* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download poster_CSHL_2007
Human genome wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Epigenomics wikipedia , lookup
Genome evolution wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Epigenetics of human development wikipedia , lookup
History of genetic engineering wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Metagenomics wikipedia , lookup
Microevolution wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Designer baby wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Microsatellite wikipedia , lookup
History of RNA biology wikipedia , lookup
Epitranscriptome wikipedia , lookup
Non-coding DNA wikipedia , lookup
Primary transcript wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Helitron (biology) wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Non-coding RNA wikipedia , lookup
Mir-92 microRNA precursor family wikipedia , lookup
RNA silencing wikipedia , lookup
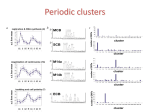
Systematic exploration of cis-regulation using a generic computational framework Olivier Elemento*, Noam Slonim* (equal contribution) and Saeed Tavazoie Lewis-Sigler Institute for Integrative Genomics, Princeton University FIRE FIRE What is FIRE ? FIRE Yeast: 78 co-expression clusters From k-mers to motifs (data from Gasch et al, 2000) FIRE (for Finding Informative Regulatory Elements) is a highly sensitive approach for motif discovery from expression data, based on mutual information. It has the following characteristics: Up-regulated Cy3/Cy5 log-ratios PAC Rpn4 Similarity to ChIPchip RAP1 motif (Lee et al, 2002) • applicable to any type of expression data, • obviates assumptions and parameter tuning often required by existing methods, Yap1 change Puf3 PAC RRPE • simultaneously finds DNA and RNA motifs and explores their functional relationships, v) scales well to mammalian genomes, • characterizes motif interactions and co-localizations Down-regulated Mutual information • highly sensitive, with very few false positive predictions, if any, • highlights the biological role of predicted motifs, their inter-species conservation, and spatial and orientation biases, Yeast: single microarray Motif conservation with S. bayanus Experiment: H2O2 treatment in ΔMsn2/ΔMsn4 background PUF4 PUF3 MSN2/4 Human gene expression atlas (clustered) Human: 78 tissues (Su et al, 2004) Statistical significance • displays the results in a user-friendly graphical format. (data from Su et al, 2004) RAP1 17 motifs in 5’ upstream regions 6 motifs in 3’UTRs Maximum of 10,000 expression-shuffled mutual information values RPN4 REB1 0 “motifs” when shuffling the gene labels of the clustering partition ELK4 73 motifs in 5’ upstream regions 42 motifs in 3’UTRs MBP1 HAP4 1129 motifs when applying AlignACE (with default parameters) to each cluster independently Sp1 0 “motifs” when shuffling the gene labels of the clustering partition miR-525/mR-526c FIRE uses mutual information to discover and characterize motifs Real mutual information value XBP1 880 “motifs” when applying AlignACE to the same shuffled clusters as above bZIP911 NF-Y Several 3’UTR motifs match the 5’ extremity of microRNAs BAS1 CBF1 All 23 motifs are highly conserved with S. bayanus E2F1 miR-200b/miR-429 Discrete Cluster index 5’ upstream region 0 6.45 0 4.39 0 3.50 0 1.98 1.54 1 0.45 1 -1.56 2 -2.32 2 -2.89 Mutual Information I ( X ;Y ) P( x, y ) log Y P ( x, y ) P( x ) P( y ) Co-occurrence 0 0 0 0 1 1 motifs informative about the phase ? -0.87 2 Cluster index (Data from Bozdech, Llinás, et al, 2003) -0.08 1 Position bias P. falciparum: intra-erythrocytic development cycle 0.01 1 X SWI4 0.12 1 5’ upstream region Log-ratio 5’ upstream region 0h Time 48h ~ 2,700 periodically expressed genes 5’ upstream region Continuous -π Phase > 50% of our predicted motifs have a non-random spatial distribution 2 2 2 2 Stastical significance Pax2 E2F CHOP-C/EBPα TCF11-MafG The RAP1 binding site has a position and orientation bias … +π -π Phase Biological insights +π 21 motifs in 5’ upstream regions 0 motifs in 3’UTRs 0 “motifs” when shuffling the gene labels of the phase profile • Importance of RNA motifs in shaping transcriptomes (~30% of yeast, worm, human, arabidopsis motifs we found are RNA motifs) 71% highly conserved with P. yoelli • In worm/human/mouse, several RNA motifs match miRNA targets • “Cooperation” between DNA and RNA motifs • Avoidance of joint-presence for certain motifs DNA replication, p<1e-4 plastid, p<0.01 • Under-representation of certain motifs ribosome, p<0.001 Bozdech, Llinás, et al, 2003 Practical aspects 1 1 TCF11-MafG PAC and the Msn2/4 binding site tend to avoid being in the same promoters PAC and RRPE tend be colocalize on the DNA Unix command line: perl fire.pl –expfile=human_clusters.txt –exptype=discrete –species=human