* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Proton-motive force
Butyric acid wikipedia , lookup
Lactate dehydrogenase wikipedia , lookup
Magnesium in biology wikipedia , lookup
Fatty acid metabolism wikipedia , lookup
Photosynthesis wikipedia , lookup
Basal metabolic rate wikipedia , lookup
Nicotinamide adenine dinucleotide wikipedia , lookup
Phosphorylation wikipedia , lookup
Mitochondrion wikipedia , lookup
Photosynthetic reaction centre wikipedia , lookup
NADH:ubiquinone oxidoreductase (H+-translocating) wikipedia , lookup
Microbial metabolism wikipedia , lookup
Light-dependent reactions wikipedia , lookup
Electron transport chain wikipedia , lookup
Biochemistry wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Citric acid cycle wikipedia , lookup
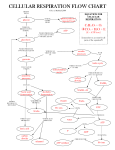
PROTON-MOTIVE FORCE Falling Water → electrical Energy Protons “Falling” → ATP Energy Balance • Overall energy balance • At cellular conditions, assume ∆G’ =54.3 kJ/mole for ATP hydrolysis to ADP + Pi • We know that the oxidation of glucose (e.g. in a calorimeter) yields ∆G’ = -2937 kJ/mole • Assuming that 32 moles of ATP are produced per mole of glucose, this gives an efficiency of: effic 32 moles ATP x 54.3 kJ / mole ATP 2937 kJ effic ca 60 % ( without O2 effic ~ 4%) • NADH + (1/2) O2 + H+ ⇄ H2O + NAD+ ΔGo’ = -220 kJ/mole • ADP + Pi + H+ ⇄ ATP + H2O ΔGo’ = -30.5 kJ/mole • Remainder “lost” as heat which we warm-blooded species use to keep warm How is NADH Oxidation Coupled to ADP Phosphorylation? First Attempts • Extend ideas from metabolism: transfer the high free energy electrons in NADH to a compound with high phosphoryl transfer potential • Researchers spent a couple of decades looking for an intermediate • Couldn’t find one Couldn’t find an Intermediate – so new hypothesis • In 1961 Peter Mitchell proposed a chemiosmotic mechanism in which entropy maximization tends to drive protons from the inter-membrane space to the matrix • Might add: he was ridiculed for his proposal, but eventually proved correct and was awarded the 1978 Nobel Prize (Chemistry) The Mitochondrian Proton Motive Force • Proton motive force is the sum of a chemical gradient and a charge gradient • pH is 1.4 units lower in inter-membrane as compared to matrix (ie inter-membrane is 25 x higher) • Charge gradient is 0.14 volts • Totaling these two gives PMF free energy of 20.8 kJ/mole e- ATP Synthase • Fo subunit “stick” proton channel • 10-14 c subunits “rotor” • Single a subunit on outside “stator” • F1 subunit “ball” • 3 units and 3 units in alternate ring to make up 33 hexamer “stator” • , stalk rotate with c ring “rotor shaft” • b2 and anchor the F1 unit to the membrane ATP Synthase • EC 3.6.3.14 ATP Synthase • The 8.5 nm “dots” are the F1 subunit of ATP Synthase • Thinnest human hair ~20m or 20,000 nm • ~2500 F1 subunits would span a human hair ATP Synthase Side view of F1 Top view of F1 subunit is pink ATP Synthase – F1 unit • ADP3- + HPO42- + H+ → ATP4- + H2O • Only sites are catalytic accept or release open= = loose, binds ADP and Pi = tight, makes ATP ATP Synthase – F1 unit • The rotation of 1 2 subunit drives the inter-conversion of the subunits • Subunit-sequence 1 - L→T →O →L →T… 2 - O→L →T →O →L… 3 - T→O →L →T →O… • unit moves counterclock wise 3 There is a Defined Order to Sequence • Note that there is a proper order to this sequence (O →L →T) • Open – molecules of ATP leave and ADP and Pi enter unit • Loose – ADP and Pi are bound to unit enzyme • Tight – the unit interacts with b enzyme to squeeze ADP and Pi together to make ATP • Open F1 Animation File_ATPsyn.htm Rotational Proof of Molecular Motor • Immobilize the 33 on glass slide • Link subunit to fluorescently labeled actin filament • Add ATP – hydrolysis of ATP caused filament to rotate Forming ATP Requires Free Energy • Remember that ATP hydrolysis to ADP and Pi is spontaneous: • ATP + H2O → ADP + Pi ΔGo’ = -30.5 kJ/mole + H2O → + Forming ATP Requires Free Energy • Reasons for spontaneity: • Phosphate groups are all negative, hence they electrostatically repelled • ADP and Pi have more resonance states than ATP, hence they are more stable. • For all of these reasons, free energy input is required to force Pi and ADP back together to make ATP • ADP3- + HPO42- + H+ → ATP4- + H2O • This energy is provided by conformationally changing enzymes driven by –subunit rotor which is in turn powered by H+ returning to the matrix How Does H+ Flow Cause to Rotate? • Take a closer look at the a and c subunits of Fo Opening at top Opening at bottom How does the c-ring turn? Animation matrix Intermembrane space How Does H+ Flow Cause to Rotate? H+ poor matrix side H+ rich cytoplasmic side How Does H+ Flow Cause to Rotate? • A closer look at the rotating c units + + + + + + The protein residues in the c-ring-a complex Aspartate, asp C-ring H H+ H+ a-unit Serine, ser Asparagine, asn Inlet H+ channel Arginine, arg Outlet H+ channel Rotational Summary Photomicrograph of C-ring Rates of Energy Production • A resting human being requires 85 kg of ATP each day • Molar mass ATP is 507.2 • Gives 167.6 moles of ATP/day • With ratio of 3 moles of electrons per mole of ATP generated • Gives 502.8 moles of electrons and with 6.02 x 1023 electrons per mole yields 3 x 1026 protons/day • Or 3.3 x 1021 protons/sec Getting ATP out into the Cytoplasm • ATP is made in the mitochondria matrix • It needs to get out into the cytoplasm in order to provide • • • • • power Remember that the inner membrane is itself only permeable to H2O, CO2 and O2 How do we get these highly charged ATP and ADP molecules in and out of matrix Need a transporter protein called ATP-ADP translocase ATP/ADP flows are coupled: ATP flows out only if ADP flows in and vice versa (antiporter) Cost approximately 1 H+ in proton motive force to make this happen Getting ATP out into the Cytoplasm Approximate ATP Yield NADH Only NADH FADH2 Total NADH = FADH2 Complex III & IV Per e- carrier Complex I 4 4 Complex II ------ 0 Complex III ------ 1 1 2 2 2 4 7 3 10 e- carriers 10 2 Protons pumped 70 6 ATP Synthase @ 3H+/ATP 23 2 Complex IV Total 0 Per 1 molecule Glucose 25 ATP Glycolysis & TCA 4 ATP Total 29 Yield of ATP from Glucose Oxidation Pathway ATP Yield per Glucose MalateGlycerol-3Phosphate Aspartate Shuttle Shuttle Glycolysis: glucose to pyruvate (cytosol) Phosphorylation of glucose Phosphorylation of fructose-6-phosphate Dephosphorylation of 2 molecules of 1,3-BPG Dephosphorylation of 2 molecules of PEP Oxidation of 2 molecules of glyceraldehyde-3phosphate yields 2 NADH (see below) Pyruvate conversion to acetyl-CoA (mitochondria) 2 NADH (see below for ATP yield) Citric acid cycle (mitochondria) 2 molecules of GTP from 2 molecules of succinyl-CoA Oxidation of 2 molecules each of isocitrate, -ketoglutarate, and malate yields 6 NADH (see below for ATP yield) Oxidation of 2 molecules of succinate yields 2 -1 -1 +2 +2 +2 +2 +3 +5 +5 +5 +3 +3 +15 30 +15 32 [FADH2] Oxidative phosphorylation (mitochondria) 2 NADH from glycolysis yield 1.5 ATP each if NADH is oxidized by glycerol-phosphate shuttle; 2.5 ATP by Oxidative decarboxylation of 2 pyruvate to 2 acetyl-CoA: 2 NADH produce 2.5 ATP each 2 [FADH2] from each citric acid cycle produce 1.5 ATP 6 NADH from citric acid cycle produce 2.5 ATP each Net Yield -1 -1 +2 +2 malate-aspartate shuttle each P/O Values • P/O ratio is the number of ATP molecules formed/pair of e• The table above assumes P/O = 2.5 for mitochondrial NADH • How do we get this? • Assume a 10 c-subunits in the ATP Synthase rotor • So 10 H+ are required for one complete rotation which makes 3 ATP or about 3H+/ATP • Add 1 H+ to power ATP-ADP translocase gives a total of 1 ATP/4 H+ • In ETC about 10 H+ pumped for every 2 e- of NADH/FADH2 → NAD+/FAD: 1 ATPcyto 10 H ATPcyto 10 2.5 4H pair e 4 pair e Control • Controlled by need of cell for ATP • Electrons flow through the electron transport chain only if protons are constantly being “used” to convert ADP to ATP