* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download TOLL-LIKE RECEPTORS Toll-like receptors & Host
Survey
Document related concepts
Cell culture wikipedia , lookup
Cell membrane wikipedia , lookup
Cytokinesis wikipedia , lookup
Cell encapsulation wikipedia , lookup
Organ-on-a-chip wikipedia , lookup
Hedgehog signaling pathway wikipedia , lookup
Cellular differentiation wikipedia , lookup
Endomembrane system wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
List of types of proteins wikipedia , lookup
Transcript
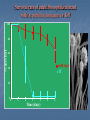
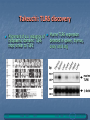
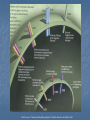
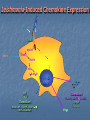
TOLL-LIKE RECEPTORS Toll-like receptors & Host-Pathogen Interaction O’Neill, Luke A.J. “Immunity’s Early-Warning System”. Scientific American, Jan (2005), 38-45. Microbe products recognized Conserved amoung microbes Known as pathogen-associated molecular patterns (PAMPs) PAMPs are recognized by plants as well as animals, meaning this innate response arose before the split Only vertebrates have evolved an adaptive immune response Pattern Recognition Receptors (PRRs) • Toll-like receptors • Natural history, function and regulation • • • • Mannose binding lectin (MBL) C-reactive protein Serum amyloid –P Functions of PRRs: – Opsonization, activation of complement and coagulation cascades, phagocytosis, activation of pro-inflammatory signaling pathways, apoptosis Nuesslein-Volhard: Drosophila Toll Identified a protein she called “Toll” meaning “weird” Helps the Drosophila embryo to differentiate its top from its bottom (Neural tube development) 1985 1988 1989 1991 http://www.nature.com/genomics/papers/drosophila.html 1996 1997 1998 1999 2000 2001 Gay: Toll and Inner Part of Human IL-1R is Similar Searching for proteins similar to Toll Shows cytoplasmic domain of Toll related to that of hIL-1R Identity extends for 135 aa Didn’t make sense Why does a protein involved in human inflammation look like one involved in fly neural tube development? 1985 1988 1989 1991 1996 1997 1998 1999 2000 2001 Toll Molecular Structure IL-1R Toll (will become TLRs) Ig-like domain LRRs Box 1 Box 2 Box 3 TIR Domain Toll receptor has an extracellular region which contains leucine rich repeats motifs (LRRs) Toll receptor has a cytoplasmic tail which contains a Toll interleukin-1 (IL-1) receptor (TIR) domain Lemaitre: Flies use Toll to Defend from Fungi Infected Tl-deficient adult flies with Aspergillus fumigatus All flies died after 2-3 days Flies use Toll to defend from fungi Thus, in Drosophila, Toll seems to be involved in embryonic development and adult immunity 1985 1988 1989 1991 1996 1997 1998 1999 2000 2001 Lemaitre: Flies use Toll to Defend from Fungi Drosophila has no adaptive immune system Therefore needs a rapid antimicrobial peptide response Two distinct pathways to activate antimicrobial peptide genes in adults Mutations in Toll pathway reduce survival after fungal infection 1985 1988 1989 1991 1996 1997 1998 1999 2000 2001 Survival rate of adult Drosophila infected with Aspergillus fumigatus in Toll100 % survival 80 60 wild type Tl- 40 20 0 0 1 2 3 4 Time (days) 5 6 Medzhitov & Janeway: Human Toll Discovery Ancient immune defence system based on the Toll signalling In insect, IL-1 receptor and the Toll protein are only similar in the segments within the cell They searched for human proteins that totally resemble to Toll 1985 1988 1989 1991 1996 1997 1998 1999 2000 2001 Medzhitov & Janeway: Human Toll Discovery Alignment of the sequences of human and Drosophila Toll proteins Homology over the entire length of the protein chains hToll gene most strongly expressed in Spleen and PBL (peripheral blood leukocytes) 1985 1988 1989 1991 1996 1997 1998 1999 2000 2001 Rock: Identification of hTLR1-5 Identified 5 human Tolls, which they called Toll like receptors (TLRs) TLR4 same as Medzhitov’s human Toll 4 complete - 1 partial hTLR 3 Drosophila TLRs 1985 1988 1989 1991 1996 1997 1998 1999 2000 2001 Poltorak: TLR4 Activated by LPS Normal mice die of sepsis after being injected with LPS C3H/HeJ mice have defective response to LPS and survive Missense mutation affecting the cytoplasmic domain of Tlr4 Major breakthrough in the field of sepsis – molecular mechanism that underlies inflammation revealed 1985 1988 1989 1991 1996 1997 1998 1999 2000 2001 Takeuchi: TLR6 discovery Alignment of a.a. sequence of cytoplasmic domains: TLR6 most similar to TLR1 1985 1988 1989 1991 Murine TLR6 expression detected in spleen, thymus, ovary and lung 1996 1997 1998 1999 2000 2001 Chuang (2000): hTLR 7, 8 and 9 Reported the cloning and characterization of 3 hTLRs Ectodomain with multiple LRRs Cytoplasmic domain homologous to that of hIL-1R Longer ectodomain (higher MW) than hTLR1-6 mRNA expression: hTLR7 - lung, placenta and spleen hTLR8 – lung and PBL hTLR9 - spleen, lymph node, bone marrow and PBL 1985 1988 1989 1991 1996 1997 1998 1999 2000 2001 Chuang (2001): Isolation of cDNA encoding hTLR10 Contains 811 aa, MW 94.6 kDA Architecture of hTLR10 same as in hTLR1-9 1985 1988 1989 1991 hTLR10 Expression of hTLR10 in human tissues and cell lines 1996 1997 1998 1999 2000 2001 Chuang: hTLR10 Phylogenetic tree of hTLR: a.a. identity with hTLR1 (50%) and hTLR6 (49%) Only 30% with hTLR2 and 25% with the remaining ones 1985 1988 1989 1991 1996 1997 1998 1999 2000 2001 TLR Roles O’Neill, Luke A.J. “Immunity’s Early-Warning System”. Scientific American, Jan (2005), 38-45. TLR Cell Type Distribution Receptor Cell Type TLR1 Ubiquitous TLR2 DCs, PMLs, and monocytes TLR3* DC and NK cells, upregulated on epithelial and endothelial cells Macrophages, PMLs, DCs, ECs, but not on lymphocytes Monocytes, immature DCs, epithelial, NK, and T cells TLR4 TLR5 TLR6 High expression in B cells, lower on monocytes and NK cells TLR7 B cells, plasmacytoid percursor DCs TLR8 Monocytes, low in NK cells and T cells TLR9 Plasmacytoid percursor DCs, B cells, macrophages, PMLs, NK cells, and microglial cells TLR10 B cells, plasmacytoid precursor DCs TLR11 Not Determined Toll-Like Receptors and their Ligands Receptor TLR1 Ligand (PAMPs) Origin of Ligand Triacyl lipopetides Soluble factors Bacteria and Mycobacteria TLR2 Heat Shock protein 70 Peptidoglycan Lipoprotein/lipopeptides HCV core and nonstructural 3 protein Host Gram-positive bacteria Various pathogens Hepatitis C Virus TLR3 Double-stranded RNA Viruses TLR4 Lipopolysaccharides Envelope protein Taxol Gram-negative bacteria Mouse mammary-tumor virus Plants TLR5 Flagellin Bacteria TLR6 Zymosan Lipoteichoic acid Diacyl lipopetides Fungi Gram-positive bacteria Mycoplasma TLR7 Single-stranded RNA (ssRNA) Imidazoquinoline Viruses Synthetic compounds TLR8 Single-stranded RNA (ssRNA) Imidazoquinoline Viruses Synthetic compounds TLR9 CpG-containing DNA Bacteria, Malaria and Viruses TLR10 Not determined Not Determined TLR11 Profilin-like molecule Toxoplasma gondii Neisseria meningitidis Converging Pathways Beutler, Nature 2004 Effects of signaling are cell specific NF-B activation is the end result of TLR-signaling TLR Signaling Pathways TLR2/TLR1 TLR2/TLR6 TLR4 TLR3 Cell membrane MAL MyD88 MAL MyD88 TRIF TRAM TRIF H+ H+ H+ TLR3 H+ TLR7 TLR8 TLR9 H+ H+ H+ H+ NF-B H+ H+ IRF3 Interferon Pathway Inflammatory Cytokines H+ H+ Endosome IRF7 TRIF MyD88 NF-B MyD88 Dependent and Independent Pathways: Major Role in Phagocyte Response LPS LBP MD-2 TLR4 MD-2 LPS sCD14 Cell membrane p50 p65 NF-kB NF-B p50 TLR4 MyD88-Independent Signaling MAL MyD88 TLR4 MyD88-Dependent Signaling p50 p65 p50 IRF-3 PP NF-kB TNF COX2 IL-18 Chemokines P P NF-B IFN- Chemokines: Rantes, IP-10 IFN LPS TLR4 MyD88-Dependent Signaling MD-2 LBP sCD14 Cell membrane IRAK1 IRAK2 MKK3 MKK7 UBC13 TRAF6 TAB2 TAK1 p38 IKK- TAB1 UBV1A IKK- IRAK4 MEKK3 IKK- MAL MyD88 TOLLIP (-) TLR4 MD-2 LPS Proteasome JNK IB IB p50 NF-B p65 TNF COX2 IL-18 Paz S., Nakhaei P,( 2005) LPS LPS TLR4 MyD88-Independent Signaling MD-2 MD-2 LBP sCD14 Cell membrane IKK- IB IB p50 IKK- IKK- TRAM TRIF TRAF6 Proteasome TLR4 TBK1 IKK P IRF-3 P p65 P P P P Late induction NF-B IFN- Paz S., Nakhaei P,( 2005) LPS dsRNA CpG DNA TLR4 ssRNA Cell membrane TRIF Tyk2 Jak1 TRAM Endosome STAT2 STAT1 ssRNA CpG DNA TLR7/8 TLR9 TBK1 STAT2 IRF-9 STAT1 MyD88 IKK- IKK- IKK- IRF-3 IRAK4 IRAK1 Proteasome TRAF6 IRF-7 IB IB p50 IFN- IFN- p65 Inflammatory Cytokines IFN Regulation NF-B Paz S., Nakhaei P,( 2005) ST2 SIGIRR (-) MD-2 MD-2 TLR4 (-) IRAK4 IRAK1 TRAF6 (-) UBC13 A20 UBV1A Proteasome IKK- IKK- IKK- (-) SOCS1 IRAK-M MyD88 MAL Cell membrane IB IB p50 p65 NF-B TNF COX2 IL-18 Negative Regulation of TLR Signaling in Phagocytes Cytoplasmic molecules: IRAK-M (restricted to monocytes and macrophages) SOCS1 (Supressor of cytokine signaling 1) A20 (TNFAIP3) Membrane bound molecules: SIGIRR (single immunoglobulin IL-1R-related molecule) ST2 Phagocyte Sabotage: Evading TLR Signaling Yersinia LcrV TLR2/TLR1 TLR2/TLR6 TLR4 Pseudomonas LPS Cell membrane TRIF TRAM MAL MyD88 NF-B Cytosolic Listeria Changing the target: Camouflaging or directly modifying the molecules that trigger TLR signaling (ex: P. aeruginosa). Crossing the wires: Interfering with downstream TLR-mediated signaling or to express TLR agonists (ex: Y. pestis). Sneaking through the back door: Bacteria such as Shigella sp. and Listeria sp. express proteins that facilitate their invasion of macrophages. Inflammatory Cytokines Nature Reviews Molecular Cell Biology 4; 385-396 (2003); Leishmania-Induced Chemokine Expression LPS TLR4 MyD88 independent MyD88 SHP-1 IRF-3 (-) IRAK-1 TRAF6 ? IKKs IB-NFB NO CD14 Chemokines (Rantes, IP-10, MCP-1, MIP-1/, MIP-2, Eotaxin) NF-B AP-1 Chemokines (MCP-1, MIP-1/, MIP-2) No NO No CD14 IFN- Chemokines, linking innate and adaptive immunity