* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download 2007 - life.illinois.edu
Mitochondrial DNA wikipedia , lookup
Designer baby wikipedia , lookup
Nutriepigenomics wikipedia , lookup
DNA profiling wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Genetic engineering wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
DNA polymerase wikipedia , lookup
Cancer epigenetics wikipedia , lookup
SNP genotyping wikipedia , lookup
Gel electrophoresis of nucleic acids wikipedia , lookup
Genealogical DNA test wikipedia , lookup
United Kingdom National DNA Database wikipedia , lookup
Primary transcript wikipedia , lookup
Non-coding DNA wikipedia , lookup
Microevolution wikipedia , lookup
DNA vaccination wikipedia , lookup
DNA damage theory of aging wikipedia , lookup
Nucleic acid double helix wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Epigenomics wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Molecular cloning wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
DNA supercoil wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Point mutation wikipedia , lookup
History of genetic engineering wikipedia , lookup
Helitron (biology) wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Genomic library wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
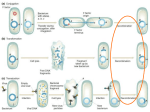
MCB 421 Exam #2 Fall 2007 There are 6 questions Be sure your name is on each page 1. 3. (10 points) You have a strain of Salmonella typhimurium that contains the plasmid pBR322. pBR322 carries a gene than encodes resistance to ampicillin (AmpR). The size of pBR322 is about 5 kb. You grow wild type P22 and P22 HT on the strain and prepare lysates. The titer of each lysate is 109 pfu/ml. When you infect an ampicillin-sensitive S. typhimurium with the lysate made with wild-type P22 you obtain no AmpR colonies. However, when you infect ampicillin-sensitive S. typhimurium with the P22 HT lysate, you obtain 500 AmpR colonies. a. (10 points) Why does the P22 HT lysate transduce AmpR while the wild type P22 lysate does not? Answer: Wild-type P22 requires a pac site to package DNA into phage heads. Apparently pBR322 does not contain a pac site or any good pac-like sites. P22 HT does not require a pac site to package DNA so it can package the pBR322 DNA and transduce it into a recipient. [P22 induces rolling circle replication of the plasmid so that concatomers are actually packaged into phage capsids]. 2. (6 points) A series of five mutants of phage T4 were isolated. Complementation tests were performed by spotting a mixture of the two phage mutants to be tested (about 106 of each) onto a lawn of bacteria at either 25°C or 40°C. The results are summarized below (+ = complete lysis; − = no lysis or only a few plaques in the spot). 1 2 3 4 5 1 + 25°C 2 + - 3 + - 4 + + + - 5 + + + - 1 2 3 4 5 1 - 40°C 2 3 - 4 + + + + 5 + + + + - (a) (3 points) How many complementation groups do these mutants affect and which mutants are in the same complementation groups? ANSWER: Two complementation groups: group I = mutants 1,2,3 and group II = mutants 4,5 (b) (3 points) Describe the nature of the mutation in each mutant (for example, nonconditional, temperature sensitive, cold sensitive)? ANSWER: Mutant 1 = TS (temperature sensitive) Mutant 2,3,5 = nonconditional (or null would be acceptable) Mutant 4 = CS (cold sensitive) 3. (15 points) Imagine you have isolated a mutant strain of Salmonella typhimurium that has a temperature sensitive mutation in the dnaA gene (dnaATS), You have isolated a Tn10 insertion (encodes resistance to tetracycline) that is 25 % linked to the dnaATS mutation by P22 HT mediated transduction. a). (5 points) How could you transfer the dnaATS mutation to a new S. typhimurium strain? How would you show that the new strain contained the dnaATS allele without DNA sequencing? Be sure to include the composition of the media and temperatures of each step. What frequency would you predict for transfer of the dnaATS allele in your experiment? Answer: Grow the dnaATS Tn10 strain at 30o, infect with P22 HT and make a lysate. Use the lysate to infect the second strain at 30o and plate out on (LB) plates supplemented with tetracycline. Incubate at 30o until colonies form. Streak or replica plate the colonies onto a new plate and incubate at 42o. The colonies that contain the dnaATS allele will not grow. You would expect that 25 % of the original TetR colonies would be TS. b). (10 points) Your lab mate has also isolated a temperature sensitive mutant. He claims that the mutation is also in the dnaA gene. How could you determine if his mutation was identical or different from your original dnaATS mutation? Answer: (If the mutation in your lab mate’s strain is at the same site as your mutation, you will not be able to obtain dnaA+ colonies when one strain is the donor and the other is the recipient in a transduction cross). Use your P22 HT lysate grown on your dnaATS Tn10 strain and infect the recipient. Select for TetR at 42o. If the mutations are at different sites, you should obtain some colonies that grow at 42o because they have a dnaA+ allele made by recombination (a crossover between the mutant sites). If the mutations are at the same site, you would expect no colonies at 42o. (Actually if you selected the tetR colonies at 30o and streaked or replica plated them and grew at 42o, the frequency of colonies that could grow at 42o would depend upon the distance between the mutant sites. If close together, they would be rare, but detectible if enough colonies were screened. If at the same site, no temperature resistant colonies would be found) 4. (30 points). The diagram below shows the immunity region of phage λ. The genes are: cI – codes for cI repressor. The cI857 allele is temperature sensitive. cro – codes for Cro protein. N – codes for the transcription anti-terminator. cII – codes for cI activator. cIII – codes for the cIII protease inhibitor. OL 1,2, and 3 and OR 1,2, and 3 are operators. PL and PR are the rightward and leftward promoters. PRE is the promoter for repressor establishment. PRM is the promoter for repressor maintenance. Calef et al. isolated a mutated lysogen that deleted most lambda DNA on the right and left sides of the immunity region as shown below. The mutant also has the cI857 TS allele (regions not shown below are deleted): The lysogen had always been maintained at 30° C previously to the experiment described below. They grew the lysogen at 30° C until cells reached mid-log growth. They split the culture in half and grew one at 30° C and the other at 42° C for several hours. They then lysed the cells from each culture as indicator bacteria for the ability of wt λ and λ imm434 to form plaques to 30° C. They obtained the results shown in the Table: Indicator Cells Grown at 30° C Grown at 42° C Cells Infected with wt no plaques clear plaques Infected with limm434 normal,turbid plaques normal , clear plaques The indicator cells grown at 42° C were returned to 30°C and grown for several generations and checked again for plaquing by wt λ and λ imm434. They obtained the same results as with the cells grown at 42° and not shifted back to 30° (i.e. they still obtained clear plaques with wt λ and turbid plaques with λ imm434). a. (5 points) Why doesn't wt λ form plaques on the indicator lysogen grown at 30° C? ANSWER: At 30°C, the cI857 repressor in the lysogen is active and will bind to the right and left operators of the infecting phage. This will prevent transcription of any phage proteins and thus prevent lysis. b. (5 points) Why can λ imm434 (as opposed to wt λ, above) form plaques on the indicator lysogen grown at 30°? Why are the plaques turbid? ANSWER: λ imm434 has the immunity region from phage 434, so the λ repressor is unable to bind its operators and repress transcription. So, the infecting phage may lyse the indicator. The plaques are turbid because the λ imm434 will eventually start making its own cI-like repressor (probably once the MOI becomes high) and form lysogens. c. (10 points) Why does wt λ form clear plaques on the indicator strain if it is grown at 42°C? ANSWER: λ is able to form plaques on the lysogen at 42° because the cI857 repressor is inactive at 42° and so does not prevent lysis by the infecting phage. The lack of cI in the recipient lysogen leads to accumulation of cro protein. Cro protein prevents transcription of cI, both in the lysogen and in the incoming phage. This ensures that incoming phage will only be able to grow lytically, and so the plaques are clear. d. (10 points) Why doesn't growth of the indicator lysogen at 30°C for several hours restore the original phenotype (no plaques with wt λ) of indicator cells once they have been grown at 42°? Answer: Once cI is deactivated at 42°, transcription of cro protein starts. Once cro protein is present, it prevents transcription of cI, even when the temperature is shifted back down to 30°C. Thus, the lysogen loses immunity even when shifted back to 30°. 5. (25 points) Arber and Dussoix (1962, J. Mol. Biol. 5:37 – 49) performed the following series of experiments to study restriction and modification in E. coli. In one experiment they grew phage lambda on an E. coli K strain and an E. coli K strain lysogenic for phage P1- E. coli K (P1). They used the lysate obtained to perform plaque forming assays on either E. coli K or E. coli K (P1) as the indicator strains. They obtained the data in the table below. Relative efficiency of PFU / ml Lysate E. coli K E. coli K (P1) Lambda on K 1 0.001 Lambda on K (P) 1 1 a, (5 points) What is the likely explanation for this result? Why? Answer: The P1 prophage has a restriction-modification system. The system is different from the K system. When lambda is grown on the K host, the DNA is K modified but not P1 modified. Thus when this phage infects the K strain, they make plaques efficiently because the DNA is not restricted. However, the P1 restriction system degrades the DNA and the frequency of plaques decreases by 1,000-fold. The lambda phage grown on the E. coli K (P1) host is modified for both K and P1 sites so they are resistant to both K and P1 restriction systems. OR the few plaques found when the K-grown phage infects the P1 lysogen are rare chromosomes that became P1 modified before the P1 restriction enzyme degrades the DNA. b. (10 points) In a second experiment, they labeled lambda DNA of phage growing in E. coli K (P1) with 32P so that the newly synthesized DNA in the phage was radioactive. They then infected E. coli K cells with the radioactive phage in medium containing nonradiactive phosphorus and found that the progeny phage that form plaques on E. coli K are resistant to inactivation by radioactive decay. However, the progeny that were able to form plaques on E. coli K (P1) were destroyed as time passed. What is the explanation for this result? Answer: The vast majority of DNA synthesized during growth in the E. coli K strain is not radioactive because the phages were made in non-radioactive medium. However, a (rare) phage that is able to form a plaque on E. coli K (P1) must contain one strand that has the original P1 modification so it can prevent the P1 restriction enzyme from destroying its DNA. This strand will also contain 32P from growth in the presence of the radioisotope and decay of the isotope over time emits radiation that destroys the infectivity of the phage DNA (i. e. radioactive decay destroys the chromosome. They also performed another experiment using density labeling of phage DNA. They grew lambda on E. coli K (P1) in medium containing deuterated water (D2O). D is a heavy isotope of hydrogen. Incorporation of D instead of H into DNA makes it denser than DNA made in the presence of H2O. They used the resulting lysate to infect E. coli K in medium containing normal H2O and used density gradient centrifugation to separate the resulting light phage form the more dense phage (Don’t worry about how this was done). They found that the phages that formed plaques on the E. coli K (P1) strain were heavy. Light phages (containing only hydrogen) could only form plaques on E. coli K. c. (10 points) What is the explanation for this result? Answer: The observation that the higher density phages (meaning that the DNA they contain is labeled with D) form plaques on E. coli K (P1) suggests that one strand of their DNA is modified. This indicates that these phage contain a strand of DNA from the original “heavy” phage and a complementary “light” strand made from DNA replication containing hydrogen in the E. coli K host. Thus, these phages can form plaques on E. coli K (P1) because the hemimethylated P1 site in the original parental strand prevents cleavage by the P1 restriction enzyme. The vast majority of the phage growing in E. coli K in the presence of H20 are light because all of their DNA is light. Since they are growing in E. coli K, their DNA is not P1 modified. Thus the light phage can only make plaques on E. coli K. 6. (14 points) Compare and Contract Generalized and Specialized Transduction for the indicated properties. Generalized Transduction Site-Specific Transduction Mechanism of Formation of Transducing Particles Packaging Chromosomal DNA Aberrant Excision Method of DNA Packaging Headful/pac Headful/cos Markers for Transducing Any (or many) Bacterial + phage DNA in capsids of Transducing Particle Bacterial Only Bacterial + phage Mechanism of Transduction HR HR or site specific Enzymes/protein required RecA mediated Int/RecA Susceptibility to host modification Susceptible Susceptible