be fully characterized. In this report, we investigated the effect of interactions between Survivin and Smac/DIABLO on the apoptosis induced by Taxol (paclitaxel). We have demonstrated that Taxol is able to trigger apoptosis significantly in HeLa cell, resulting in the activation of cellular caspases-3, -7, and -9 and release of Smac/ DIABLO and cytochrome c from the mitochondrial. Ectopic expression of Survivin was able to significantly block the Taxolinduced cell death. Our GST pull-down binding assay and immunoprecipitation demonstrate for the first time that Survivin directly interacted with Smac/DIABLO but not with caspases. We also found mutant Survivin with a single amino acid change at residue Asp-71 (D71R) or truncated Survivin lacking the C terminus (residues 1–97) failed to bind to Smac/ DIABLO and abrogated its ability to inhibit apoptosis. On the contrary, these two Survivin mutants displayed some apoptotic effects. In addition, we have shown that wild type Survivin does not interact with Smac mutants M-Smac (a methionine 23130 This paper is available on line at http://www.jbc.org The Role of Survivin in Apoptosis was added to the begin of mature Smac) or ⌬74Smac (residues 75–239). We examined subcellular distributions of Survivin and Smac/DIABLO within living cells at different stages of the cell cycle and found that Survivin and Smac are co-localized in the cytosol during interphase, and when cells enter into M phase Survivin acts as a chromosome passenger protein. In addition, by using the cell-free system, we were able to demonstrate that both Smac/DIABLO and XIAP are required in order for inhibition of caspase cleavage by Survivin. Combined, our data strongly support our proposed hypothetical mode for Survivin in the inhibition of apoptosis. During apoptosis Survivin binds to Smac, which is released from induced mitochondria. Therefore, by reducing Smac/DIABLO antagonism to IAPs, such as XIAP, the free XIAP directly interacts with caspases and cell death is blocked. MATERIALS AND METHODS Oligonucleotides—The sequences of the oligonucleotides used in this study are listed as follows. P1, 5⬘-CGGGATCCGCATGGGTGCCCCGACGTTG-3⬘; P2, 5⬘-CGGTCGACCCATCCATGGCAGCCAGCTG-3⬘; P3, 5⬘-GCAAGCTTATGGCGGCTCTGAAGAGTTG-3⬘; P4, 5⬘-GCGGATCCTCCTCACGCAGGTAGGCCT-3⬘; P5, 5⬘-CGGTCGACGTTAATTCTTCAAACTGCTT-3⬘; P6, 5⬘-GCGATTAATATGGCGGTTCCTATTGCAC-3⬘; P7, 5⬘-CGGCGGCCGCATCCTCACGCAGGTAGGCCT-3⬘; P8, 5⬘-CGGGATCCGATTAATATGAGGAGAGCAGTGTCTTT-3⬘; P9, 5⬘-CGGGATCCTGATGGCGGTTCCTATTGCACAG-3⬘; P10, 5⬘-GGGAGCCAGATCGCGACCCCATAGAGGA-3⬘; P11, 5⬘-CTATGGGGTCGCGATCTGGCTCCCAGCC-3⬘; and P12, 5⬘-CGGGATCCTCAATCCATGGCAGCCAGCT-3⬘. Reagents and Antibodies—The following antibodies were used in this study: polyclonal antibodies Ab-caspase-7, Ab-caspase-3, Ab-survivin, and Ab-actin (Santa Cruz Biotechnology, Santa Cruz, CA); monoclonal antibodies: mAb-bcl2 (Oncogene, Manhasset, NY), mAb-GFP (MBL, Japan), mAb-cytochrome c (R&D Systems, Inc., Minneapolis, MN), mAb-caspase-9 (Immunotech, France), and mAb-Smac/DIABLO (Calbiochem, La Jolla, CA). Taxol used in our experiments is labeled as GCP grade. Restriction enzymes were mostly purchased from New England BioLabs. Medium compounds were obtained from Oxid. The majority of biochemical reagents were ordered from Sigma. Trypan blue was purchased from Invitrogen. Cell Culture and Transfection—HeLa cells were maintained in Dulbecco’s modified Eagle’s medium containing 10% heat-inactivated fetal bovine serum, 1 ⫻ nonessential amino acid, 1 ⫻ minimal essential medium sodium pyruvate, 100 g/ml penicillin, 100 g/ml streptomycin (Invitrogen, Grand Island, NY). Cultured cells were incubated in a humidified atmosphere containing 5% CO2 at 37 °C. Transfection of cells with various mammalian expression constructs by LipofectAMINE 2000 (Invitrogen) was carried out according to the methods provided by the manufacturer’s instructions. PCR-mediated Mutagenesis—To obtain the mutant survivin, we have employed the PCR-mediated mutagenesis method. Two pairs of primers, P1/P10 (complementing the regions from nucleotides 1 to 18 and nucleotides 196 to 224, respectively) and P11/P2 (complementing the regions from nucleotides 200 to 228 and nucleotides 408 to 426, respectively), were used to amplify two Survivin fragments. After gel purification, the resultant two overlapping PCR fragments were mixed with equal amounts. This mixture was incubated first at 94 °C for 4 min, followed by first PCR (PCR1) of 94 °C denaturing for 1 min, 56 °C annealing for 1 min, and 72 °C extension for 1 min. After 10 cycles, another 8-min extension was added to ensure the completion of all the extension at 72 °C. The PCR1 products were used as a template in a second PCR (PCR2), primed by oligonucleotides P10 and P11 to carry out another 20 cycles of PCR with the same PCR conditions used in PCR1. A prominent band with an expected size of 0.42 kb was visible on 1% agarose gel. A point mutation, GAC (Asp) to CGC (Arg), was further verified by DNA sequencing determination. Plasmids Construction—The full-length Survivin coding sequence (nucleotides 1– 426) and its mutant variant with a point mutation at amino acid residue 71 (D71R) were cloned into BamHI/SalI sites of pGEX-5X-3 (Amersham Biosciences, UK), respectively, to generate recombinant expression vectors pGEX-5X-3/Survivin and pGEX-5X-3/ Surv-D71R, and both fragments were inserted in-frame with the GST gene. The gene fragment (nucleotides 1–291), containing only the Survivin BIR domain (residues 1–97), was generated by the PCR method using primers P1 and P5 (see “Oligonucleotides”) and cloned into 23131 BamHI/SalI sites of pGEX-5X-3 to yield pGEX-5X-3/Surv-BIR. Gene fragments coding for wild type Survivin, for mutant Survivin (SurvD71R) with a point mutation at amino acid residue 71, for the truncated Survivin (Surv-BIR) lacking its C-terminal and Smac/DIABLO were also cloned into pEGFP-C1 plasmid, respectively. In addition, the gene fragments (reacted by PCR for primer pairs P12/P1) was used to generated pTRE2/A-Surv (coding for antisense Survivin). The DNA fragment coding for mature Smac/DIABLO (residues 56 –239) and DNA fragment coding for mutant Smac/DIABLO (⌬74Smac, residues75–239) were generated by PCR techniques using primer pairs P6/P7 and P8/P7, respectively. The resultant PCR products were digested with restriction enzymes VspI and NotI and cloned into NdeI/NotI sites of pET-22b (Novagen, Madison, WI) to generate pET-22b/Smac and pET-22b/ ⌬74Smac with an His6 tag fusion at its C terminus. A full-length Smac/DIABLO gene fragment (residues1–239) resulted from PCR amplification with primers P3 and P4 inserted into HindIII/BamHI sites of the mammalian expression vector pEGFP-N1 (Clontech, Palo Alto, CA) to fuse to the EGFP gene, and the resultant recombinant plasmid was designated as pEGFP-N1/F-Smac. DNA fragments, coding for mutant Smac/DIABLO with an methionine residue added at the N terminus of mature Smac/DIABLO (designated as M-Smac) and for mutant Smac/ DIABLO lacking 73 amino acid residues at its N terminus (designated as ⌬73Smac), were reacted by PCR away from Smac/DIABLO cDNA using primer pairs P9/P4 and P8/P4, respectively, and inserted into the BamHI site of pEGFP-N1 in fusion with EGFP to generate plasmids pEGFP-N1/M-Smac and pEGFP-N1/⌬73Smac. Expression and Purification of Fusion Proteins—Overnight cultured Escherichia coli cells DH5␣ or BL21/DE3 containing the fusion plasmids were grown at a 1:100 dilution in an Amp-LB medium until the optical density reached A600 ⫽ 0.5. Isopropyl-1-thio--D-galactopyranoside was then added to a final concentration of 0.4 mM. Cells were induced for another 3 h before they were harvested and subjected to sonication for 10 shot pulses of 20 s each. The maximum protein release was determined by the Bradford assay (Bio-Rad) by comparing the known concentration of bovine serum albumin (BSA). The GST fusion proteins were purified through the glutathione-SepharoseTM 4B beads (Amersham Biosciences). The solubilized His6-Smac or His6-⌬74Smac fusion proteins were purified by incubating with chelating SepharoseTM Fast Flow beads (Amersham Biosciences) according to the manufacturer’s instructions. The ultimate elution products were dialyzed with 1⫻ PBS and were used for in vitro interaction assay. In Vitro Interaction Assay—An in vitro interaction assay was performed as described by Suzuki et al. (50). Taxol-stimulated HeLa cells (2 ⫻ 106) were lysed with the 200-l lysis buffer as described above. Cell lysates or bacterially expressed and purified His6 tag proteins were incubated with GST or GST fusion protein immobilized on glutathioneSepharose 4B beads (Amersham Biosciences) for overnight at 4 °C and washed three times with 500 l of 1⫻ PBS. The bound proteins were eluted by elution buffer, and eluted products were subjected to SDSPAGE for Western blot analysis or Coomassie Blue staining. Cell Death Assay—The ability of Survivin, Smac/DIABLO, or their mutants to affect cell viability was assayed by transfecting or HeLa cells (2 ⫻ 104 cells/well) in 24-well plates with 0.3 g of mammalian expression vectors using LipofectAMINE 2000. Twenty hours after transfection, cells continued to be incubated with 100 nM Taxol for 36 h, and the viability of the cells was measured with the standard trypan blue exclusion method by counting blue dead cells. Data are expressed as percentages of control and are the means of three independent experiments. Western Blot Analysis—Cells were washed with 1⫻ PBS and resuspended with 5 volumes of cold lysis buffer (50 mM Tris-HCl (pH 7.5), 250 mM NaCl, 5 mM EDTA, 50 mM NaF, 0.5% Nonidet P-40) supplemented with protease inhibitor mixture (Roche Applied Science). The cell lysate was incubated on ice for 30 min and was then centrifuged for 10 min at 4 °C. The protein content of the supernatant was determined by using a BCA-200 protein assay kit (Pierce). Equal amounts of proteins (10 –20 g) were loaded onto the gel and separated by SDS-PAGE, and the resolved proteins were transferred to nitrocellulose membrane. After blocking with 5% nonfat milk in TBST (20 mM Tris-HCl, pH 8.0, 150 mM NaCl, 0.1% Tween 20) for overnight at 4 °C, the blot was incubated with primary antibody for 1 h at room temperature. The membrane was then washed with TBST and probed with horseradish peroxidase-conjugated secondary antibody for 1 h. The membrane was washed three times in TBST and developed by ECL using the manufacturer’s protocol. Digitonin fractionation of cells into membrane and cytosolic fractions used for detection of cytochrome c and Smac/DIABLO was performed as described by Ekert et al. (23). Immunoprecipitation—Cells were lysed in a Triton X-100-based lysis 23132 The Role of Survivin in Apoptosis buffer (1% Triton X-100, 10% glycerol, 150 mM NaCl, 20 mM Tris, pH 7.5, 2 mM EDTA, protease inhibitor mixture) for 1 h, and the nuclear and cellular debris was cleared by centrifugation. Then the cytosolic lysis was incubated with Smac monoclonal antibody bound to Protein A/G-Sepharose. After a 4-h incubation at 4 °C, the immunoprecipitates were washed five times in lysis buffer, and proteins were recovered by boiling the beads in SDS sample buffer and analyzed using a Western blot. Immunofluorescence Confocal Microscopy—HeLa cells were grown in six-well chamber slides, and 24 h later, cells were either left untreated or exposed to 100 nM Taxol for another 24-h incubation before 100 nM MitoTracker Red was added and treated for 60 min at 37 °C. The cells were then washed with PBS, fixed with 3.7% paraformaldehyde, permeabilized with 0.1% Triton X-100 in PBS, and blocked for 1 h in 3% BSA in PBS at 4 °C. Cells were incubated with anti-Smac (1:400 in 3% BSA) or anti-cyto c antibody (1:400 in 3% BSA) for 1 h at room temperature. The primary antibody was recognized by secondary FITC-conjugated antibody. Finally, the stained cells were analyzed with a laser scanning confocal microscopy system (Fluoview, Olympus). RESULTS Smac/DIABLO Is Released from the Mitochondria Accompanied with Cytochrome c upon Induction of Apoptosis by Taxol— Taxol is a natural product with potent anti-tumor activity and has been approved for the treatment of breast, ovarian, and lung cancers (25, 30 –32). We incubated HeLa cells with Taxol at a concentration of 100 nM, which is known to induce apoptosis in a number of different cell types. After 24-h incubation, the clear morphological changes characteristic of apoptosis were observed. Taxol-treated cells became rounded and detached from the substratum of the flask, followed by cellular shrinkage and membrane blebbing (data not shown). Caspase activation was believed to be the key factor during apoptosis. To evaluate which of the caspases is activated during the process of Taxol-induced apoptosis in HeLa cells, we analyzed the cleavage pattern of caspase-9, caspase-7, and caspase-3 by Western analysis methods. The results, shown in Fig. 1A, indicate that treatment with this drug results in the activation of caspase-9, caspase-7, and caspase-3 as evidenced by the appearance of caspase-active forms. One apoptotic signaling pathway leading into caspase activation involves the translocation of cytochrome c into the cytosol from the mitochondrial intermembrane space (33). To further investigate whether the mitochondria are involved in this event, Taxol-treated HeLa cells were first lysed and divided into membrane pellet and cytosolic fraction. The pellet was then treated with 0.5% bile acid deoxycholate and separated by centrifugation into a soluble and membrane fraction. Cytosolic proteins and membrane proteins solubilized by the deoxycholate treatment were subjected to Western blot analysis. As shown in Fig. 1B (panel a), an increasing amount of cytochrome c was detected in the cytosolic fractions, whereas a decreasing amount of cytochrome c remained in the mitochondrial membrane, and after 48 h, the majority of cytochrome c was detected in the cytosolic fractions. We also observed that Smac/DIABLO was predominantly present within the membrane fractions prior to treatment with Taxol, and by 48 h post-treatment, there was a significant loss of Smac/DIABLO from the membrane fractions and a concomitant increase of Smac/DIABLO in the cytosolic fractions (Fig. 1B, panel a). In addition, to further elucidate the behavior of Smac/DIABLO or cyto c during Taxol-induced apoptosis, we examined its subcellular localization in both untreated cells and treated cells by immunostaining and confocal microscopy (Fig. 1B, panel b). HeLa cells were stained with MitoTracker Red to define the location of mitochondria and with anti-Smac or anti-cyto c antibody plus an FITC-conjugated secondary antibody. In uninduced HeLa cells, the localization of FITC and MitoTracker Red staining gave rise to the yellow color indicating that both Smac/DIABLO and cyto c were localized within the mitochondria. However, after induction with Taxol for 24 h, FIG. 1. Mitochondrial cytochrome c and Smac are released during induction of apoptosis by Taxol. A, HeLa cells were treated with 100 nM Taxol for the indicated times before being harvested and lysed in lysis buffer. The processing of caspase-9 (a), -7 (b), and -3 (c) was then analyzed by Western blot probing with their respective antibodies. In B: a, HeLa cells were exposed to Taxol (100 nM) for the indicated times, and harvested cells were separated into membrane and cytosolic fractions as described previously (33). The two fractions were assessed for the contents of cytochrome c and Smac by immunoblot. Equal loading of proteins was measured by using a BCA-200 protein assay kit. b, the behaviors of cytochrome c and Smac were investigated by immunostaining in untreated HeLa cells and cells induced by Taxol for 24 h. Cells in i and ii were untreated; those in iii and iv were treated. The Role of Survivin in Apoptosis FIG. 2. Ectopic overexpression of Smac/DIABLO sensitizes HeLa cells to Taxol-induced apoptosis. A, schematic diagrams of Smac and its N-terminal deletion mutants. MTS, mitochondria targeting sequence, the first amino acid residue in each Smac or mutant Smac is numbered. Smac and the ⌬74Smac were expressed in bacteria as His6 fusion proteins, whereas F-Smac, ⌬73Smac, and M-Smac were expressed in HeLa cells as C-terminal EGFP fusion proteins. B, the expressed Smac and mutant Smac proteins were verified by Western analysis using anti-GFP antibody. Lane 1, EGFP; lane 2, F-SmacEGFP; lane 3, Smac-EGFP; lane 4, M-Smac-EGFP; lane 5, ⌬73SmacEGFP. Smac-EGFP was generated by removing the N terminus of F-Smac-EGFP during Taxol-induced apoptosis. The molecular sizes of the proteins are indicated at the left. C, HeLa cells were transfected with EGFP, F-Smac-EGFP, M-Smac-EGFP, or ⌬73Smac-EGFP expression vectors, and 20 h after transfection the cells were treated with 100 nM Taxol for another 36 h. The percentage of cell death was determined by the trypan blue exclusion method. both Smac/DIABLO and cyto c were partially released into cytoplasm from mitochondria evidenced by their changed localization pattern different from MitoTracker Red staining. Thus we have demonstrated that Taxol treatment results in the release of cyto c and Smac/DIABLO, suggesting that the mitochondria pathway is involved. Ectopic Overexpression of Smac/DIABLO Sensitizes HeLa Cells to Taxol-induced Apoptosis—Smac/DIABLO is capable of promoting apoptosis induced by various apoptotic stimuli, including TRAIL, UV irradiation, and etoposide (22, 23). We have demonstrated that treatment of Taxol could result in the release of Smac/DIABLO from mitochondria (Fig. 1B); it is therefore rational to propose that released cytosolic Smac/DIABLO may increase the susceptibility of HeLa cells to Taxol induction. To test this hypothesis, we transiently transfected HeLa cells with plasmids pEGFP-N1/F-Smac, pEGFP-N1/M-Smac, pEGFP-N1/⌬73Smac, and pEGFP-N1 separately (Fig. 2A). After 24 h of transfection, cell lysates were immunoblotted with 23133 anti-GFP antibody. Fig. 2B shows that full-length Smac (F-Smac) and its deleted mutants (M-Smac and ⌬73Smac) were overexpressed in the transfected cells but not in the control cells transfected with vector. Mature Smac resulted from F-Smac cleavage from mitochondria upon Taxol induction was also examined by Western blot (Fig. 2B, lane 3). We then determined whether overexpressed Smac/DIABLO promoted apoptosis induced by Taxol. 20 h after transfection, cells were treated with 100 nM Taxol and incubated for another 36 h then collected and subjected to trypan blue staining. As shown in Fig. 2C, elevated expression of full-length Smac, which was cleaved after mitochondria import into the mature Smac/DIABLO, significantly increased Taxol-induced apoptosis, whereas two Smac-deleted mutants, M-Smac and ⌬73Smac, which were reported to be unable to bind to XIAP (26 –29), were less proapoptotic than full-length Smac. These results suggest that, although the N terminus of mature Smac/DIABLO is likely to play a major role in the pro-apoptotic effect on cell death, the N-terminal sequence per se is not essential, because the Smac mutant lacking the first 18 amino acid residues (⌬73Smac) or the mutant with Met added in ahead of the first amino acid (A) of mature Smac (M-Smac) are able to partially compromise its pro-apoptotic activity. Combined, these data suggest that Smac/DIABLO has yet an uncharacterized mechanism underlying its pro-apoptotic activity and are in accord with previous reports that Smac/DIABLO has a dual role in the caspase cascade (27). Survivin Physically Interacts with Smac/DIABLO—It has been shown that Smac/DIABLO promotes apoptosis by neutralizing several IAP family members, particularly XIAP (20, 21). Survivin is a special IAP family member that contains a single IAP repeat (BIR) and lacks the RING finger motif. Survivin is found to be up-regulated in most transformed cell lines and in nearly all human tumors, but it is rarely present in normal adult tissues (3, 16, 34). Overexpression of Survivin was able to block cell death induced by various stimuli such as TRAIL and tumor necrosis factor (13, 15), yet the detailed mechanism remains controversial (16 –18). Particularly, whether Survivin interacts with Smac/DIABLO in vivo and how they interact to each other have not been documented. To determine whether Smac/DIABLO can bind to Survivin, we expressed GFP/Survivin, GFP/XIAP, or vector control in HeLa cells then treated transfected cells with 100 nM Taxol for 48 h to generate mature Smac, which is an active form (the first 55 amino acid residues of full-length Smac are cleaved), for interaction with IAPs. Immunoprecipitation of endogenous Smac demonstrated that Smac strongly interacts with GFP/XIAP or GFP/Survivin but not with control GFP protein (Fig. 3C, panel a). This suggests that Survivin is able to interact with mature Smac in vivo. However, the direct binding of Survivin to Smac/DIABLO has not yet been confirmed, because the co-immunoprecipitation experiment using cell lysate does not exclude the possibility that additional cellular factors may be involved in the binding. To study the direct interaction between Survivin and Smac/ DIABLO, we performed an in vitro binding assay that did not rely on proteins present in the eukaryotic cell lysate. We successfully expressed and purified GST/Survivin fusion protein and Smac/DIABLO with an His6 tag fused at its C terminus from bacteria. The soluble GST/Survivin and mature Smac/DIABLOHis6 were mixed and incubated at 4 °C for overnight, and this mixture was used for interaction assay. As shown in Fig. 3C (panel b), GST/Survivin protein was bound specifically to the mature Smac protein as evidenced by appearance of an eluted 23-kDa mature Smac band, whereas GST protein alone (control) was unable to bind to Smac. This result indisputably demonstrated that Survivin directly interacts with Smac/DIABLO. 23134 The Role of Survivin in Apoptosis FIG. 3. Survivin physically interacts with Smac. A, homology of the BIR domain between Survivin and other IAP family members was determined using NCBI BLAST software. The highly conserved residues in BIR of IAPs are shaded. B, schematic diagrams of Survivin and Survivin mutants Surv71 (D71R) and Surv-BIR. N, N-terminal. The coiled-coil structure is in lightly shaded color. The asterisk indicates point The Role of Survivin in Apoptosis To investigate which region of Survivin or Smac is responsible for interaction, we generated several mutants to perform interaction assay both in vitro and in vivo. Recently, numbers of reports have shown that XIAP directly interacts with Smac/ DIABLO, and this interaction involves the BIR domain of XIAP and the amino terminus of mature Smac (24, 26, 27). Both Survivin and XIAP are members of the IAP family characterized by the presence of one or more BIR domains. We thereby reasoned that the pattern of binding of Survivin to Smac/ DIABLO might be similar to that of XIAP to Smac/DIABLO. A BLAST program search for comparisons between the BIR domain of Survivin and BIR domains of other IAPs revealed that Survivin BIR domain displays high homology to BIR3 and BIR2 domains of XIAP (Fig. 3A). Based on the crystal structural analysis of complex of Smac/DIABLO䡠XIAP (BIR3), Fesik and coworkers (35) proposed that Glu-314 in the BIR3 domain of XIAP may directly contact with the N-terminal amine of mature Smac and that the E314S mutant of BIR3 abolishes almost all binding to mature Smac. In addition, it was proposed that D214S of the BIR2 mutant domain also lost all the capability of binding to Smac (35). As shown in Fig. 3A, amino acids Glu-314 in BIR3 domain or Asp-214 in BIR2 domain of XIAP are equivalent to Asp-71 of Survivin, because these amino acids are highly conserved. To demonstrate whether the amino acid Asp-71 of Survivin is critical for binding to Smac/DIABLO, we mutated Asp-71 to Arg-71 (Fig. 3B) and performed an interaction assay as described above. As shown in Fig. 3D (panel a), no eluted Smac band was detected (lane 6), indicating that the D71R mutant Survivin lost its binding ability to interact with mature Smac, and further suggested that Asp-71 of Survivin is vital for the binding to Smac. We further performed a coprecipitation experiment and confirmed that mature Smac does not bind to Surv-D71R in vivo (Fig. 3D, panel b). Both the BIR2 and BIR3 domains of XIAP were shown to directly interact with Smac/DIABLO, so we then examined whether the Survivin BIR domain alone (Fig. 3B) is able to bind to Smac/ DIABLO. To our surprise, there was no expected Smac band to be detected (lane 8) using an in vitro interaction assay (Fig. 3D, panel a), and an immunoprecipitation assay also failed to detect the GFP/Surv-BIR protein in the final eluted complex (Fig. 3D, panel b), implying that, unlike XIAP, Survivin BIR domain alone is unable to bind to Smac/DIABLO. The discrepancy may be explained by analogy: although XIAP is able to bind to HtrA2/Omi (a homolog of Smac/DIABLO), Survivin is unable to bind to it (36). In addition, the coiled-coil structure at the C terminus of Survivin is reported to be essential for inhibition of apoptosis (5), therefore, it is not unwise to argue that this coiled-coil structure, which is missing in the Survivin BIR domain, may be responsible for the interaction with Smac/ DIABLO. The N-terminal of Smac/DIABLO is essential for binding to XIAP (26 –29), we thereby checked whether this N terminus is also crucial for binding to Survivin. We made a series of Smac mutant constructs as depicted in Fig. 2A. Proteins ⌬74Smac and mature Smac were expressed and purified from bacteria E. coli cells, and proteins F-Smac, M-Smac, and ⌬73Smac were expressed in HeLa cells. Among them, proteins ⌬74Smac, M- 23135 Smac, and mature Smac were used in GST pull-down assay (see “Material and Methods”). Fig. 3E (panel a) shows that mutant variant ⌬74Smac is not detected in eluted products (lane 2), indicating that Survivin does not bind to ⌬74Smac, which lacks 18 amino acid residues at its N terminus of mature Smac. A further experiment showed that M-Smac-GFP in cell lysate was not able to bind to GST/Survivin-coupled beads (Fig. 3E, panel b). Taken together, these data suggest that the interactions between Survivin and Smac/DIABLO involve the N terminus of mature Smac, and, moreover, we propose that wild type Survivin may directly contact with Ala-1 of mature Smac, because M-Smac (with a methionine added to the beginning of mature Smac) nullifies the ability of mature Smac to interact with Survivin. Survivin Is Co-localized in Cytosol with Mature Smac during Interphase—Survivin not only acts as an inhibitor of apoptosis but also regulates cell division (5, 14, 15). Skoufias et al. (42) reported that human Survivin is a kinetochore-associated passenger protein and determined its localization by using an immunostaining method. However, different immunostaining protocols or antibodies produce different results because of the co-existence of Survivin splice variants (46). We thereby constructed a series of plasmids that express GFP, GFP/Survivin, GFP/Surv-BIR, GFP/Surv-D71R, and GFP/Smac (Fig. 4B) to study their distributions in living cells. We first transiently transfected HeLa cells with constructed plasmids; 48 h later, cells were incubated with DNA-staining dye Hoechst 33342 (2 g/ml) for 30 min, and the living cells were then examined by fluorescence microscope. As shown in Fig. 4A, GFP protein displayed an evenly diffused localization throughout the whole cells transfected with pEGFP-C1, and its distribution was found to be independent of the cell cycle. GFP/Survivin was firstly found to localize in the cytoplasm during interphase and then translocate into the central spindle midzone when cell cycle division entered into anaphase. Finally, GFP/Survivin was detected in the midbody during telophase (Fig. 4A). These data clearly demonstrate that Survivin is a bona fide chromosomal passenger protein. Our result is in good agreement with a previous report by Li et al. (5) in which they demonstrate that Survivin was able to bind to microtubules during interphase and to be translocated to the nucleus to regulate cell division at the G2/M phase. To investigate the subcellular distribution of Survivin mutants, we expressed GFP fusion proteins GFP/ Surv-D71R and GFP/Surv-BIR in HeLa cells and examined their distributions by fluorescence microscopy. Fig. 4A shows the intracellular localization of GFP/Surv-D71R in different phases of the cell cycle in HeLa cells. During interphase mutant GFP/Surv-D71R was distributed mainly in the cytoplasm similar to GFP/Survivin. As the cell cycle entered into anaphase, it spread evenly throughout the whole cells and was unable to localize at the spindle midzone. Interestingly enough, GFP/Surv-D71R displayed a midbody localization pattern identical to that of wild type GFP/Survivin during the late telophase. We also studied the localization of the GFP/Surv-BIR mutant and found that its localization was the same as that of GFP protein, which distributed uniformly throughout the cells and did not localize to either spindle midzone or the midbody. mutation of Asp-71 to Arg-71 (D71R). In C: a, lysates were prepared from GFP, GFP/XIAP, or GFP/Survivin stably transfected HeLa cells (treatment by Taxol for 48 h), respectively. Endogenous mature Smacs were immunoprecipitated (IP) from the lysates and examined for interaction with GFP, GFP/XIAP, and GFP/Survivin by Western blot (WB) with anti-GFP antibody. b, the interactions between GST/Survivin and mature Smac were examined by GST pull-down binding assay, and GST was used as mock control. In D: a, results from GST mediated pull-down assay. Survivin mutants did not interact with mature Smac. Lanes 1, 3, 5, and 7 indicate input Smac, and lanes 2, 4, 6, and 8 indicate the final eluted complex. b, the interaction of GFP-tagged Survivin or Survivin mutants with mature Smac was examined as in C, panel a. In E: a, the interaction of GST/Survivin with ⌬74Smac was investigated by in vitro interaction assay. Lane 1 denotes input ⌬74Smac, and lane 2 indicates the final eluted complex. b, lane 1 is a cytosolic fraction containing transfected expressed, which could not be pulled down by immobilized GST/Survivin (lane 2) and GST control (lane 3). Anti-GFP antibody was used to deleted M-Smac-EGFP. 23136 The Role of Survivin in Apoptosis FIG. 4. Survivin is co-localized with Smac/DIABLO during interphase. A, HeLa cells were transfected with plasmids, including pEGFP-C1, pEGFP-C1/Survivin, pEGFP-C1/Surv-D71R, pEGFP-C1/Surv-BIR, and pEGFP-C1/Smac. 48 h after transfection, cells were incubated with Hoechst 33342 (2 g/ml) for another 30 min, and the living cells were then analyzed with a fluorescence microscope. GFP or GFP fusion proteins are represented by the green color, and the blue color was generated by Hoechst 33342 staining. Interphase, anaphase, and telophase are denoted at the top. B, the expressions of five transfected recombinant plasmids indicated in A were verified by Western blot, and the primary antibody used was anti-GFP monoclonal antibody. These results suggest that the ␣-helix region at the C-terminal of Survivin contributes to its binding to microtubule in interphase and to its proper dynamic redistribution during mitosis. In addition, data from localization of mutant GFP/Surv-D71R (BIR domain was mutated) indicated that the BIR domain was absolutely required for the association of Survivin with spindle midzone. It has been reported that Survivin interacts physically with two kinetochore proteins INCENP and Aurora-B, which are known to be localized to both spindle midzone and midbody and thus to control cell cycle (43). Koufias et al. (42) reported that Survivin mutant Surv-C84A did not localize to either spindle midzone at anaphase or midbody in telophase. If complexing with INCENP and Aurora-B was the only mode for Survivin to localize to the chromosome during M phase, we would expect that the Survivin mutant Surv-D71R to have the same localization pattern with that of Survivin mutant SurvC84A or wild type Survivin. However, GFP/Surv-D71R protein, although unable to locate to the spindle midzone at anaphase, was able to localize to the midbody in telophase, suggesting that Survivin could utilize an as yet uncharacterized mechanism to become a chromosomal passenger protein during mi- tosis. In addition, we have demonstrated that Smac directly interacts with Survivin (Fig. 3C). Therefore, we expected that Smac/DIABLO should co-localize with Survivin in certain phase of cell cycle. We constructed a plasmid expressing mature GFP/Smac fusion protein in HeLa cells, and 24 h after transfection, GFP/Smac was found to locate exclusively in the cytosol and was not exhibited in the nucleus during the interphase (Fig. 4A). As the cell cycle enters into anaphase, unlike GFP/Survivin, GFP/Smac was completely excluded from the spindle midzone and did not associate with midbody in late telophase (Fig. 4A). Thus Survivin and Smac were co-localized in the cytoplasm during interphase, but their subsequent distributions are distinctive during mitosis. However, the possibility that Survivin interacts with Smac during M phase still cannot be completely excluded. Taking together, our findings suggest that Survivin is a chromosomal passenger protein during mitosis and it co-localizes with mature Smac in cytosol during interphase when Smac/DIABLO is released from mitochondria. Survivin Is Not a Direct Inhibitor of Caspase—Numbers of reports have demonstrated that Survivin can inhibit apoptosis The Role of Survivin in Apoptosis FIG. 5. Survivin is unable to bind to caspase. Taxol-treated (100 nM, 36 h) HeLa cells were collected and lysed with lysis buffer, and the cytosolic extracts were then used for GST-mediated pull-down assay as described under “Materials and Methods.” The cytosolic extracts and final eluted products were subject to Western blot analysis and probed with antibodies of caspases-9, -7, and -3 and Smac as needed. Lanes 1, 4, 7, and 10 were cytosolic fractions alone; lane 2, 5, 8, and 11 were final eluted products pulled from immobilized GST/Survivin. Lane 3, 6, 9, and 12 were eluted complex pulled from the immobilized GST control. In the left panel (lanes 1–3), Smac was shown to be successfully pulled down using the same conditions as those used in GST pull-down for caspase-9, -7, and -3. Molecular sizes of proteins are indicated at the left. induced by various stimuli (3, 5, 15). The mechanism whereby Survivin inhibits apoptosis still remains controversial. Survivin belongs to the IAP family; therefore, some groups reported that Survivin blocks cell death through binding to active caspase and in turn to inhibit caspases activity. Tamm et al. (16) reported that Survivin can bind to the effectors caspase-3 and -7 in vitro and inhibit cell death. O’Connor et al. (39) demonstrated that the inhibitory effect of Survivin appears through its binding to caspase-9. However, some other groups reported contradictory results: they claimed that, in the process of inhibition of cell death, Survivin does not directly interact with caspase-3 and -9 (18, 19, 37). To test whether Survivin physically interacts with caspases, we performed an in vitro interaction assay (see “Materials and Methods”). GST/Survivin fusion protein immobilized on glutathione-Sepharose 4B beads was incubated for 1 h with Taxol-treated HeLa cytosolic extracts, and the beads were then washed and eluted. The final eluted products were subject to Western analysis, and the results are shown in Fig. 5. Neither GST/Survivin nor GST control was found able to pull down caspase-3, -7, and -9 from Taxol-stimulated cell lysates, whereas both pro-caspase and processed caspase were detected in the cell lysate (input) lanes. As a positive control, mature Smac could be pulled down by the GST/Survivin as shown in Fig. 5. These results illustrate that Survivin interacts with Smac but not with caspase-3, -7, and -9, and therefore Survivin is not a direct suppressor of caspases. Recently, Takahashi and coworkers have pointed out that the linker region between BIR1 and BIR2 within XIAP is responsible for the active site-directed inhibition of caspase-3 and -7 (40), whereas a single BIR2 domain (residues 157–242) without a linker region does not bind to caspase-7 (29). Chai et al. (29) have further demonstrated that the BIR domains of XIAP are dispensable for the inhibition of caspase-3 and -7 and that a fusion protein between GST and the linker peptide of XIAP tightly binds to and potently inhibits caspase-7 and -3, whereas the GST-BIR2 (residues 156 –240) fusion protein lacking the linker region does not. Wu et al. (41) also reported that the linker of XIAP is the major determinant of binding and inhibitor for the caspases. Unlike XIAP, Survivin does not contain a linker region, and therefore it is not surprising for us to find that Survivin does not bind to caspase-3 and -7. In addition, results from a comparison of the x-ray crystallographic structures between Survivin and the XIAP (BIR2)䡠caspase-3 complex also support the hypothesis that Survivin does not suppress caspase-3 (17). These data are in good agreement with our conclusion that Survivin is unable to bind to caspases. Survivin Blocks Apoptosis May through Antagonizing the Activity of Smac/DIABLO—Both Survivin and XIAP were able 23137 to inhibit apoptosis, however, XIAP was found to be more potent than Survivin (16). In a co-transfection experiment conducted by Tamm et al. (16), although XIAP was shown to almost completely block cell death induced by Bax or Fas/ CD95, cell death could only be partially inhibited by Survivin under the same conditions. This more complete inhibition of apoptosis by XIAP was not due to higher levels of XIAP protein compared with that of Survivin (16) but to its higher activity, because in Bax or Fas/CD95-induced cells, the Survivin protein level was even higher than that of XIAP. All these observations, plus our finding that Survivin is unable to bind to caspases, suggest that Survivin blocks apoptosis, maybe through some other bridge of proteins to inhibit caspase activity indirectly. One of the possible bridge protein candidates could be the Smac/DIABLO based on our new finding that Survivin can directly interact with Smac/DIABLO (Fig. 3C). We reasoned that Survivin inhibits cell death, maybe through protecting other IAPs such as XIAP from being neutralized by Smac/DIABLO, thus allowing them to maintain their suppression on caspases. If this hypothesis is correct, mutant Survivin incapable of binding to Smac/DIABLO would be expected to lose its inhibitory activity. We have demonstrated previously that Survivin mutants Surv-D71R and Surv-BIR had lost their ability to bind to Smac/DIABLO (Fig. 3D); we therefore investigated whether the lack of interaction between Survivin mutant and Smac/DIABLO could result in the loss of the inhibitory effect of Survivin. We constructed a series of mammalian expression vectors to express Survivin, Surv-D71R, or SurvBIR in HeLa cells, and the transfected cells were then treated with 100 nM Taxol for 36 h, the cytotoxic effects were measured using the trypan blue exclusion method, and protein expression was detected by Western blot analysis (Fig. 6A, panel b). In this assay, although Survivin reduced Taxol-induced cell death by 22%, protein Surv-D71R or Surv-BIR was unable to protect cells from apoptosis (Fig. 6A, panel a). This suggests that abolishment of the inhibitory function for Surv-D71R and SurvBIR may be due to the loss of their interactions with Smac/ DIABLO. It is interesting to note that Survivin mutant SurvD71R causes more cell death compared with empty vector control (Fig. 6A, panel a), suggesting Surv-D71R is a novel dominant negative mutant. The conversion of anti-apoptotic wild type Survivin to pro-apoptotic mutant Survivin (SurvD71R) is not unusual, because Survivin mutations T34A or C84A or Survivin antisense were reported to be pro-apoptotic (39, 5), but the exact mechanism still awaits further investigation. Recently, Silke et al. (38) reported that XIAP mutants that were unable to bind to caspase and Smac/DIABLO completely lost the inhibitory function of XIAP, whereas XIAP mutants that were unable to bind to caspase but could bind to Smac/ DIABLO retained their inhibitory effects. These experimental data strongly support our notion that Survivin blocks apoptosis mainly through antagonizing the activity of Smac/DIABLO but not directly interacting with caspases (Fig. 5). As described above, we have illustrated that Survivin blocked Taxol-induced apoptosis through its binding to Smac/ DIABLO, because disruption of Smac䡠Survivin complex formation leads to a failure of apoptotic inhibition (Fig. 6A, panel a). To further verify this result, we established a cell-free system to examine whether Smac/DIABLO is a mediator involving in the inhibition of apoptosis by Survivin. We purified recombinant Smac fusion (with an His6 tag at its C terminus) and GST fusion proteins GST/Survivin, GST/Surv-D71R, and GST/Survivin-BIR for a cell-free assay system. The effects of Smac/ DIABLO or GST/Survivin were evaluated by studying the processing of caspase-9 and caspase-7 in cytosolic extracts from empty vector-transfected or XIAP-overexpressed HeLa cells 23138 The Role of Survivin in Apoptosis FIG. 6. Survivin antagonizes the pro-apoptotic activity of Smac/DIABLO. In A: Survivin mutants fail to inhibit apoptosis. a, HeLa cells were transfected with plasmid pEGFP-C1 (first bar), pEGFP-C1/XIAP (second bar), pEGFP-C1/Survivin (third bar), pEGFPC1/Surv-D71R (fourth bar), pEGFP-C1/Surv-BIR (fifth bar), or p EGFPC1/ant-Surv (sixth bar), respectively. 20 h later, cells were incubated with Taxol (100 nM) for 36 h, and the percentage of cell death was calculated as the means of three independent experiments. b, expression of proteins from transfected constructs in A (panel a) was confirmed by Western analysis. In B: the analysis of dATP/cyto c-dependent caspase-9 (a) and caspase-7 (b) processing in cytosolic extracts from mock or stable XIAP-expressed HeLa cells. The purified proteins GST/ Survivin and His tag/Smac and two Survivin mutants GST/Surv-D71R and GST/Surv-BIR were added to cytosolic extracts in different combinations indicated above the Western blot. The concentrations of various following the addition of cytochrome c and dATP. As shown in Fig. 6B (panel a), cell extracts without exogenously added cyto c and dATP left procaspase-9 uncleaved, and only one band of 45 kDa corresponding to procaspase-9 could be detected (Fig. 6B, panel a, lane 1). When cyto c and dATP were added, two cleaved bands (37 and 35 kDa) of caspase-9 were detected in the blot (Fig. 6B, panel a, lane 2). We found that overexpression of XIAP strongly prevents the appearance of p37 form (which is generated by caspase-3 cleavage) of caspase-9 but not of the p35 form (which is generated by auto-cleavage of caspase-9) (Fig. 6B, panel a, lane 3), indicating XIAP had blocked the activation of caspase-9. Addition of GST/Survivin or GST/SurvD71R or GST/Survivin-BIR alone into a cell-free mixture from mock cytosolic extracts was unable to block the appearance of the p37 form of caspase-9 (Fig. 6B, panel a, lanes 4 – 6), suggesting Survivin by itself is unable to inhibit caspase-9. To exclude the possibility that bacterially expressed Survivin fusion proteins may compromise their inhibitory functions, we used the cytosolic extracts from cells in which Survivin, SurvD71R, or Surv-BIR was overexpressed and obtained the same results, indicating a failure to block caspase-9 cleavage by Survivin alone is not due to Survivin expressed from Escherichia coli (data not shown). In addition, adding Smac protein into the cell-free mixture from XIAP-overexpressed cytosolic extracts abrogated the prevention of caspase activation by XIAP (Fig. 6B, panel a, lane 7) and resulted in the appearance of the p37 form, thus confirming that Smac is able to stimulate caspase-9 activation by removing the inhibition of XIAP. If the GST-Survivin was added to the mixture containing both XIAP and Smac, the p37 form will not be generated (Fig. 6B, panel a, lane 8). In contrast, when the mutant Survivin GST-SR71 or GST-BIR was added, the p37 form was detected (Fig. 6B, panel a, lanes 9 and 10). From these results we have reached two conclusions. First, Survivin is capable of blocking caspase-9 activation in the presence of Smac and XIAP. Second, Survivin mutants, which are unable to bind to Smac, fail to inhibit activation of caspase-9 even in the presence of XIAP and Smac. The same cell-free system was used to examine the effect of Survivin and Smac by evaluating the processing of caspase-7. We found that the presence of XIAP, but not of GST/Survivin, GST/Surv-D71R, or GST/Surv-BIR, was able to inhibit cleavage of caspase-7 (Fig. 6B, panel b, lanes 3– 6), whereas Smac protein was found to antagonize the inhibition of XIAP and generate a p19-active form of caspase-7 (Fig. 6B, panel b, lane 7). When XIAP, Smac, and GST/Survivin were mixed altogether in the cell-free system, cleavage of caspase-7 was blocked (Fig. 6B, panel b, lane 8), whereas the simultaneous presence of XIAP, Smac, and Survivin mutants GST/SurvD71R or GST/Surv-BIR were unable to block the cleavage of caspase-7, resulting in the appearance of p19 (Fig. 6B, panel b, lanes 9 and 10). XIAP is known to block the activities of caspase-9 and caspase-7 by its binding to caspases (29, 40). However, the inhibitory activity of XIAP can be eliminated by Smac through its direct interaction with XIAP, thus freeing the caspase from the XIAP䡠caspase complex (20, 21). In this cellfree system, we have demonstrated that Survivin alone was unable to block caspase activation due to its inability to bind to purified proteins and compounds used in the cell-free experiment were the following: cyto c (1 g/ml), dATP (1 mM), Smac (100 nM), and GST/Survivin, GST/Surv-D71R, and GST/Surv-BIR were all 200 nM. “⫹” represents presence and “⫺” is absence. C, HeLa cells were transfected or co-transfected with plasmids pEGFP-N1, pEGFP-C1/Survivin, pEGFP-C1/Survivin:pEGFP-N1/F-Smac(1:1), pEGFP-C1/Surv-D71R: pEGFP-N1/F-Smac (1:1), pEGFP-C1/Surv-BIR:pEGFP-N1/F-Smac (1: 1), or pEGFP-N1/F-Smac, 20 h after transfection the cells were treated with 100 nM Taxol for another 36 h. The percentage of cell death was determined by the trypan blue exclusion method. The Role of Survivin in Apoptosis caspase (Fig. 5). In addition, Survivin was able to rescue inhibition of XIAP through its binding to Smac, thus freeing XIAP from the XIAP䡠Smac complex. However, Survivin mutants Surv-D71R and Surv-BIR were not able to rescue the inhibitory effect of XIAP, because these two Survivin mutants were unable to bind to Smac (Fig. 3D). To further verify whether the results obtained from the cell-free system actually occurred in the cells in vivo, we co-expressed Survivin and Smac (1:1) in HeLa cells and found that co-overexpression of Survivin and Smac remarkably reduced the cell death induced by Taxol compared with overexpression of Smac alone, whereas co-overexpression of Survivin mutant (Surv-D71R or Surv-BIR) and Smac (1:1) were unable to decrease the cell death (Fig. 6C). This suggested that Survivin was able to antagonize the proapoptotic activity of Smac, in particular the BIR domain or C-terminal domain of Survivin were required for its antagonism. Taken together, data from the transfection experiment in vivo were consistent with that from cell-free system in vitro. Although our data have provided strong evidence that Survivin had inhibitory function with the help of XIAP and Smac, the possibility that Survivin involves yet uncharacterized pathways to prevent cell death cannot be dismissed. Additionally, the competitiveness of Survivin and XIAP for binding to Smac is required for further investigation. DISCUSSION In this study, we have shown that ectopic overexpression of Smac/DIABLO sensitizes HeLa cells to apoptosis induced by Taxol. We demonstrated for the first time that Smac/DIABLO physically interacts with Survivin both in vitro and in vivo. Mutational analysis revealed that amino acid Asp-71 in Survivin is critical for its binding to Smac/DIABLO, and a single amino acid change (D71R) converts Survivin from being antiapoptotic to pro-apoptotic. The deletion experiments showed that the C-terminal coiled-coil domain of Survivin is responsible for the interaction between Survivin and Smac/DIABLO. Similarly, we found that by removing 18 amino acids (AVPIAQKSEPHSLSSEALM) or adding one amino acid (Met) at its N terminus Smac/DIABLO abolishes its ability to interact with Survivin. In addition, Survivin is co-localized in cytosol with Smac/DIABLO during the interphase in living cells. More importantly, the Survivin䡠Smac complex formation contributes to Survivin inhibitory function during Taxol-induced apoptosis in HeLa cells. Smac/DIABLO, a mitochondria protein that is released to cytosol in response to a number of apoptotic stimuli, was found to bind IAPs and prevent them from inhibiting caspases (20 – 23). Previous reports (22–24, 44) have clearly shown that the BIR domains of IAPs, such as BIR2 or BIR3 of XIAP and BIR of ML-IAP, are sufficient to interact with Smac/DIABLO. However, in the present study, we found that the single BIR domain of Survivin failed to bind to Smac/DIABLO (Fig. 3D), indicating that the way Survivin binds to Smac/DIABLO is different from XIAP or ML-IAP. Smac/DIABLO and its newly identified homolog HtrA2/Omi were reported to be able to bind to XIAP in the same manner, but, unlike Smac/DIABLO, HtrA2/Omi was unable to interact with Survivin (36). Our result may shed some light on explaining why Survivin does not bind to HtrA2/Omi. Recently, Survivin has attracted growing attention due to its expression in various tumors and its potential application in tumor therapy (3, 45). However, its subcellular localization remains controversial: different immunostaining protocols or different antibodies used give rise to diverse results because of the presence of Survivin splice variants (46). To remedy these discrepancies, we analyzed the localizations of GFP/Survivin in living HeLa cells by using confocal laser microscopy. GFP/ Survivin was excluded from the interphase nucleus and was 23139 FIG. 7. Hypothetical model of Survivin in the inhibition of apoptosis. Smac is released from Taxol-induced mitochondria, then binds to overexpressed Survivin, which reduces neutralizing effect of Smac on XIAP. XIAP in turn was able to interact with caspases, and cell death was blocked. detected in the central spindle midzone at anaphase and localized to the remnants of a mitotic apparatus, midbody at telophase (Fig. 4A). Interestingly enough, Survivin mutant GFP/ Surv-D71R, which is unable to bind to Smac/DIABLO, lost its ability to localize at the spindle midzone at anaphase. This observation may explain the possible mechanism for the proapoptotic effect of Surv-D71R, because the other two dominant negative mutants, T34A and C84A, reported elsewhere displayed the same localization pattern as D71R (42, 47). Although it has been confirmed that Survivin is able to inhibit apoptosis induced by various stimuli, the mechanism by which Survivin inhibits apoptosis has not been conclusively determined. As with other IAPs, it has been reported that Survivin physically interacts with initiator or effector caspases (16, 37, 48). However, some groups (49, 50) demonstrated that this did not seem to translate into physiologically meaningful inhibition of caspase activity, and the x-ray crystal structure of Survivin also fails to reveal the presence of a “hook and sinker” region that mediates caspase binding in other IAPs (18, 37). Results from the present study have shown that Survivin inhibits apoptosis mainly through antagonizing the pro-apoptotic ability of Smac/DIABLO rather than through binding to caspases. We demonstrated that Survivin does not interact with caspases in vitro and that its mutants Surv-D71R and Surv-BIR, which are unable to bind to Smac/DIABLO, had lost their anti-apoptotic effect (Fig. 6A, panel a). By using a cell-free system, we have shown that XIAP but not Survivin, Surv-D71R or Surv-BIR, prevents cyto c induced caspases activation. This prevention can be blocked by adding Smac/DIABLO which neutralizes XIAP through the formation of Smac*XIAP complex (20, 21). When additional Survivin was subsequently added, prevention of caspases activation could be restored. However, Surv-D71R or Surv-BIR, when added into the cellfree system containing both XIAP and Smac, were unable to restore this prevention (Fig. 6B). These data suggest that Survivin is able to antagonize the pro-apoptotic effect of Smac/ DIABLO in vitro. We also demonstrated that Survivin blocks the pro-apoptotic activity of Smac/DIABLO in HeLa cells (Fig. 6C). Recently, it was reported that XIAP mutants, which had lost their ability to inhibit caspase-9 and caspase-3 yet maintained their ability to interact with Smac/DIABLO, were able to inhibit cell death and that this inhibition could be explained by showing that endogenous XIAP is sufficient to block apoptosis provided it is not antagonized by an IAP antagonist such as Smac/DIABLO (20, 21). In the present study, we have demonstrated that, although Survivin is unable to inhibit caspases, it possesses the ability to bind to Smac/DIABLO, thereby allowing endogenous IAPs such as XIAP to block caspases without being antagonized. Taken together, we provided a working model for the inhibition of apoptosis by Survivin. During apoptosis Survivin binds to Smac/DIABLO released from induced mitochondria, thus reducing antagonism of Smac/ 23140 The Role of Survivin in Apoptosis DIABLO to XIAP, the free XIAP from Smac䡠XIAP complex directly interacts with caspases, and cell death is blocked (Fig. 7). This mode suggests that the anti-apoptotic effect of Survivin may be mitochondria-dependent, because mature Smac is released only from mitochondria. This view is strongly consistent with ideas proposed by some other groups; they pointed to a more selective role of Survivin in antagonizing mitochondrialdependent apoptosis, because Survivin blocks mitochondrialinduced but not death-receptor-induced apoptosis in transgenic animals (53). Recent genetic data showed the heterozygous Survivin mice (surv ⫺/surv⫹) were more sensitive to mitochondrial-dependent cell death (52). Furthermore, cell death following loss/interference of Survivin showed the characteristics of mitochondrial-dependent apoptosis (39, 45, 51). In addition, it is worth noting that Survivin mutant Surv-D71R constructed by our group represents a novel dominant negative mutant, because overexpression of this mutant not only abolishes the inhibitory effect of Survivin but also promotes Taxol-induced apoptosis. This new finding may have clinical usefulness in designing novel therapeutic drugs. Acknowledgments—We thank Yi Wang and Shixin Liu for their excellent technical help. REFERENCES 1. Crook, N. E., Clem, R. J., and Miller, L. K. (1993) J. Virol. 67, 2168 –2174 2. Uren, A. G., Wong, L., Pakusch, M., Fowler, K. J., Burrows, F. J., Vaux, D. L., and Choo, K. H. (2000) Curr. Biol. 10, 1319 –1328 3. Ambrosini, G., Adida, C., and Altieri, D. C. (1997) Nat. Med. 3, 917–921 4. Adida, C., Crotty, P. L., McGrath, J., Berrebi, D., Diebold, J., and Altieri, D. C. (1998) Am. J. Pathol. 152, 43– 49 5. Li, F., Ambrosini, G., Chu, E. Y., Plescia, J., Tognin, S., Marchisio, P. C., and Altieri, D. C. (1998) Nature. 396, 580 –584 6. Zhao, J., Tenev, T., Martins, L. M., Downward, J., and Lemoine, N. R. (2000) J. Cell Sci. 113, 4363– 4371 7. Fraser, A. G., James, C., Evan, G. I., and Hengartner, M. O. (1999) Curr. Biol. 9, 292–301 8. Li, F., Ackermann, E. J., Bennett, C. F., Rothermel, A. L., Plescia, J., Tognin, S., Villa, A., Marchisio, P. C., and Altieri, D. C. (1999) Nat. Cell Biol. 1, 461– 466 9. Miller, L. K. (1999) Trends. Cell Biol. 9, 323–328 10. Reed, J. C., and Reed, S. I. (1999) Nat. Cell Biol. 1, E199 –E200 11. Uren, A. G., Beilharz, T., O’Connell, M. J., Bugg, S. J., van, Driel R, Vaux, D. L., and Lithgow, T. (1999) Proc. Natl. Acad. Sci. U. S. A. 96, 10170 –10175 12. Uren, A. G., Coulson, E. J., and Vaux, D. L. (1998) Trends. Biochem. Sci. 23, 159 –162 13. Reed, J. C., and Bischoff, J. R. (2000) Cell. 102, 545–548 14. Fortugno, P., Wall, N. R., Giodini, A., O’Connor, D. S., Plescia, J., Padgett, K. M., Tognin, S., Marchisio, P. C., and Altieri, D. C. (2002) J. Cell Sci. 115, 575–585 15. Altieri, D. C., Marchisio, P. C., and Marchisio, C. (1999) Lab. Invest. 79, 1327–1333 16. Tamm, I., Wang, Y., Sausville, E., Scudiero, D. A., Vigna, N., Oltersdorf, T., and Reed, J. C. (1998) Cancer Res. 58, 5315–5320 17. Riedl, S. J., Renatus, M., Schwarzenbacher, R., Zhou, Q., Sun, C., Fesik, S. W., Liddington, R. C., and Salvesen, G. S. (2001) Cell. 104, 791– 800 18. Verdecia, M. A., Huang, H., Dutil, E., Kaiser, D. A., Hunter, T., and Noel, J. P. (2000) Nat. Struct. Biol. 7, 602– 608 19. Banks, D. P., Plescia, J., Altier, D. C., Chen, J., Rosenberg, S. H., Zhang, H., and Ng, S. C. (2000) Blood 96, 4002– 4003 20. Du, C., Fang, M., Li, Y., Li, L., and Wang, X. (2000) Cell 102, 33– 42 21. Verhagen, A. M., Ekert, P. G., Pakusch, M., Silke, J., Connolly, L. M., Reid, G. E., Moritz, R. L., Simpson, R. J., and Vaux, D. L. (2000) Cell 102, 43–53 22. Chauhan, D., Hideshima, T., Rosen, S., Reed, J. C., Kharbanda, S., and Anderson, K. C. (2001) J. Biol. Chem. 276, 24453–24456 23. Ekert, P. G., Silke, J., Hawkins, C. J., Verhagen, A. M., and Vaux, D. L. (2001) J. Cell Biol. 152, 483– 490 24. Chai, J., Du, C., Wu, J. W., Kyin, S., Wang, X., and Shi, Y. (2000) Nature 406, 855– 862 25. Marone, M., D’Andrilli, G., Das, N., Ferlini, C., Chatterjee, S., and Scambia, G. (2001) Exp. Cell Res. 270, 1–12 26. Wu, G., Chai, J., Suber, T. L., Wu, J. W., Du, C., Wang, X., and Shi, Y. (2000) Nature 408, 1008 –1012 27. Srinivasula, S. M., Datta, P., Fan, X. J., Fernandes-Alnemri, T., Huang, Z., and Alnemri, E. S. (2000) J. Biol. Chem. 275, 36152–36157 28. Srinivasula, S. M., Hegde, R., Saleh, A., Datta, P., Shiozaki, E., Chai, J., Lee, R. A., Robbins, P. D., Fernandes-Alnemri, T., Shi, Y., and Alnemri, E. S. (2001) Nature 410, 112–116 29. Chai, J., Shiozaki, E., Srinivasula, S. M., Wu, Q., Datta, P., Alnemri, E. S., Shi, Y., and Dataa, P. (2001) Cell 104, 769 –780 30. Rowinsky, E. K., Cazenave, L. A., and Donehower, R. C. (1990) J. Natl. Cancer. Inst. 82, 1247–1259 31. Panvichian, R., Orth, K., Day, M. L., Day, K. C., Pilat, M. J., and Pienta, K. J. (1998) Cancer Res. 58, 4667– 4672 32. Nicolini, G., Rigolio, R., Miloso, M., Bertelli, A. A., and Tredici, G. (2001) Neurosci. Lett. 302, 41– 44 33. Liu, X., Kim, C. N., Yang, J., Jemmerson, R., and Wang, X. (1996) Cell 86, 147–157 34. Lu, C. D., Altieri, D. C., and Tanigawa, N. (1998) Cancer Res. 58, 1808 –1812 35. Liu, Z., Sun, C., Olejniczak, E. T., Meadows, R. P., Betz, S. F., Oost, T., Herrmann, J., Wu, J. C., and Fesik, S. W. (2000) Nature 408, 1004 –1008 36. Verhagen, A. M., Silke, J., Ekert, P. G., Pakusch, M., Kaufmann, H., Connolly, L. M., Day, C. L., Tikoo, A., Burke, R., Wrobel, C., Moritz, R. L., Simpson, R. J., and Vaux, D. L. (2002) J. Biol. Chem. 277, 445– 454 37. Kasof, G. M., and Gomes, B. C. (2001) J. Biol. Chem. 276, 3238 –3246 38. Silke, J., Hawkins, C. J., Ekert, P. G., Chew, J., Day, C. L., Pakusch, M., Verhagen, A. M., and Vaux, D. L. (2002) J. Cell Biol. 157, 115–124 39. O’Connor, D. S., Grossman, D., Plescia, J., Li, F., Zhang, H., Villa, A., Tognin, S., Marchisio, P. C., and Altieri, D. C. (2000) Proc. Natl. Acad. Sci. U. S. A. 97, 13103–13107 40. Suzuki, Y., Nakabayashi, Y., Nakata, K., Reed, J. C., and Takahashi, R. (2001) J. Biol. Chem. 276, 27058 –27063 41. Huang, Y., Park, Y. C., Rich, R. L., Segal, D., Myszka, D. G., and Wu, H. (2001) Cell 104, 781–790 42. Skoufias, D. A., Mollinari, C., Lacroix, F. B., and Margolis, R. L. (2000) J. Cell Biol. 151, 1575–1582 43. Wheatley, S. P., Carvalho, A., Vagnarelli, P., and Earnshaw, W. C. (2001) Curr. Biol. 11, 886 – 890 44. Vucic, D., Deshayes, K., Ackerly, H., Pisabarro, M. T., Kadkhodayan, S., Fairbrother, W. J., and Dixit, V. M. (2002) J. Biol. Chem. 277, 12275–12279 45. Mesri, M., Wall, N. R., Li, J., Kim, R. W., and Altieri, D. C. (2001) J. Clin. Invest. 108, 981–990 46. Mahotka, C., Wenzel, M., Springer, E., Gabbert, H. E., and Gerharz, C. D. (1999) Cancer Res. 59, 6097– 6102 47. Bolton, M. A., Lan, W., Powers, S. E., McCleland, M. L., Kuang, J., and Stukenberg, P. T. (2002) Mol. Biol. Cell. 13, 3064 –3077 48. Kobayashi, K., Hatano, M., Otaki, M., Ogasawara, T., and Tokuhisa, T. (1999) Proc. Natl. Acad. Sci. U. S. A. 96, 1457–1462 49. Conway, E. M., Pollefeyt, S., Cornelissen, J., DeBaere, I., Steiner-Mosonyi, M., Ong, K., Baens, M., Collen, D., and Schuh, A. C. (2000) Blood 95, 1435–1442 50. Shin, S., Sung, B. J., Cho, Y. S., Kim, H. J., Ha, N. C., Hwang, J. I., Chung, C. W., Jung, Y. K., and Oh, B. H. (2001) Biochemistry 40, 1117–1123 51. O’Connor, D. S., Wall, N. R., Porter, A. C., and Altieri, D. C. (2002) Cancer Cell 2, 43–54 52. Conway, E. M., Pollefeyt, S., Steiner-Mosonyi, M., Luo, W., Devriese, A., Lupu, F., Bono, F., Leducq, N., Dol, F., Schaeffer, P., Collen, D., and Herbert, J. M. (2002) Gastroenterology 123, 619 – 631 53. Grossman, D., Kim, P. J., Blanc-Brude, O. P., Brash, D. E., Tognin, S., Marchisio, P. C., and Altieri, D. C. (2001) J. Clin. Invest. 108, 991–999