* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Genetic Epidemiological Strategies to the Search for Osteoporosis
Human genome wikipedia , lookup
Genetic testing wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Medical genetics wikipedia , lookup
Pharmacogenomics wikipedia , lookup
Essential gene wikipedia , lookup
Genetic engineering wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Pathogenomics wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Human genetic variation wikipedia , lookup
Gene expression programming wikipedia , lookup
Population genetics wikipedia , lookup
Genomic imprinting wikipedia , lookup
History of genetic engineering wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Ridge (biology) wikipedia , lookup
Designer baby wikipedia , lookup
Genome evolution wikipedia , lookup
Public health genomics wikipedia , lookup
Minimal genome wikipedia , lookup
Gene expression profiling wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Behavioural genetics wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Microevolution wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
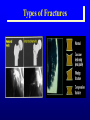
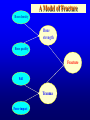
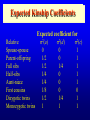
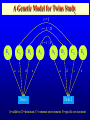
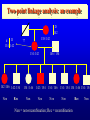
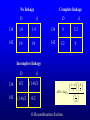
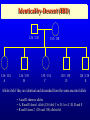
Genetic Epidemiological Strategies to the Search for Osteoporosis Genes Dr. Tuan V. Nguyen Bone and Mineral Research Program Garvan Institute of Medical Research Sydney, Australia Contents of Presentation • • • • • Epidemiologic results Clinical features of osteoporosis Determinants of fracture risk Genetics of bone mineral density The search for osteoporosis genes Definition Osteoporosis is a metabolic bone disease, which is characterised by low bone mass, microarchitectural deterioration of bone tissue leading to enhanced bone fragility and a consequent increase in fracture risk (Consensus Development Conference, 1991) Magnitude of the Problem Prevalence of Osteoporosis Incidence of all Fractures % Per 100,000 80 12000 70 Males 10000 60 Females 8000 50 6000 40 30 4000 20 2000 10 0 0 60-64 65-69 70-74 75-79 80+ 60-64 65-69 70-74 75-79 80+ Incidence of Hip Fx Worldwide Type of Fractures Types of Fractures A Model of Fracture Bone density Bone strength Bone quality Fracture Fall Trauma Force impact Risks Factors for Hip Fractures Females Unit Relative Risk Femoral neck BMD 0.12 g/cm2 2.1 (1.6 - 2.6) Falls Each fall 2.4 (1.5 - 3.8) Postural sway 2000 mm2 1.3 (1.1 - 1.5) Femoral neck BMD 0.12 g/cm2 2.4 (1.6 - 3.7) Falls Each fall 3.9 (1.7 - 9.3) Age 5 years 1.7 (1.2 - 2.3) Males Relationship between Fracture and Bone Mineral Density Change in BMC and BMD with Age 5 0 1 4 Hip BMC 4 0 3 0 Hip BMD 1 0 Percentperyear Percentperyear 2 0 1 0 6 2 0 -1 0 5 1 0 1 5 2 0 -2 4 2 5 8 1 2 B a se lin ea g e 1 6 2 0 2 4 2 8 B a se lin ea g e 5 0 Spine BMC 4 0 Spine BMD 1 4 3 0 1 0 Percentperyear Percentperyear 2 0 1 0 6 2 0 -1 0 5 1 0 1 5 B a se lin ea g e 2 0 2 5 -2 5 1 0 1 5 B a se lin ea g e 2 0 2 5 Determinants of Peak Bone Mass Genetic factors Nutritional factors Peak Bone Mass 16-25 yr of age Exercise and environmental factors Hormonal factors Risk Factors for Osteoporosis Genetics Race, Sex, Familial prevalence Hormones Menopause, Oophorectomy, Body composition Nutrition Low calcium intake, High caffeine intake, High sodium intake, High animal protein intake Lifestyles Cigarette use, High alcoholic intake, Low level of physical activity Drug Heparin, Anticonculsants, Immunosuppressants Chemotherapy, Corticosteroids, Thyroid hormone Risk factors for Low Bone Density Smoker Age (per 5 years) Maternal history of fx Steroid use Caffeine intake Activity score Age at menopause Milk intake Ever pregnant Surgical menopause Waist/hip ratio Weight Grip strength Height Thiazide use Oestrogen use -8 -6 -4 -2 0 2 Percent change in BMD 4 6 8 Genetics of Bone Mineral Density Clues to Genetics and Environment Epidemiol characteristics Geographic variation Ethnic variation Temporal variation Epidemics Social class variation Gender variation Age Family variables History of disease Birth order Birth interval Co-habitation Genetics + + +/+ +/+ +/- Environment + + + + + + + + + + + Methods of Investigation • Family studies. Examine phenotypes (diseases) in the relatives of affected subjects (probands). • Twin studies. Examine the intraclass correlation between MZ (who share 100% genotypes) and DZ twins (who share 50% genotypes). • Adoption studies. Seek to distinguish genetic from environmental effects by comparing phenotypes in children more closely resemble their biological than adoptive parents. • Offspring of discordant MZ twins. Control for environmental effect; test for large genetic contribution to etiology. Basic Genetic Model Phenotype (P) = Genetics + Environment Genetics = Additive (A) + Dominant (D) Environment = Common (C) + Specific (E) => P = A + D + C + E Statistical Genetic Model Cov(Yi,Yj) = 2Fijs2(a) + Dijs2(d) + gijs2(c) + dijs2(e) Fij : kinship coefficient Dij : Jacquard’s coefficient of identical-by-descent gij : Probability of sharing environmental factors dij : Residual coefficient VP = VA + VD + VC + VE V = variance; P = Phenotype; A, D, C, E = as defined Expected Kinship Coefficients Expected coefficient for Relative Spouse-spouse Parent-offspring Full sibs Half-sibs Aunt-niece First cousins Dizygotic twins Monozygotic twins s2(a) 0 1/2 1/2 1/4 1/4 1/8 1/2 1 s2(d) 0 0 1/4 0 0 0 1/4 1 s2(c) 1 1 1 1 1 0 1 1 A Genetic Model for Twins Study r=1 r = .5 / .25 r = 1 / .5 E1 C1 a c D1 d Twin 1 A1 e A2 D2 a C2 d c E2 e Twin 2 A=additive; D=dominant; C=common environment; E=specific environment Intraclass Correlation: Femoral Neck BMD MZ 1.4 1.4 rMZ = 0.73 rMZ = 0.47 1.3 1.2 1.2 1.1 1.1 1 Twin 2 Twin 2 1.3 DZ 0.9 1 0.9 0.8 0.8 0.7 0.7 0.6 0.6 0.5 0.5 0.5 0.6 0.7 0.8 0.9 1 1.1 1.2 1.3 1.4 0.5 0.6 0.7 0.8 0.9 1 1.1 1.2 1.3 1.4 Twin 1 Twin 1 Genetic Determination of Lean, Fat and Bone Mass rMZ rDZ H2 (%) Lumar spine BMD 0.74 (0.06) 0.48 (0.10) 77.8 Femoral neck BMD 0.73 (0.06) 0.47 (0.11) 76.4 Total body BMD 0.80 (0.05) 0.48 (0.10) 78.6 Lean mass 0.72 (0.06) 0.32 (0.12) 83.5 Fat mass 0.62 (0.08) 0.30 (0.12) 64.8 rMZ and rDZ are shown in coefficient of correlation and standard error in brackets; H2, Heritability index: proportion of variance of a traited attributed to genetic factors Multivariate Analysis: The Cholesky Decomposition Model G1 G2 G3 G4 Lean mass Fat mass LS BMD FN BMD TB BMD E3 E4 E5 E1 E2 G5 LS=lumbar spine, FN=femoral neck, TB=total body, BMD = bone mineral density Genetic and Environmental Correlation between Lean, Fat and Bone Mass LM Lean mass (LM) FM LS 0.52 0.39 0.23 0.51 0.41 0.36 0.70 0.57 0.70 Ft mass (FM) 0.16 Lumbar spine BMD (LS) 0.08 0.02 Femoral neck BMD (FN) 0.16 0.05 0.64 Total body BMD (TB) 0.09 0.31 0.75 FN TB 0.61 0.58 Upper diagonal: genetic correlation; lower diagonal: environmental correlation LS=lumbar spine, FN=femoral neck, TB=total body, BMD = bone mineral density Strategies for Finding Genes How many genes ? • Initial estimate: 120,000. • DNA sequence: 60,000 - 70,000. • Estimates from the Human Genome Project: 32,000 - 39,000 (including non-functional genes = inactive genes). • Osteporosis genes = 50 - 70 (?) Distribution of the number of genes Polygenes Number of genes Oligogenes Major genes Effect size Finding genes: a challenge One of the most difficult challenges ahead is to find genes involved in diseases that have a complex pattern of inheritance, such as those that contribute to osteoporosis, diabetes, asthma, cancer and mental illness. Why search for genes? • Scientific value • Study genes’ actions at the molecular level • Therapeutic value • Gene product and development of new drugs; • Gene therapy • Public health value • Identification of “high-risk” individuals • Interaction between genes and environment Genomewise screening vs Candidate gene • Genome-wide screening approach • No physiological assumption • Systematic screening for chromosomal regions of interest in the entire genome • Candidate gene approach • Proven or hypothetical physiological mechanism • Direct test for individual genes Linkage vs Association • Linkage – traces cosegregation and recombination phenomena between observed markers and unobserved putative trait. Significance is shown by a LOD (log-odds) score. • Association – compares the frequencies of alleles between unrelated cases (diseased) and controls. • Transmission disequilibrium test (TDT) – examines the transmission of alleles from heterozygous parents to those children exhibiting the phenotype of interest. Two-point linkage analysis: an example D 142 D d 134 142 138 /142 ?? 134 /142 142 /146 142 /154 Non Rec 134 / 146 Non 142 / 154 Non 146 / 154 134 / 146 Non 134 / 154 134 / 146 134 / 154 Non Non = non-recombination; Rec = recombination Rec Non No linkage D Complete linkage d D d 134 1/4 1/4 134 0 1/2 142 1/4 1/4 142 1/2 0 Incomplete linkage 134 D d q/2 (1-q)/2 6 LOD log 10 142 (1-q)/2 1 θ θ 2 2 8 1 4 q/2 q: Recombination fraction 2 Estimation of the recombination fraction q Max LOD score +6 +4 LOD score +2 0 -2 -4 -6 0 0.1 0.2 0.3 0.4 Estimated value of q 0.5 A model for sibpair linkage analysis Xi1 = value of sib 1; Xi2 = value of sib 2 Di = abs(Xi1 - Xi2)2 pi = probability of genes shared identical-by-descent E(Di | pi) = a + b pi If b = 0 If b < 0 => => s2(g) = 0; q = 0.5, i.e. No linkage s2(g) > 0; q ne 0.5, i.e. Linkage Behav Genet 1972; 2:3-19 Identical-by-Descent (IBD) 126 / 130 126 / 134 A 126 / 138 B 134 / 138 130 / 134 C 130 / 138 D 126 / 138 E Alleles ibd if they are identical and descended from the same ancestral allele • A and D share no alleles • A, B and E share 1 allele (126) ibd; C vs D; A vs C; B, D and E • B and E share 2 (126 and 138) alleles ibd Sibpair linkage analysis: an example 25 20 o o ooo oo o o o oo oo oo o o o oo oo oo o o Intrapair difference in BMD (%) Squared difference in BMD among siblings 15 10 5 0 0 1 2 Number of alleles shared IBD 0 1 Alleles shared IBD Nature 1994; 367:284-287 2 Association analysis: an example 1.1 g/cm2 1 0.9 0.8 BB Bb bb VDR genotype Association between vitamin D receptor gene and bone mineral density Candidate BMD genes : association analysis Location Name Symbol 1q25 2q13 3q21-24 3q27 4q11-13 4q21 5q31 6q25.1 7p21 7q21.3 7q22 11p15 12q13 17q22 19q13 19q13 Osteocalcin IL-1 Receptor Antagonist Calcium Sensing Receptor a2HS Glycoprotein Vitamin D binding protein Osteopontin Osteonectin Estrogen receptor a Interleukin-6 Calcitonin receptor Collagen type Ia2 Parathyroid hormone Vitamin D receptor Collagen Type Ia1 Transforming growth factor b1 Apolipoprotein E BGLAP CASR CASR AHSG DBP/GCv SPP1 SPOCK ESRa IL-6 CALCR COLIA2 PTH VDR COLIA1 TGF-b1 ApoE Localization of BMD genes in humans Some notable genes • • • • • Vitamin D receptor (VDR) Collagen I alpha 1 (COLIA1) Estrogen receptor (ER) Interleukin-6 (IL6) Transforming growth factor b (TGFb) Problems • None of the candidate genes have clinically meaningful effect on BMD. • Inconsistent (even conflicting) results. • Past studies have suffered serious problems in experimental design and methodology. – – – – Association Inadequate sample size Univariate analysis Sibpair analysis New paradigms • Sampling design – large multi-generational families • Phenotypes – consideration of multitraits rather than a single trait. • Analysis – Combine linkage and association analyses • Animal model – Mouse genome and transgenic model Summary • Fracture is an ultimate and clinically relevant outcome of osteoporosis. • BMD is a primary predictor of fracture. • Variation in BMD is largely determined by genetic factors. • The search for specific genes that are linked to BMD has not been successful nor productive. Perspective • Can genes be found? – Definitely. • The Human Genome Project role? – Very helpful. • Influences of biotechnology? – Great realization. • Gene therapy? – Quite possible. • Lôøi queâ (genes) chaép nhaët doâng daøi • Mua vui cuõng ñöôïc moät vaøi troáng canh (phuùt giaây) • Nguyeãn Du